Key Points
-
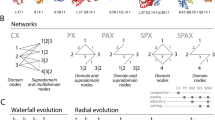
Biomolecular networks, such as protein–protein interaction (PPI) or metabolic networks, organize the 'parts lists' generated by various large-scale approaches and are therefore frameworks that facilitate many discoveries in molecular biology. Nodes represent proteins (specifically enzymes in metabolic networks), whereas PPIs in PPI networks and enzyme–enzyme interactions through shared metabolites in metabolic networks are considered as links.
-
Using this framework, general knowledge on the topology of networks can be applied. However, specification of biomolecular networks, such as the impact of the environment and experimental conditions, also have to be taken into account. The experimental conditions raise important issues regarding the accuracy and coverage of these networks, which also have an impact on the conclusions about the evolution of the networks.
-
The evolutionary dynamics of PPI and metabolic networks is mostly based on two classes of genetic events. The first is duplication and loss of regions encompassing complete genes, which implies the addition and loss of nodes and links. The second is more fine-tuned and includes point mutations, small insertions or deletions, and mutations that affect the regulation of genes, which implies the addition and loss of links.
-
Owing to different biological functions and distinct topological features of PPI and metabolic networks, changes of nodes and links in each are subject to different selection.
-
So far, most of the research on networks is devoted to in vitro and static networks, and these are usually considered in two dimensions (2D networks) — that is, without spatial (3D) or temporal (4D) resolution. Many network features and their evolution can be understood only when taking spatiotemporal resolution into account.
Abstract

Despite only becoming popular at the beginning of this decade, biomolecular networks are now frameworks that facilitate many discoveries in molecular biology. The nodes of these networks are usually proteins (specifically enzymes in metabolic networks), whereas the links (or edges) are their interactions with other molecules. These networks are made up of protein–protein interactions or enzyme–enzyme interactions through shared metabolites in the case of metabolic networks. Evolutionary analysis has revealed that changes in the nodes and links in protein–protein interaction and metabolic networks are subject to different selection pressures owing to distinct topological features. However, many evolutionary constraints can be uncovered only if temporal and spatial aspects are included in the network analysis.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Watts, D. J. & Strogatz, S. H. Collective dynamics of 'small-world' networks. Nature 393, 440–442 (1998).
Uetz, P. et al. A comprehensive analysis of protein–protein interactions in Saccharomyces cerevisiae. Nature 403, 623–627 (2000).
Ito, T. et al. A comprehensive two-hybrid analysis to explore the yeast protein interactome. Proc. Natl Acad. Sci. USA 98, 4569–4574 (2001).
Rain, J. C. et al. The protein–protein interaction map of Helicobacter pylori. Nature 409, 211–215 (2001).
Reboul, J. et al. C. elegans ORFeome version 1.1: experimental verification of the genome annotation and resource for proteome-scale protein expression. Nature Genet. 34, 35–41 (2003).
Giot, L. et al. A protein interaction map of Drosophila melanogaster. Science 302, 1727–1736 (2003).
Rual, J. F. et al. Towards a proteome-scale map of the human protein–protein interaction network. Nature 437, 1173–1178 (2005).
Stelzl, U. et al. A human protein–protein interaction network: a resource for annotating the proteome. Cell 122, 957–968 (2005).
Butland, G. et al. Interaction network containing conserved and essential protein complexes in Escherichia coli. Nature 433, 531–537 (2005).
Arifuzzaman, M. et al. Large-scale identification of protein–protein interaction of Escherichia coli K-12. Genome Research 16, 686–691 (2006).
Gavin, A. C. et al. Functional organization of the yeast proteome by systematic analysis of protein complexes. Nature 415, 141–147 (2002).
Ho, Y. et al. Systematic identification of protein complexes in Saccharomyces cerevisiae by mass spectrometry. Nature 415, 180–183 (2002).
Gavin, A. C. et al. Proteome survey reveals modularity of the yeast cell machinery. Nature 440, 631–636 (2006).
Krogan, N. J. et al. Global landscape of protein complexes in the yeast Saccharomyces cerevisiae. Nature 440, 637–643 (2006).
Tarassov, K. et al. An in vivo map of the yeast protein interactome. Science 320, 1465–1470 (2008).
Fell, D. A. & Sauro, H. M. Metabolic control and its analysis. Additional relationships between elasticities and control coefficients. Eur. J. Biochem. 148, 555–561 (1985).
Thomas, S. & Fell, D. A. A computer program for the algebraic determination of control coefficients in metabolic control analysis. Biochem. J. 292, 351–360 (1993).
Durek, P. & Walther, D. The integrated analysis of metabolic and protein interaction networks reveals novel molecular organizing principles. BMC systems biology 2, 100 (2008). Provides the topological differences between PPI and metabolic networks.
Díaz-Mejía, J. J., Pérez-Rueda, E. & Segovia, L. A network perspective on the evolution of metabolism by gene duplication. Genome Biology 8, R26 (2007).
Jensen, R. A. Enzyme recruitment in evolution of new function. Annu. Rev. Microbiol 30, 409–425 (1976).
Feist, A. M. et al. A genome-scale metabolic reconstruction for Escherichia coli K-12 MG1655 that accounts for 1260 ORFs and thermodynamic information. Mol. Syst. Biol. 3, 121 (2007).
Herrgård, M. J. et al. A consensus yeast metabolic network reconstruction obtained from a community approach to systems biology. Nature Biotechnol. 26, 1155–1160 (2008).
Duarte, N. et al. Global reconstruction of the human metabolic network based on genomic and bibliomic data. Proc. Natl Acad. Sci. USA 104, 1777–1782 (2007).
Sharan, R. et al. Conserved patterns of protein interaction in multiple species. Proc. Natl Acad. Sci. USA 102, 1974–1979 (2005).
Kanehisa, M. et al. KEGG for linking genomes to life and the environment. Nucleic Acids Res. 36, D480–D484 (2008).
Joshi-Tope, G. et al. Reactome: a knowledgebase of biological pathways. Nucleic Acids Res. 33, D428–D432 (2005).
Salwinski, L. et al. The database of interacting proteins: 2004 update. Nucleic Acids Res. 32, D449–D451 (2004).
Jensen, L. J. et al. STRING 8 — a global view on proteins and their functional interactions in 630 organisms. Nucleic Acids Res. 37, D412 (2009).
Jensen, L. J. & Bork, P. Biochemistry. Not comparable, but complementary. Science 322, 56–57 (2008).
Yu, H. et al. High-quality binary protein interaction map of the yeast interactome network. Science 322, 104–110 (2008).
von Mering, C. et al. Comparative assessment of large-scale data sets of protein-protein interactions. Nature 417, 399–403 (2002).
Bader, J. S., Chaudhuri, A., Rothberg, J. M. & Chant, J. Gaining confidence in high-throughput protein interaction networks. Nature Biotechnol. 22, 78–85 (2004).
Feist, A. M. & Palsson, B. Ø. The growing scope of applications of genome-scale metabolic reconstructions using Escherichia coli. Nature Biotechnol. 26, 659–667 (2008).
Jeong, H., Tombor, B., Albert, R., Oltvai, Z. N. & Barabási, A. L. The large-scale organization of metabolic networks. Nature 407, 651–654 (2000).
Ravasz, E., Somera, A. L., Mongru, D. A., Oltvai, Z. N. & Barabási, A. L. Hierarchical organization of modularity in metabolic networks. Science 297, 1551–1555 (2002). The first demonstration of a topological analysis for biomolecular networks, suggesting that the metabolic network is scale free.
Barabási, A. L. & Oltvai, Z. N. Network biology: understanding the cell's functional organization. Nature Rev. Genet. 5, 101–113 (2004).
van Noort, V., Snel, B. & Huynen, M. A. The yeast coexpression network has a small-world, scale-free architecture and can be explained by a simple model. EMBO Rep. 5, 280–284 (2004).
Wagner, A. How the global structure of protein interaction networks evolves. Proc. Biol. Sci. 270, 457–466 (2003).
Rison, S. C. & Thornton, J. Pathway evolution, structurally speaking. Current Opinion in Structural Biology 12, 374–382 (2002).
Janga, S. C. & Babu, M. M. Network-based approaches for linking metabolism with environment. Genome Biology 9, 239 (2008).
Schmidt, S., Sunyaev, S., Bork, P. & Dandekar, T. Metabolites: a helping hand for pathway evolution? Trends Biochem. Sci. 28, 336–341 (2003).
Horowitz, N. H. On the evolution of biochemical syntheses. Proc. Natl Acad. Sci. USA 31, 153–157 (1945). Together with reference 43, this paper provides the first evolutionary models of biochemical networks.
Ycas, M. On earlier states of the biochemical system. J. Theor. Biol. 44, 145–160 (1974).
Lazcano, A. & Miller, S. L. On the origin of metabolic pathways. J. Mol. Evol. 49, 424–431 (1999).
Copley, R. & Bork, P. Homology among (βα)8 barrels: implications for the evolution of metabolic pathways. J. Mol. Biol. 303, 627–641 (2000).
Teichmann, S. A. et al. The evolution and structural anatomy of the small molecule metabolic pathways in Escherichia coli. J. Mol. Biol. 311, 693–708 (2001).
Alves, R., Chaleil, R. A. & Sternberg, M. J. Evolution of enzymes in metabolism: a network perspective. J. Mol. Biol. 320, 751–770 (2002).
Raymond, J. & Segrè, D. The effect of oxygen on biochemical networks and the evolution of complex life. Science 311, 1764–1767 (2006).
Borenstein, E., Kupiec, M., Feldman, M. W. & Ruppin, E. Large-scale reconstruction and phylogenetic analysis of metabolic environments. Proc. Natl Acad. Sci. USA 105, 14482–14487 (2008).
Gianoulis, T. A. et al. Quantifying environmental adaptation of metabolic pathways in metagenomics. Proc. Natl Acad. Sci. USA 106, 1374–1379 (2009).
Snel, B., Bork, P. & Huynen, M. A. Genomes in flux: the evolution of archaeal and proteobacterial gene content. Genome Research 12, 17–25 (2002).
Berg, J., Lässig, M. & Wagner, A. Structure and evolution of protein interaction networks: a statistical model for link dynamics and gene duplications. BMC Evol. Biol. 4, 51 (2004).
Campillos, M., Doerks, T., Shah, P. K. & Bork, P. Computational characterization of multiple Gag-like human proteins. Trends Genet. 22, 585–589 (2006).
Liang, H. & Li, W. H. Gene essentiality, gene duplicability and protein connectivity in human and mouse. Trends Genet. 23, 375–378 (2007).
Rambaldi, D., Giorgi, F., Capuani, F., Ciliberto, A. & Ciccarelli, F. Low duplicability and network fragility of cancer genes. Trends Genet. 24, 427–430 (2008).
Molina, N. & van Nimwegen, E. The evolution of domain-content in bacterial genomes. Biology Direct 3, 51 (2008).
Maslov, S., Krishna, S., Pang, T. Y. & Sneppen, K. Toolbox model of evolution of prokaryotic metabolic networks and their regulation. Proc. Natl Acad. Sci. USA 106, 9743–9748 (2009).
Raes, J., Korbel, J. O., Lercher, M. J., von Mering, C. & Bork, P. Prediction of effective genome size in metagenomic samples. Genome Biology 8, R10 (2007).
Sorek, R. et al. Genome-wide experimental determination of barriers to horizontal gene transfer. Science 318, 1449 (2007).
Prachumwat, A. & Li, W. H. Protein function, connectivity, and duplicability in yeast. Mol. Biol. Evo. 23, 30–39 (2006).
Han, J. D. et al. Evidence for dynamically organized modularity in the yeast protein-protein interaction network. Nature 430, 88–93 (2004).
Jeong, H., Mason, S. P., Barabási, A. L. & Oltvai, Z. N. Lethality and centrality in protein networks. Nature 411, 41–42 (2001). First demonstration of a large-scale analysis of protein–protein physical interactions as a biomolecular network.
Wuchty, S. Evolution and topology in the yeast protein interaction network. Genome Research 14, 1310–1314 (2004).
Fraser, H. B. Modularity and evolutionary constraint on proteins. Nature Genet. 37, 351–352 (2005).
Drummond, D. A., Raval, A. & Wilke, C. O. A single determinant dominates the rate of yeast protein evolution. Mol. Biol. Evo. 23, 327–337 (2006).
Ekman, D. et al. What properties characterize the hub proteins of the protein-protein interaction network of Saccharomyces cerevisiae? Genome Biology 7, R45 (2006).
Lu, C. et al. Impacts of yeast metabolic network structure on enzyme evolution. Genome Biology 8, 407 (2007).
Ciccarelli, F. et al. Complex genomic rearrangements lead to novel primate gene function. Genome Research 15, 343–351 (2005).
Kim, P. M., Korbel, J. O. & Gerstein, M. B. Positive selection at the protein network periphery: evaluation in terms of structural constraints and cellular context. Proc. Natl Acad. Sci. USA 104, 20274–20279 (2007).
Roguev, A. et al. Conservation and rewiring of functional modules revealed by an epistasis map in fission yeast. Science 322, 405–410 (2008). Shows rewiring events on the genetic interaction network with large-scale experiments and analysis.
Conaway, R. & Conaway, J. The INO80 chromatin remodeling complex in transcription, replication and repair. Trends Biochem. Sci. 34, 71–77 (2009).
Jin, J. et al. In and out: histone variant exchange in chromatin. Trends Biochem. Sci. 30, 680–687 (2005).
Shevchenko, A. et al. Chromatin central: towards the comparative proteome by accurate mapping of the yeast proteomic environment. Genome Biology 9, R167 (2008).
Lorch, Y., Zhang, M. & Kornberg, R. Histone octamer transfer by a chromatin-remodeling complex. Cell 96, 389–392 (1999).
Park, Y., Chodaparambil, J. V., Bao, Y., McBryant, S. J. & Luger, K. Nucleosome assembly protein 1 exchanges histone H2A-H2B dimers and assists nucleosome sliding. J. Biol. Chem. 280, 1817–1825 (2005).
Park, Y. J. & Luger, K. The structure of nucleosome assembly protein 1. Proc. Natl Acad. Sci. USA 103, 1248–1253 (2006).
Walfridsson, J., Khorosjutina, O., Matikainen, P., Gustafsson, C. M. & Ekwall, K. A genome-wide role for CHD remodelling factors and Nap1 in nucleosome disassembly. EMBO J. 26, 2868–2879 (2007).
Hahn, M. W. & Kern, A. D. Comparative genomics of centrality and essentiality in three eukaryotic protein-interaction networks. Mol. Biol. Evo. 22, 803–806 (2005).
Barton, N. H. & Keightley, P. D. Understanding quantitative genetic variation. Nature Rev. Genet. 3, 11–21 (2002).
Maslov, S. & Sneppen, K. Specificity and stability in topology of protein networks. Science 296, 910–913 (2002).
Zhu, D. & Qin, Z. S. Structural comparison of metabolic networks in selected single cell organisms. BMC Bioinformatics 6, 8 (2005).
Wolf, D. M. & Arkin, A. P. Motifs, modules and games in bacteria. Curr. Opin. Microbiol. 6, 125–134 (2003).
Kreimer, A., Borenstein, E., Gophna, U. & Ruppin, E. The evolution of modularity in bacterial metabolic networks. Proc. Natl Acad. Sci. USA 105, 6976–6981 (2008).
Spirin, V. & Mirny, L. A. Protein complexes and functional modules in molecular networks. Proc. Natl Acad. Sci. USA 100, 12123–12128 (2003).
Spirin, V., Gelfand, M. S., Mironov, A. A. & Mirny, L. A. A metabolic network in the evolutionary context: Multiscale structure and modularity. Proc. Natl Acad. Sci. USA 103, 8774–8779 (2006).
Snel, B. & Huynen, M. A. Quantifying modularity in the evolution of biomolecular systems. Genome Research 14, 391–397 (2004).
Ihmels, J., Levy, R. & Barkai, N. Principles of transcriptional control in the metabolic network of Saccharomyces cerevisiae. Nature Biotechnol. 22, 86–92 (2004).
von Mering, C. et al. Genome evolution reveals biochemical networks and functional modules. Proc. Natl Acad. Sci. USA 100, 15428–15433 (2003).
Yamada, T., Kanehisa, M. & Goto, S. Extraction of phylogenetic network modules from the metabolic network. BMC Bioinformatics 7, 130 (2006).
Campillos, M., von Mering, C., Jensen, L. J. & Bork, P. Identification and analysis of evolutionarily cohesive functional modules in protein networks. Genome Research 16, 374–382 (2006).
Kelley, B. P. et al. Conserved pathways within bacteria and yeast as revealed by global protein network alignment. Proc. Natl Acad. Sci. USA 100, 11394–11399 (2003).
Fokkens, L. & Snel, B. Cohesive versus flexible evolution of functional modules in eukaryotes. PLoS Comput. Biol. 5, e1000276 (2009).
Parter, M., Kashtan, N. & Alon, U. Environmental variability and modularity of bacterial metabolic networks. BMC Evol. Biol. 7, 169 (2007).
Kashtan, N. & Alon, U. Spontaneous evolution of modularity and network motifs. Proc. Natl Acad. Sci. USA 102, 13773–13778 (2005).
Bork, P. & Serrano, L. Towards Cellular Systems in 4D. Cell 121, 507–509 (2005).
Laub, M. T., McAdams, H. H., Feldblyum, T., Fraser, C. M. & Shapiro, L. Global analysis of the genetic network controlling a bacterial cell cycle. Science 290, 2144–2148 (2000).
de Lichtenberg, U., Jensen, L. J., Brunak, S. & Bork, P. Dynamic complex formation during the yeast cell cycle. Science 307, 724–727 (2005). Provides a time-dependent protein interaction network by gene expression, leading to the study of protein complex dynamics.
Jensen, L. J., Jensen, T. S., de Lichtenberg, U., Brunak, S. & Bork, P. Co-evolution of transcriptional and post-translational cell-cycle regulation. Nature 443, 594–597 (2006).
Hooper, S. D. et al. Identification of tightly regulated groups of genes during Drosophila melanogaster embryogenesis. Mol. Syst. Biol. 3, 72 (2007).
Tomancak, P. et al. Systematic determination of patterns of gene expression during Drosophila embryogenesis. Genome Biology 3, 0088 (2002).
Schmid, M. et al. A gene expression map of Arabidopsis thaliana development. Nature Genet. 37, 501–506 (2005).
Haudry, Y. et al. 4DXpress: a database for cross-species expression pattern comparisons. Nucleic Acids Res. 36, D847–D853 (2008).
Berg, J., Tymoczko J., Stryer L. & Clarke N. Biochemistry (W. H. Freeman).
Shyamsundar, R. et al. A DNA microarray survey of gene expression in normal human tissues. Genome Biology 6, R22 (2005).
Saito-Hisaminato, A. et al. Genome-wide profiling of gene expression in 29 normal human tissues with a cDNA microarray. DNA Res. 9, 35–45 (2002).
Erdo˝s, P. & Renyi, A. On the strength of connectedness of a random graph. Acta Math. Hung. 12, 261–267 (1961).
Barabasi, A. L. & Albert, R. Emergence of scaling in random networks. Science 286, 509–512 (1999).
Karp, P. D. et al. Expansion of the BioCyc collection of pathway/genome databases to 160 genomes. Nucleic Acids Res. 33, 6083–6089 (2005).
Roguev, A., Wiren, M., Weissman, J. S. & Krogan, N. J. High-throughput genetic interaction mapping in the fission yeast Schizosaccharomyces pombe. Nature Methods 4, 861–866 (2007).
Siegert, R., Leroux, M. R., Scheufler, C., Hartl, F. U. & Moarefi, I. Structure of the molecular chaperone prefoldin: unique interaction of multiple coiled coil tentacles with unfolded proteins. Cell 103, 621–632 (2000).
Acknowledgements
The authors thank V. van Noort for supporting statistical analysis of the network, J. Muller for supporting gene annotation procedures and all members of the Bork group for critical discussions. The work in the author's laboratory is supported by the BMBF grant Neuronet (17282) and the EU grant Metahit (17286).
Author information
Authors and Affiliations
Supplementary information
41580_2009_BFnrm2787_MOESM3_ESM.pdf
Supplementary information S1 (box) | Data coverage of interaction networks for selected studies and resources (PDF 577 kb)
Related links
Glossary
- Graph theory
-
The study of the properties of graphs. A graph is a mathematical structure used to model the pairwise relationships between objects. It is composed of a collection of nodes (vertices) and links (edges).
- TAP–MS
-
A method used to detect physical protein–protein interactions by a series of affinity column purifications, followed by mass spectrometry for their identification.
- Protein fragment complementation assay
-
A method used to measure protein–protein physical interactions. Protein interactions are coupled to the refolding of β-lactamase, which is fragmented and each of the two fragments is fused to the two proteins of interest. The reconstitution of β-lactamase activity acts as an interaction detector.
- Power law
-
A statistical model that describes that one quantity is proportional to the power of another quantity.
- Triosephosphate isomerase (TIM)-barrel fold
-
The most frequent and conserved protein fold, comprising eight α-helices and eight β-sheets.
- Orthologue
-
A gene present in different species that evolved from a common ancestral gene by speciation.
Rights and permissions
About this article
Cite this article
Yamada, T., Bork, P. Evolution of biomolecular networks — lessons from metabolic and protein interactions. Nat Rev Mol Cell Biol 10, 791–803 (2009). https://doi.org/10.1038/nrm2787
Issue Date:
DOI: https://doi.org/10.1038/nrm2787
This article is cited by
-
Degeneracy measures in biologically plausible random Boolean networks
BMC Bioinformatics (2022)
-
1H NMR metabolomics analysis of oil palm stem tissue infected by Ganoderma boninense based on field severity Indices
Scientific Reports (2022)
-
System-Wide Pollution of Biomedical Data: Consequence of the Search for Hub Genes of Hepatocellular Carcinoma Without Spatiotemporal Consideration
Molecular Diagnosis & Therapy (2021)
-
The Coevolution of Cellularity and Metabolism Following the Origin of Life
Journal of Molecular Evolution (2020)
-
NetControl4BioMed: a pipeline for biomedical data acquisition and analysis of network controllability
BMC Bioinformatics (2018)