Key Points
-
Transcription factors of the interferon (IFN)-regulatory factor (IRF) family were originally identified by their capacity to mediate the effects of type I IFNs. Ten IRF-family members have now been defined on the basis of sequence homology and the ability to bind a common DNA motif in various gene promoters.
-
In the past seven years, many IRFs have been found to be involved in T helper (TH)-cell differentiation, a consequence of their capacity to influence the function of accessory cells. This influence leads to alterations in gene products that are important for differentiation into TH1 or TH2 cells, such as interleukin-12 (IL-12), IL-18, IL-23, type I IFNs and nitric oxide.
-
Recent evidence has shown an additional T-cell-intrinsic role for IRF1, -2 and -4 during TH-cell differentiation. This role can be shown even when highly purified naive TH cells are studied in the absence of any accessory cell.
-
IRF1 and IRF2 interact with each other and bind elements in the IL-4 promoter, suppressing its activity.
-
Evidence is accumulating that IRF1 is a multifunctional transcription factor that influences several different cell types and genes to modify the differentiation of TH cells into either TH1 or TH2 cells. Importantly, all of the activities of IRF1 are directed towards a TH1 response, making IRF1 an attractive target for therapeutic intervention in diseases with increased or deficient TH1-cell responses, such as multiple sclerosis or asthma.
-
IRF4 mainly drives TH2-cell development but might also influence TH1-cell development. IRF4 acts intrinsically on TH cells, functions upstream of GATA-binding protein 3 (GATA3) and might interact with signal transducer and activator of transcription 6 (STAT6), B-cell lymphoma 6 (BCL-6) and/or nuclear factor of activated T cells 1 (NFAT1) and/or NFAT2.
Abstract
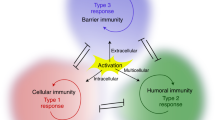
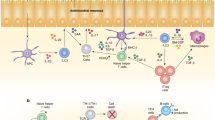
Members of the interferon-regulatory factor family of transcription factors have long been known to be intracellular mediators of the effects of interferons. In recent years, interferon-regulatory factors have also been shown to have an essential role in the differentiation of T helper cells, both by modulating the functions of antigen-presenting cells and by having direct effects on the T helper cells themselves. Depending on the interferon-regulatory factor involved, the differentiation of T helper cells to either T helper 1 cells or T helper 2 cells can be influenced. In this article, we provide an overview of this relatively new and still underappreciated role of interferon-regulatory factors.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Miyamoto, M. et al. Regulated expression of a gene encoding a nuclear factor, IRF-1, that specifically binds to IFN-β gene regulatory elements. Cell 54, 903–913 (1988).
Harada, H. et al. Structurally similar but functionally distinct factors, IRF-1 and IRF-2, bind to the same regulatory elements of IFN and IFN-inducible genes. Cell 58, 729–739 (1989).
Tanaka, N., Kawakami, T. & Taniguchi, T. Recognition DNA sequences of interferon regulatory factor 1 (IRF-1) and IRF-2, regulators of cell growth and the interferon system. Mol. Cell. Biol. 13, 4531–4538 (1993).
Darnell, J. E., Kerr, I. M. & Stark, G. R. Jak–STAT pathways and transcriptional activation in response to IFNs and other extracellular signaling proteins. Science 264, 1415–1421 (1994).
Au, W. C., Moore, P. A., Lowther, W., Juang, Y. T. & Pitha, P. M. Identification of a member of the interferon regulatory factor family that binds to the interferon-stimulated response element and activates expression of interferon-induced genes. Proc. Natl Acad. Sci. USA 92, 11657–11661 (1995).
Barnes, B. J., Moore, P. A. & Pitha, P. M. Virus-specific activation of a novel interferon regulatory factor, IRF-5, results in the induction of distinct interferon α genes. J. Biol. Chem. 276, 23382–23390 (2001).
Driggers, P. H. et al. An interferon γ-regulated protein that binds the interferon-inducible enhancer element of major histocompatibility complex class I genes. Proc. Natl Acad. Sci. USA 87, 3743–3747 (1990).
Eisenbeis, C. F., Singh, H. & Storb, U. Pip, a novel IRF family member, is a lymphoid-specific, PU.1-dependent transcriptional activator. Genes Dev. 9, 1377–1387 (1995).
Grant, C. E., Vasa, M. Z. & Deeley, R. G. cIRF-3, a new member of the interferon regulatory factor (IRF) family that is rapidly and transiently induced by dsRNA. Nucleic Acids Res. 23, 2137–2146 (1995).
Hatada, S. et al. An interferon regulatory factor-related gene (xIRF-6) is expressed in the posterior mesoderm during the early development of Xenopus laevis. Gene 203, 183–188 (1997).
Matsuyama, T. et al. Molecular cloning of LSIRF, a lymphoid-specific member of the interferon regulatory factor family that binds the interferon-stimulated response element (ISRE). Nucleic Acids Res. 23, 2127–2136 (1995).
Nehyba, J., Hrdlickova, R., Burnside, J. & Bose, H. R. A novel interferon regulatory factor (IRF), IRF-10, has a unique role in immune defense and is induced by the v-Rel oncoprotein. Mol. Cell. Biol. 22, 3942–3957 (2002).
Schindler, C., Fu, X. Y., Improta, T., Aebersold, R. & Darnell, J. E. Proteins of transcription factor ISGF-3: one gene encodes the 91- and 84-kDa ISGF-3 proteins that are activated by interferon α. Proc. Natl Acad. Sci. USA 89, 7836–7839 (1992).
Yamagata, T. et al. A novel interferon regulatory factor family transcription factor, ICSAT/Pip/LSIRF, that negatively regulates the activity of interferon-regulated genes. Mol. Cell. Biol. 16, 1283–1294 (1996).
Zhang, L. & Pagano, J. S. IRF-7, a new interferon regulatory factor associated with Epstein–Barr virus latency. Mol. Cell. Biol. 17, 5748–5757 (1997).
Hobart, M., Ramassar, V., Goes, N., Urmson, J. & Halloran, P. F. IFN regulatory factor-1 plays a central role in the regulation of the expression of class I and II MHC genes in vivo. J. Immunol. 158, 4260–4269 (1997).
Kamijo, R. et al. Requirement for transcription factor IRF-1 in NO synthase induction in macrophages. Science 263, 1612–1615 (1994).
Tanaka, N. et al. Cooperation of the tumour suppressors IRF-1 and p53 in response to DNA damage. Nature 382, 816–818 (1996).
Taniguchi, T., Ogasawara, K., Takaoka, A. & Tanaka, N. IRF family of transcription factors as regulators of host defense. Annu. Rev. Immunol. 19, 623–655 (2001).
Holtschke, T. et al. Immunodeficiency and chronic myelogenous leukemia-like syndrome in mice with a targeted mutation of the ICSBP gene. Cell 87, 307–317 (1996).
Meraro, D. et al. Protein–protein and DNA–protein interactions affect the activity of lymphoid-specific IFN regulatory factors. J. Immunol. 163, 6468–6478 (1999).
Barber, S. A., Fultz, M. J., Salkowski, C. A. & Vogel, S. N. Differential expression of interferon regulatory factor 1 (IRF-1), IRF-2, and interferon consensus sequence binding protein genes in lipopolysaccharide (LPS)-responsive and LPS-hyporesponsive macrophages. Infect. Immun. 63, 601–608 (1995).
Galon, J., Sudarshan, C., Ito, S., Finbloom, D. & O'Shea, J. J. IL-12 induces IFN regulating factor-1 (IRF-1) gene expression in human NK and T cells. J. Immunol. 162, 7256–7262 (1999).
Coccia, E. M., Stellacci, E., Marziali, G., Weiss, G. & Battistini, A. IFN-γ and IL-4 differently regulate inducible NO synthase gene expression through IRF-1 modulation. Int. Immunol. 12, 977–985 (2000).
Elser, B. et al. IFN-γ represses IL-4 expression via IRF-1 and IRF-2. Immunity 17, 703–712 (2002).
Pine, R., Canova, A. & Schindler, C. Tyrosine phosphorylated p91 binds to a single element in the ISGF2/IRF-1 promoter to mediate induction by IFN α and IFN γ, and is likely to autoregulate the p91 gene. EMBO J. 13, 158–167 (1994).
Remoli, M. E. et al. Selective expression of type I IFN genes in human dendritic cells infected with Mycobacterium tuberculosis. J. Immunol. 169, 366–374 (2002).
Lin, R. & Hiscott, J. A role for casein kinase II phosphorylation in the regulation of IRF-1 transcriptional activity. Mol. Cell. Biochem. 191, 169–180 (1999).
Nelson, N. et al. Expression of IFN regulatory factor family proteins in lymphocytes. Induction of Stat-1 and IFN consensus sequence binding protein expression by T cell activation. J. Immunol. 156, 3711–3720 (1996).
Watanabe, N., Sakakibara, J., Hovanessian, A. G., Taniguchi, T. & Fujita, T. Activation of IFN-β element by IRF-1 requires a posttranslational event in addition to IRF-1 synthesis. Nucleic Acids Res. 19, 4421–4428 (1991).
Harada, H. et al. Structure and regulation of the human interferon regulatory factor 1 (IRF-1) and IRF-2 genes: implications for a gene network in the interferon system. Mol. Cell. Biol. 14, 1500–1509 (1994).
Matsuyama, T. et al. Targeted disruption of IRF-1 or IRF-2 results in abnormal type I IFN gene induction and aberrant lymphocyte development. Cell 75, 83–97 (1993).
Yamamoto, H., Lamphier, M. S., Fujita, T., Taniguchi, T. & Harada, H. The oncogenic transcription factor IRF-2 possesses a transcriptional repression and a latent activation domain. Oncogene 9, 1423–1428 (1994).
Jesse, T. L., LaChance, R., Iademarco, M. F. & Dean, D. C. Interferon regulatory factor-2 is a transcriptional activator in muscle where it regulates expression of vascular cell adhesion molecule-1. J. Cell Biol. 140, 1265–1276 (1998).
Childs, K. S. & Goodbourn, S. Identification of novel co-repressor molecules for interferon regulatory factor-2. Nucleic Acids Res. 31, 3016–3026 (2003).
Mamane, Y. et al. Interferon regulatory factors: the next generation. Gene 237, 1–14 (1999).
Qin, B. Y. et al. Crystal structure of IRF-3 reveals mechanism of autoinhibition and virus-induced phosphoactivation. Nature Struct. Biol. 10, 913–921 (2003).
Takahasi, K. et al. X-ray crystal structure of IRF-3 and its functional implications. Nature Struct. Biol. 10, 922–927 (2003).
Fitzgerald, K. A. et al. IKKε and TBK1 are essential components of the IRF3 signaling pathway. Nature Immunol. 4, 491–496 (2003).
McWhirter, S. M. et al. IFN-regulatory factor 3-dependent gene expression is defective in Tbk1-deficient mouse embryonic fibroblasts. Proc. Natl Acad. Sci. USA 101, 233–238 (2004).
Sharma, S. et al. Triggering the interferon antiviral response through an IKK-related pathway. Science 300, 1148–1151 (2003).
Karpova, A. Y., Trost, M., Murray, J. M., Cantley, L. C. & Howley, P. M. Interferon regulatory factor-3 is an in vivo target of DNA-PK. Proc. Natl Acad. Sci. USA 99, 2818–2823 (2002).
Marecki, S., Atchison, M. L. & Fenton, M. J. Differential expression and distinct functions of IFN regulatory factor 4 and IFN consensus sequence binding protein in macrophages. J. Immunol. 163, 2713–2722 (1999).
Grossman, A. et al. Cloning of human lymphocyte-specific interferon regulatory factor (hLSIRF/hIRF4) and mapping of the gene to 6p23–p25. Genomics 37, 229–233 (1996).
Grumont, R. J. & Gerondakis, S. Rel induces interferon regulatory factor 4 (IRF-4) expression in lymphocytes: modulation of interferon-regulated gene expression by Rel/nuclear factor κB. J. Exp. Med. 191, 1281–1292 (2000).
Gupta, S., Jiang, M., Anthony, A. & Pernis, A. B. Lineage-specific modulation of interleukin 4 signaling by interferon regulatory factor 4. J. Exp. Med. 190, 1837–1848 (1999).
Lau, J. F., Parisien, J. P. & Horvath, C. M. Interferon regulatory factor subcellular localization is determined by a bipartite nuclear localization signal in the DNA-binding domain and interaction with cytoplasmic retention factors. Proc. Natl Acad. Sci. USA 97, 7278–7283 (2000).
Brass, A. L., Kehrli, E., Eisenbeis, C. F., Storb, U. & Singh, H. Pip, a lymphoid-restricted IRF, contains a regulatory domain that is important for autoinhibition and ternary complex formation with the Ets factor PU.1. Genes Dev. 10, 2335–2347 (1996).
Barnes, B. J. et al. Global and distinct targets of IRF-5 and IRF-7 during innate response to viral infection. J. Biol. Chem. 279, 45194–45207 (2004).
Barnes, B. J., Kellum, M. J., Field, A. E. & Pitha, P. M. Multiple regulatory domains of IRF-5 control activation, cellular localization, and induction of chemokines that mediate recruitment of T lymphocytes. Mol. Cell. Biol. 22, 5721–5740 (2002).
Barnes, B. J., Field, A. E. & Pitha-Rowe, P. M. Virus-induced heterodimer formation between IRF-5 and IRF-7 modulates assembly of the IFNA enhanceosome in vivo and transcriptional activity of IFNA genes. J. Biol. Chem. 278, 16630–16641 (2003).
Kondo, S. et al. Mutations in IRF6 cause Van der Woude and popliteal pterygium syndromes. Nature Genet. 32, 285–289 (2002).
Izaguirre, A. et al. Comparative analysis of IRF and IFN-α expression in human plasmacytoid and monocyte-derived dendritic cells. J. Leukoc. Biol. 74, 1125–1138 (2003).
Lu, R., Au, W. C., Yeow, W. S., Hageman, N. & Pitha, P. M. Regulation of the promoter activity of interferon regulatory factor-7 gene. Activation by interferon and silencing by hypermethylation. J. Biol. Chem. 275, 31805–31812 (2000).
Hemmi, H., Kaisho, T., Takeda, K. & Akira, S. The roles of Toll-like receptor 9, MyD88, and DNA-dependent protein kinase catalytic subunit in the effects of two distinct CpG DNAs on dendritic cell subsets. J. Immunol. 170, 3059–3064 (2003).
Kerkmann, M. et al. Activation with CpG-A and CpG-B oligonucleotides reveals two distinct regulatory pathways of type I IFN synthesis in human plasmacytoid dendritic cells. J. Immunol. 170, 4465–4474 (2003).
Lin, R., Mamane, Y. & Hiscott, J. Multiple regulatory domains control IRF-7 activity in response to virus infection. J. Biol. Chem. 275, 34320–34327 (2000).
Marie, I., Durbin, J. E. & Levy, D. E. Differential viral induction of distinct interferon-α genes by positive feedback through interferon regulatory factor-7. EMBO J. 17, 6660–6669 (1998).
Sato, M. et al. Positive feedback regulation of type I IFN genes by the IFN-inducible transcription factor IRF-7. FEBS Lett. 441, 106–110 (1998).
Au, W. C., Yeow, W. S. & Pitha, P. M. Analysis of functional domains of interferon regulatory factor 7 and its association with IRF-3. Virology 280, 273–282 (2001).
Lu, R. & Pitha, P. M. Monocyte differentiation to macrophage requires interferon regulatory factor 7. J. Biol. Chem. 276, 45491–45496 (2001).
Tsujimura, H., Tamura, T. & Ozato, K. IFN consensus sequence binding protein/IFN regulatory factor 8 drives the development of type I IFN-producing plasmacytoid dendritic cells. J. Immunol. 170, 1131–1135 (2003). This paper shows that IRF8 is associated with the generation of T H 1-cell-inducing plasmacytoid DCs.
Tamura, T., Nagamura-Inoue, T., Shmeltzer, Z., Kuwata, T. & Ozato, K. ICSBP directs bipotential myeloid progenitor cells to differentiate into mature macrophages. Immunity 13, 155–165 (2000).
Sharf, R. et al. Phosphorylation events modulate the ability of interferon consensus sequence binding protein to interact with interferon regulatory factors and to bind DNA. J. Biol. Chem. 272, 9785–9792 (1997).
Nelson, N., Marks, M. S., Driggers, P. H. & Ozato, K. Interferon consensus sequence-binding protein, a member of the interferon regulatory factor family, suppresses interferon-induced gene transcription. Mol. Cell. Biol. 13, 588–599 (1993).
Sharf, R. et al. Functional domain analysis of interferon consensus sequence binding protein (ICSBP) and its association with interferon regulatory factors. J. Biol. Chem. 270, 13063–13069 (1995).
Weisz, A. et al. Human interferon consensus sequence binding protein is a negative regulator of enhancer elements common to interferon-inducible genes. J. Biol. Chem. 267, 25589–25596 (1992).
Eklund, E. A., Jalava, A. & Kakar, R. PU. 1, interferon regulatory factor 1, and interferon consensus sequence-binding protein cooperate to increase gp91phox expression. J. Biol. Chem. 273, 13957–13965 (1998).
Levy, D. E., Lew, D. J., Decker, T., Kessler, D. S. & Darnell, J. E. Synergistic interaction between interferon-α and interferon-γ through induced synthesis of one subunit of the transcription factor ISGF3. EMBO J. 9, 1105–1111 (1990).
Lohoff, M. et al. Interferon regulatory factor-1 is required for a T helper 1 immune response in vivo. Immunity 6, 681–689 (1997).
Taki, S. et al. Multistage regulation of TH1-type immune responses by the transcription factor IRF-1. Immunity 6, 673–679 (1997). References 70 and 71 are the first reports to associate IRFs with the T H 1-cell versus T H 2-cell differentiation concept. They describe the marked T H 2-response-prone phenotype of Irf1−/− mice.
Sommer, F. et al. Lack of gastritis and of an adaptive immune response in interferon regulatory factor-1-deficient mice infected with Helicobacter pylori. Eur. J. Immunol. 31, 396–402 (2001).
Liu, J., Cao, S., Herman, L. M. & Ma, X. Differential regulation of interleukin (IL)-12 p35 and p40 gene expression and interferon (IFN)-γ-primed IL-12 production by IFN regulatory factor 1. J. Exp. Med. 198, 1265–1276 (2003).
Maruyama, S. et al. Identification of IFN regulatory factor-1 binding site in IL-12 p40 gene promoter. J. Immunol. 170, 997–1001 (2003).
Salkowski, C. A. et al. IL-12 is dysregulated in macrophages from IRF-1 and IRF-2 knockout mice. J. Immunol. 163, 1529–1536 (1999).
Oppmann, B. et al. Novel p19 protein engages IL-12p40 to form a cytokine, IL-23, with biological activities similar as well as distinct from IL-12. Immunity 13, 715–725 (2000).
Fantuzzi, G. et al. Role of interferon regulatory factor-1 in the regulation of IL-18 production and activity. Eur. J. Immunol. 31, 369–375 (2001).
Niedbala, W. et al. Nitric oxide preferentially induces type 1 T cell differentiation by selectively up-regulating IL-12 receptor β2 expression via cGMP. Proc. Natl Acad. Sci. USA 99, 16186–16191 (2002).
Laskay, T., Rollinghoff, M. & Solbach, W. Natural killer cells participate in the early defense against Leishmania major infection in mice. Eur. J. Immunol. 23, 2237–2241 (1993).
Scharton, T. M. & Scott, P. Natural killer cells are a source of interferon γ that drives differentiation of CD4+ T cell subsets and induces early resistance to Leishmania major in mice. J. Exp. Med. 178, 567–577 (1993).
Ogasawara, K. et al. Requirement for IRF-1 in the microenvironment supporting development of natural killer cells. Nature 391, 700–703 (1998).
Hida, S. et al. CD8+ T cell-mediated skin disease in mice lacking IRF-2, the transcriptional attenuator of interferon-α/β signaling. Immunity 13, 643–655 (2000).
Honda, K., Mizutani, T. & Taniguchi, T. Negative regulation of IFN-α/β signaling by IFN regulatory factor 2 for homeostatic development of dendritic cells. Proc. Natl Acad. Sci. USA 101, 2416–2421 (2004).
Ichikawa, E. et al. Defective development of splenic and epidermal CD4+ dendritic cells in mice deficient for IFN regulatory factor-2. Proc. Natl Acad. Sci. USA 101, 3909–3914 (2004). References 83 and 84 are the first reports to describe the effects of IRF2 on the generation of the B220− CD8α− subset of DCs.
Lohoff, M. et al. Deficiency in the transcription factor interferon regulatory factor (IRF)-2 leads to severely compromised development of natural killer and T helper type 1 cells. J. Exp. Med. 192, 325–336 (2000). This paper describes the T H 2-response-prone phenotype of Irf2−/− mice. It also describes the dysregulated maturation of NK cells in these mice.
Wang, I. M. et al. An IFN-γ-inducible transcription factor, IFN consensus sequence binding protein (ICSBP), stimulates IL-12 p40 expression in macrophages. J. Immunol. 165, 271–279 (2000).
Bovolenta, C. et al. Molecular interactions between interferon consensus sequence binding protein and members of the interferon regulatory factor family. Proc. Natl Acad. Sci. USA 91, 5046–5050 (1994).
Salkowski, C. A., Barber, S. A., Detore, G. R. & Vogel, S. N. Differential dysregulation of nitric oxide production in macrophages with targeted disruptions in IFN regulatory factor-1 and -2 genes. J. Immunol. 156, 3107–3110 (1996).
Lohoff, M. et al. Dysregulated T helper cell differentiation in the absence of interferon regulatory factor 4. Proc. Natl Acad. Sci. USA 99, 11808–11812 (2002).
Rengarajan, J. et al. Interferon regulatory factor 4 (IRF4) interacts with NFATc2 to modulate interleukin 4 gene expression. J. Exp. Med. 195, 1003–1012 (2002). References 89 and 90 are the first descriptions of the requirement for IRF4 in mouse T H 2-cell development. Reference 89 also shows that IRF4 upregulates the expression of GATA3. Reference 90 also indicates that an IRF4–NFAT1 interaction might be the underlying molecular mechanism.
Tominaga, N. et al. Development of TH1 and not TH2 immune responses in mice lacking IFN-regulatory factor-4. Int. Immunol. 15, 1–10 (2003).
Hu, C. M., Jang, S. Y., Fanzo, J. C. & Pernis, A. B. Modulation of T cell cytokine production by interferon regulatory factor-4. J. Biol. Chem. 277, 49238–49246 (2002). This paper shows the relevance of IRF4 for the production of human T H 2 cells.
Lohoff, M. et al. Enhanced TCR-induced apoptosis in interferon regulatory factor 4-deficient CD4+ TH cells. J. Exp. Med. 200, 247–253 (2004).
Serfling, E. et al. The role of NF-AT transcription factors in T cell activation and differentiation. Biochim. Biophys. Acta 1498, 1–18 (2000).
Zhu, J. et al. Growth factor independent-1 induced by IL-4 regulates TH2 cell proliferation. Immunity 16, 733–744 (2002).
Dent, A. L., Hu-Li, J., Paul, W. E. & Staudt, L. M. T helper type 2 inflammatory disease in the absence of interleukin 4 and transcription factor STAT6. Proc. Natl Acad. Sci. USA 95, 13823–13828 (1998).
Rosenbauer, F. et al. Interferon consensus sequence binding protein and interferon regulatory factor-4/Pip form a complex that represses the expression of the interferon-stimulated gene-15 in macrophages. Blood 94, 4274–4281 (1999).
Mittrucker, H. W. et al. Requirement for the transcription factor LSIRF/IRF4 for mature B and T lymphocyte function. Science 275, 540–543 (1997).
Fanzo, J. C., Hu, C. M., Jang, S. Y. & Pernis, A. B. Regulation of lymphocyte apoptosis by interferon regulatory factor 4 (IRF-4). J. Exp. Med. 197, 303–314 (2003).
Suzuki, S. et al. Critical roles of interferon regulatory factor 4 in CD11bhighCD8α− dendritic cell development. Proc. Natl Acad. Sci. USA 101, 8981–8986 (2004).
Fehr, T. et al. Crucial role of interferon consensus sequence binding protein, but neither of interferon regulatory factor 1 nor of nitric oxide synthesis for protection against murine listeriosis. J. Exp. Med. 185, 921–931 (1997).
Giese, N. A. et al. Interferon (IFN) consensus sequence-binding protein, a transcription factor of the IFN regulatory factor family, regulates immune responses in vivo through control of interleukin 12 expression. J. Exp. Med. 186, 1535–1546 (1997). This paper and reference 104 are the first reports to describe the T H 2-response-prone phenotype of Irf8−/− mice. This paper also describes the susceptibility of these mice to infection with L. major and their deficit in IL-12 production. Reference 104 also describes the susceptibility of these mice to infection with Toxoplasma gondii and their deficit in IL-12 production.
Hein, J. et al. Interferon consensus sequence binding protein confers resistance against Yersinia enterocolitica. Infect. Immun. 68, 1408–1417 (2000).
Scharton, K. T., Contursi, C., Masumi, A., Sher, A. & Ozato, K. Interferon consensus sequence binding protein-deficient mice display impaired resistance to intracellular infection due to a primary defect in interleukin 12 p40 induction. J. Exp. Med. 186, 1523–1534 (1997).
Kim, Y. M. et al. Roles of IFN consensus sequence binding protein and PU.1 in regulating IL-18 gene expression. J. Immunol. 163, 2000–2007 (1999).
Wu, C. Y., Maeda, H., Contursi, C., Ozato, K. & Seder, R. A. Differential requirement of IFN consensus sequence binding protein for the production of IL-12 and induction of TH1-type cells in response to IFN-γ. J. Immunol. 162, 807–812 (1999).
Doyle, S. et al. IRF3 mediates a TLR3/TLR4-specific antiviral gene program. Immunity 17, 251–263 (2002).
Fitzgerald, K. A. et al. LPS–TLR4 signaling to IRF-3/7 and NF-κB involves the Toll adapters TRAM and TRIF. J. Exp. Med. 198, 1043–1055 (2003).
Yamamoto, M. et al. A novel Toll/IL-1 receptor domain-containing adapter that preferentially activates the IFN-β promoter in the Toll-like receptor signaling. J. Immunol. 169, 6668–6672 (2002).
Yamamoto, M. et al. Role of adaptor TRIF in the MyD88-independent Toll-like receptor signaling pathway. Science 301, 640–643 (2003).
Barnes, B. J. et al. Global and distinct targets of IRF-5 and IRF-7 during innate response to viral infection. J. Biol. Chem. 279, 45194–45207 (2004).
Fujita, T., Kimura, Y., Miyamoto, M., Barsoumian, E. L. & Taniguchi, T. Induction of endogenous IFN-α and IFN-β genes by a regulatory transcription factor, IRF-1. Nature 337, 270–272 (1989).
Wathelet, M. G. et al. Virus infection induces the assembly of coordinately activated transcription factors on the IFN-β enhancer in vivo. Mol. Cell 1, 507–518 (1998).
Sasaki, S., Amara, R. R., Yeow, W. S., Pitha, P. M. & Robinson, H. L. Regulation of DNA-raised immune responses by cotransfected interferon regulatory factors. J. Virol. 76, 6652–6659 (2002).
Nakao, F. et al. Association of IFN-γ and IFN regulatory factor 1 polymorphisms with childhood atopic asthma. J. Allergy Clin. Immunol. 107, 499–504 (2001).
Cherwinski, H. M., Schumacher, J. H., Brown, K. D. & Mosmann, T. R. Two types of mouse helper T cell clone. III. Further differences in lymphokine synthesis between TH1 and TH2 clones revealed by RNA hybridization, functionally monospecific bioassays, and monoclonal antibodies. J. Exp. Med. 166, 1229–1244 (1987).
Fiorentino, D. F., Bond, M. W. & Mosmann, T. R. Two types of mouse T helper cell. IV. TH2 clones secrete a factor that inhibits cytokine production by TH1 clones. J. Exp. Med. 170, 2081–2095 (1989).
Mosmann, T. R., Cherwinski, H., Bond, M. W., Giedlin, M. A. & Coffman, R. L. Two types of murine helper T cell clone. I. Definition according to profiles of lymphokine activities and secreted proteins. J. Immunol. 136, 2348–2357 (1986).
Sacks, D. & Noben-Trauth, N. The immunology of susceptibility and resistance to Leishmania major in mice. Nature Rev. Immunol. 2, 845–858 (2002).
Murphy, K. M. & Reiner, S. L. The lineage decisions of helper T cells. Nature Rev. Immunol. 2, 933–944 (2002).
Zheng, W. & Flavell, R. A. The transcription factor GATA-3 is necessary and sufficient for TH2 cytokine gene expression in CD4 T cells. Cell 89, 587–596 (1997).
Ouyang, W. et al. Stat6-independent GATA-3 autoactivation directs IL-4-independent TH2 development and commitment. Immunity 12, 27–37 (2000).
Takemoto, N. et al. Chromatin remodeling at the IL-4/IL-13 intergenic regulatory region for TH2-specific cytokine gene cluster. J. Immunol. 165, 6687–6691 (2000).
Noben-Trauth, N. et al. An interleukin 4 (IL-4)-independent pathway for CD4+ T cell IL-4 production is revealed in IL-4 receptor-deficient mice. Proc. Natl Acad. Sci. USA 94, 10838–10843 (1997).
Kim, J. I., Ho, I. C., Grusby, M. J. & Glimcher, L. H. The transcription factor c-Maf controls the production of interleukin-4 but not other TH2 cytokines. Immunity 10, 745–751 (1999).
Lighvani, A. A. et al. T-bet is rapidly induced by interferon-γ in lymphoid and myeloid cells. Proc. Natl Acad. Sci. USA 98, 15137–15142 (2001).
Szabo, S. J. et al. A novel transcription factor, T-bet, directs TH1 lineage commitment. Cell 100, 655–669 (2000).
Afkarian, M. et al. T-bet is a STAT1-induced regulator of IL-12R expression in naive CD4+ T cells. Nature Immunol. 3, 549–557 (2002).
Mullen, A. C. et al. Hlx is induced by and genetically interacts with T-bet to promote heritable TH1 gene induction. Nature Immunol. 3, 652–658 (2002).
Pflanz, S. et al. IL-27, a heterodimeric cytokine composed of EBI3 and p28 protein, induces proliferation of naive CD4+ T cells. Immunity 16, 779–790 (2002).
Smeltz, R. B., Chen, J., Ehrhardt, R. & Shevach, E. M. Role of IFN-γ in TH1 differentiation: IFN-γ regulates IL-18Rα expression by preventing the negative effects of IL-4 and by inducing/maintaining IL-12 receptor β2 expression. J. Immunol. 168, 6165–6172 (2002).
Bradley, L. M., Dalton, D. K. & Croft, M. A direct role for IFN-γ in regulation of TH1 cell development. J. Immunol. 157, 1350–1358 (1996).
Farrar, J. D. et al. Selective loss of type I interferon-induced STAT4 activation caused by a minisatellite insertion in mouse Stat2. Nature Immunol. 1, 65–69 (2000).
Rogge, L. et al. The role of Stat4 in species-specific regulation of TH cell development by type I IFNs. J. Immunol. 161, 6567–6574 (1998).
Cella, M. et al. Plasmacytoid monocytes migrate to inflamed lymph nodes and produce large amounts of type I interferon. Nature Med. 5, 919–923 (1999).
Maldonado-Lopez, R. et al. CD8α+ and CD8α− subclasses of dendritic cells direct the development of distinct T helper cells in vivo. J. Exp. Med. 189, 587–592 (1999).
Sato, M. et al. Distinct and essential roles of transcription factors IRF-3 and IRF-7 in response to viruses for IFN-α/β gene induction. Immunity 13, 539–548 (2000). This paper provides clear evidence of the interplay of IRF3, -7 and -9 in the production of type I IFNs in response to viral infection. It describes the susceptibility of Irf3−/− mice to viral infection.
Kimura, T. et al. Essential and non-redundant roles of p48 (ISGF3 γ) and IRF-1 in both type I and type II interferon responses, as revealed by gene targeting studies. Genes Cells 1, 115–124 (1996).
Acknowledgements
We thank G. S. Duncan (Ontario Cancer Institute, Toronto, Canada), H.-W. Mittrücker (Max–Planck-Institut, Berlin, Germany), T. Matsuyama (Nagasaki University, Nagasaki, Japan) and D. Ferrick (University of California, Davis, United States) for their excellent cooperation during the past few years. We also thank M. E. Saunders for scientific editing. This work was supported by grants from the Deutsche Forschungsgemeinschaft (Germany) and the Canadian Institutes of Health Research.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Glossary
- VIRUS–IFN AXIS
-
(Virus–interferon axis). After viral infection, constitutively expressed cytoplasmic IFN-regulatory factor 3 (IRF3) translocates to the nucleus to induce transcription of the IFN-α4 and IFN-β genes. In turn, IFN-α4 and IFN-β induce the expression of IRF7 and thereby the production of other subtypes of IFN-α.
- PEST DOMAIN
-
(Proline-, glutamic-acid-, serine- and threonine-rich domain). A domain that supports the interaction of transcription factors of the ETS family with other transcription factors.
- CONCANAVALIN A
-
(con A). A plant lectin that functions as a T-cell mitogen.
- NUCLEAR-LOCALIZATION SIGNAL
-
A positively charged region of a target protein that binds a cytoplasmic transporter protein. It is responsible for directing the transport of the target protein through nuclear membrane pores and into the nucleus.
- PLASMACYTOID DCS
-
(Plasmacytoid dendritic cells). Immature DCs with a plasmacytoid morphology (that is, similar to plasmablasts), which produce type I interferons in response to viral infection.
- ACTIVATION-INDUCED CELL DEATH
-
(AICD). Apoptotic cell death that is triggered by cellular activation. For example, the death undergone by T helper cells that are activated by CD3-specific antibodies.
- ETS SITE
-
A DNA-binding motif that is recognized by transcription factors of the ETS family. It is a purine-rich sequence that contains the core motif GGAA/T.
Rights and permissions
About this article
Cite this article
Lohoff, M., Mak, T. Roles of interferon-regulatory factors in T-helper-cell differentiation. Nat Rev Immunol 5, 125–135 (2005). https://doi.org/10.1038/nri1552
Issue Date:
DOI: https://doi.org/10.1038/nri1552
This article is cited by
-
Targeting USP2 regulation of VPRBP-mediated degradation of p53 and PD-L1 for cancer therapy
Nature Communications (2023)
-
Regulation of effector and memory CD8 + T cell differentiation: a focus on orphan nuclear receptor NR4A family, transcription factor, and metabolism
Immunologic Research (2023)
-
IRF2 maintains the stemness of colonic stem cells by limiting physiological stress from interferon
Scientific Reports (2020)
-
Genetically engineered mesenchymal stem cells: targeted delivery of immunomodulatory agents for tumor eradication
Cancer Gene Therapy (2020)
-
Uncovering Sub-Structure and Genomic Profiles in Across-Countries Subpopulations of Angus Cattle
Scientific Reports (2020)