Key Points
-
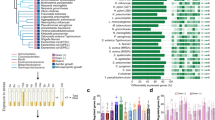
We have re-analysed and compared microarray data from 32 published studies and uncovered a common host-transcriptional-response to pathogens.
-
Sub-clusters of functionally-related genes are preferentially induced by different host cell types.
-
Toll-like receptors (TLRs) are responsible for both common and pathogen-specific programmes of gene expression.
-
Some pathogens actively repress portions of the host transcriptional response and such activities can be correlated with pathogen virulence and disease severity.
-
The use of material from the clinic and from animal models provides information about the dynamics of infection and its resolution in vivo.
-
Gene expression profiling of pathogen-associated cancers provides clues about the mechanisms of tumorigenesis.
Abstract
DNA microarrays have allowed us to monitor the effects of pathogens on host-cell gene expression programmes in great depth and on a broad scale. The comparison of results that have been generated by these studies is complex, and such a comparison has not previously been attempted in a systematic manner. In this review, we have collated and compared published transcriptional-profiling data from 32 studies that involved 77 different host–pathogen interactions, and have defined a common host-transcriptional-response. We outline gene expression patterns in the context of Toll-like receptor and pathogen-mediated signalling pathways, and summarize the contributions that transcriptional-profiling studies have made to our understanding of the infectious disease process.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Der, S. D., Zhou, A., Williams, B. R. & Silverman, R. H. Identification of genes differentially regulated by interferon α, β, or γ using oligonucleotide arrays. Proc. Natl Acad. Sci. USA 95, 15623–15628 (1998).
Zhu, H., Cong, J. P., Mamtora, G., Gingeras, T. & Shenk, T. Cellular gene expression altered by human cytomegalovirus: global monitoring with oligonucleotide arrays. Proc. Natl Acad. Sci. USA 95, 14470–14475 (1998). This was the first study to use microarrays to monitor the host response to infection.
Boldrick, J. C. et al. Stereotyped and specific gene expression programs in human innate immune responses to bacteria. Proc. Natl Acad. Sci. USA 99, 972–977 (2002).
Nau, G. J. et al. Human macrophage activation programs induced by bacterial pathogens. Proc. Natl Acad. Sci. USA 99, 1503–1508 (2002).
Kobayashi, S. D. et al. Bacterial pathogens modulate an apoptosis differentiation program in human neutrophils. Proc. Natl Acad. Sci. USA 100, 10948–10953 (2003).
Huang, Q. et al. The plasticity of dendritic cell responses to pathogens and their components. Science 294, 870–875 (2001). References 3, 4 and 6 demonstrate that human immune cells respond to pathogens with both common and pathogen-specific responses and that individual pathogen components can recapitulate these gene expression changes.
Izmailova, E. et al. HIV-1 Tat reprograms immature dendritic cells to express chemoattractants for activated T cells and macrophages. Nature Med. 9, 191–197 (2003). This paper revealed how an HIV protein selectively induces only part of the normal DC response to pathogens, which results in the recruitment of T cells in the absence of DC activation.
Hertzog, P. J., O'Neill, L. A. & Hamilton, J. A. The interferon in TLR signaling: more than just antiviral. Trends Immunol. 24, 534–539 (2003).
Hooper, L. V. et al. Molecular analysis of commensal host–microbial relationships in the intestine. Science 291, 881–884 (2001).
Chaussabel, D. et al. Unique gene expression profiles of human macrophages and dendritic cells to phylogenetically distinct parasites. Blood 102, 672–681 (2003).
Beutler, B. Inferences, questions and possibilities in Toll-like receptor signalling. Nature 430, 257–263 (2004).
Doyle, S. et al. IRF3 mediates a TLR3/TLR4-specific antiviral gene program. Immunity 17, 251–263 (2002).
Taniguchi, T. & Takaoka, A. The interferon-α/β system in antiviral responses: a multimodal machinery of gene regulation by the IRF family of transcription factors. Curr. Opin. Immunol. 14, 111–116 (2002).
Geiss, G. et al. A comprehensive view of regulation of gene expression by double-stranded RNA-mediated cell signaling. J. Biol. Chem. 276, 30178–30182 (2001).
Rosenberger, C. M., Scott, M. G., Gold, M. R., Hancock, R. E. & Finlay, B. B. Salmonella typhimurium infection and lipopolysaccharide stimulation induce similar changes in macrophage gene expression. J. Immunol. 164, 5894–5904 (2000).
Poltorak, A. et al. Defective LPS signaling in C3H/HeJ and C57BL/10ScCr mice: mutations in Tlr4 gene. Science 282, 2085–2088 (1998).
Palliser, D. et al. A role for toll-like receptor 4 in dendritic cell activation and cytolytic CD8+ T cell differentiation in response to a recombinant heat shock fusion protein. J. Immunol. 172, 2885–2893 (2004).
Weighardt, H. et al. Identification of a TLR4- and TRIF-dependent activation program of dendritic cells. Eur. J. Immunol. 34, 558–564 (2004).
Rogers, P. D. et al. Pneumolysin-dependent and-independent gene expression identified by cDNA microarray analysis of THP-1 human mononuclear cells stimulated by Streptococcus pneumoniae. Infect. Immun. 71, 2087–2094 (2003).
Malley, R. et al. Recognition of pneumolysin by Toll-like receptor 4 confers resistance to pneumococcal infection. Proc. Natl Acad. Sci. USA 100, 1966–1971 (2003).
Kurt-Jones, E. A. et al. Pattern recognition receptors TLR4 and CD14 mediate response to respiratory syncytial virus. Nature Immunol. 1, 398–401 (2000).
Hayashi, F. et al. The innate immune response to bacterial flagellin is mediated by Toll-like receptor 5. Nature 410, 1099–1103 (2001).
Zeng, H. et al. Flagellin is the major proinflammatory determinant of enteropathogenic Salmonella. J. Immunol. 171, 3668–3674 (2003).
Nau, G. J., Schlesinger, A., Richmond, J. F. & Young, R. A. Cumulative Toll-like receptor activation in human macrophages treated with whole bacteria. J. Immunol. 170, 5203–5209 (2003).
Toshchakov, V. et al. TLR4, but not TLR2, mediates IFN-β-induced STAT1α/β-dependent gene expression in macrophages. Nature Immunol. 3, 392–398 (2002).
Lund, J. M. et al. Recognition of single-stranded RNA viruses by Toll-like receptor 7. Proc. Natl Acad. Sci. USA 101, 5598–5603 (2004).
Underhill, D. M. & Ozinsky, A. Toll-like receptors: key mediators of microbe detection. Curr. Opin. Immunol. 14, 103–110 (2002).
Simmen, K. A. et al. Global modulation of cellular transcription by human cytomegalovirus is initiated by viral glycoprotein B. Proc. Natl Acad. Sci. USA 98, 7140–7145 (2001). This paper identified the specific virus protein that is responsible for initiating the host transcriptional response against a virus and defined this response as being mediated by IFN.
Boehme, K. W., Singh, J., Perry, S. T. & Compton, T. Human cytomegalovirus elicits a coordinated cellular antiviral response via envelope glycoprotein B. J. Virol. 78, 1202–1211 (2004).
Compton, T. et al. Human cytomegalovirus activates inflammatory cytokine responses via CD14 and Toll-like receptor 2. J. Virol. 77, 4588–4596 (2003).
Wang, X., Huong, S. M., Chiu, M. L., Raab-Traub, N. & Huang, E. S. Epidermal growth factor receptor is a cellular receptor for human cytomegalovirus. Nature 424, 456–461 (2003).
Mossman, K. L. et al. Herpes simplex virus triggers and then disarms a host antiviral response. J. Virol. 75, 750–758 (2001).
Browne, E. P., Wing, B., Coleman, D. & Shenk, T. Altered cellular mRNA levels in human cytomegalovirus-infected fibroblasts: viral block to the accumulation of antiviral mRNAs. J. Virol. 75, 12319–12330 (2001).
Browne, E. P. & Shenk, T. Human cytomegalovirus UL83-coded pp65 virion protein inhibits antiviral gene expression in infected cells. Proc. Natl Acad. Sci. USA 100, 11439–11444 (2003).
Chang, Y. E. & Laimins, L. A. Microarray analysis identifies interferon-inducible genes and Stat-1 as major transcriptional targets of human papillomavirus type 31. J. Virol. 74, 4174–4182 (2000).
Nees, M. et al. Papillomavirus type 16 oncogenes downregulate expression of interferon-responsive genes and upregulate proliferation-associated and NF-κB-responsive genes in cervical keratinocytes. J. Virol. 75, 4283–4296 (2001). References 34 and 36 show the potential of using microarrays to identify pathogen proteins that inhibit the host immune response.
Simmons, A., Aluvihare, V. & McMichael, A. Nef triggers a transcriptional program in T cells imitating single-signal T cell activation and inducing HIV virulence mediators. Immunity 14, 763–777 (2001). The authors used microarrays to generate an expression signature of T-cell activation and to show that this is mimicked by HIV Nef.
Corbeil, J. et al. Temporal gene regulation during HIV-1 infection of human CD4+ T cells. Genome Res. 11, 1198–11204 (2001).
van 't Wout, A. B. et al. Cellular gene expression upon human immunodeficiency virus type 1 infection of CD4+-T-cell lines. J. Virol. 77, 1392–1402 (2003).
Ehrt, S. et al. Reprogramming of the macrophage transcriptome in response to interferon-γ and Mycobacterium tuberculosis: signaling roles of nitric oxide synthase-2 and phagocyte oxidase. J. Exp. Med. 194, 1123–1140 (2001).
Sauvonnet, N., Pradet-Balade, B., Garcia-Sanz, J. A. & Cornelis, G. R. Regulation of mRNA expression in macrophages after Yersinia enterocolitica infection. Role of different Yop effectors. J. Biol. Chem. 277, 25133–25142 (2002).
Bohn, E. et al. Gene expression patterns of epithelial cells modulated by pathogenicity factors of Yersinia enterocolitica. Cell. Microbiol. 6, 129–141 (2004).
Geiss, G. K. et al. Cellular transcriptional profiling in influenza A virus-infected lung epithelial cells: the role of the nonstructural NS1 protein in the evasion of the host innate defense and its potential contribution to pandemic influenza. Proc. Natl Acad. Sci. USA 99, 10736–10741 (2002).
Girard, S. et al. An altered cellular response to interferon and up-regulation of interleukin-8 induced by the hepatitis C viral protein NS5A uncovered by microarray analysis. Virology 295, 272–283 (2002).
Geiss, G. K. et al. Gene expression profiling of the cellular transcriptional network regulated by α/β interferon and its partial attenuation by the hepatitis C virus nonstructural 5A protein. J. Virol. 77, 6367–6375 (2003).
Polyak, S. J., Khabar, K. S., Rezeiq, M. & Gretch, D. R. Elevated levels of interleukin-8 in serum are associated with hepatitis C virus infection and resistance to interferon therapy. J. Virol. 75, 6209–6211 (2001).
Geimonen, E. et al. Pathogenic and nonpathogenic hantaviruses differentially regulate endothelial cell responses. Proc. Natl Acad. Sci. USA 99, 13837–13842 (2002).
Guerra, S. et al. Cellular gene expression survey of vaccinia virus infection of human HeLa cells. J. Virol. 77, 6493–6506 (2003).
Guerra, S. et al. Microarray analysis reveals characteristic changes of host cell gene expression in response to attenuated modified vaccinia virus Ankara infection of human HeLa cells. J. Virol. 78, 5820–5834 (2004).
Ichikawa, J. K. et al. Interaction of Pseudomonas aeruginosa with epithelial cells: identification of differentially regulated genes by expression microarray analysis of human cDNAs. Proc. Natl Acad. Sci. USA 97, 9659–9664 (2000).
Detweiler, C. S., Cunanan, D. B. & Falkow, S. Host microarray analysis reveals a role for the Salmonella response regulator phoP in human macrophage cell death. Proc. Natl Acad. Sci. USA 98, 5850–5855 (2001).
Pathan, N. et al. Role of interleukin 6 in myocardial dysfunction of meningococcal septic shock. Lancet 363, 203–209 (2004). This paper coupled microarray gene expression data with other functional annotation to identify an unknown factor that was responsible for impaired heart function during sepsis.
Taylor, L. A. et al. Host gene regulation during coxsackievirus B3 infection in mice: assessment by microarrays. Circ. Res. 87, 328–334 (2000).
Sui, Y. et al. Microarray analysis of cytokine and chemokine genes in the brains of macaques with SHIV-encephalitis. J. Med. Primatol. 32, 229–239 (2003).
Roberts, E. S. et al. Induction of pathogenic sets of genes in macrophages and neurons in NeuroAIDS. Am. J. Pathol. 162, 2041–2057 (2003).
Domachowske, J. B., Bonville, C. A., Easton, A. J. & Rosenberg, H. F. Differential expression of proinflammatory cytokine genes in vivo in response to pathogenic and nonpathogenic pneumovirus infections. J. Infect. Dis. 186, 8–14 (2002).
Johnston, C., Jiang, W., Chu, T. & Levine, B. Identification of genes involved in the host response to neurovirulent alphavirus infection. J. Virol. 75, 10431–10445 (2001).
Bigger, C. B., Brasky, K. M. & Lanford, R. E. DNA microarray analysis of chimpanzee liver during acute resolving hepatitis C virus infection. J. Virol. 75, 7059–7066 (2001). This study followed the gene expression changes that accompany the course of infection by a human pathogen.
George, M. D., Sankaran, S., Reay, E., Gelli, A. C. & Dandekar, S. High-throughput gene expression profiling indicates dysregulation of intestinal cell cycle mediators and growth factors during primary simian immunodeficiency virus infection. Virology 312, 84–94 (2003).
Guadalupe, M. et al. Severe CD4+ T-cell depletion in gut lymphoid tissue during primary human immunodeficiency virus type 1 infection and substantial delay in restoration following highly active antiretroviral therapy. J. Virol. 77, 11708–11717 (2003).
Chun, T. W. et al. Gene expression and viral prodution in latently infected, resting CD4+ T cells in viremic versus aviremic HIV-infected individuals. Proc. Natl Acad. Sci. USA 100, 1908–1913 (2003).
Mueller, A. et al. Protective immunity against Helicobacter is characterized by a unique transcriptional signature. Proc. Natl Acad. Sci. USA 100, 12289–12294 (2003).
Wu, C. G. et al. Distinctive gene expression profiles associated with Hepatitis B virus x protein. Oncogene 20, 3674–3682 (2001).
Xu, X. R. et al. Insight into hepatocellular carcinogenesis at transcriptome level by comparing gene expression profiles of hepatocellular carcinoma with those of corresponding noncancerous liver. Proc. Natl Acad. Sci. USA 98, 15089–15094 (2001).
Okabe, H. et al. Genome-wide analysis of gene expression in human hepatocellular carcinomas using cDNA microarray: identification of genes involved in viral carcinogenesis and tumor progression. Cancer Res. 61, 2129–2137 (2001).
Iizuka, N. et al. Comparison of gene expression profiles between hepatitis B virus- and hepatitis C virus-infected hepatocellular carcinoma by oligonucleotide microarray data on the basis of a supervised learning method. Cancer Res. 62, 3939–3944 (2002).
Wang, H. W. et al. Kaposi sarcoma herpesvirus-induced cellular reprogramming contributes to the lymphatic endothelial gene expression in Kaposi sarcoma. Nature Genet. 36, 687–693 (2004).
Hong, Y. K. et al. Lymphatic reprogramming of blood vascular endothelium by Kaposi sarcoma-associated herpesvirus. Nature Genet. 36, 683–685 (2004). Reference 67 shows that the unusual gene expression pattern of Kaposi's sarcoma is caused by the reprogramming of endothelial cells by KSHV. Reference 68 suggests that this virus activity might reside in LANA1.
Jenner, R. G. et al. Kaposi's sarcoma-associated herpesvirus-infected primary effusion lymphoma has a plasma cell gene expression profile. Proc. Natl Acad. Sci. USA 100, 10399–10404 (2003).
Klein, U. et al. Gene expression profile analysis of AIDS-related primary effusion lymphoma (PEL) suggests a plasmablastic derivation and identifies PEL-specific transcripts. Blood 101, 4115–4121 (2003).
Moses, A. V. et al. Kaposi''s sarcoma-associated herpesvirus-induced upregulation of the c-kit proto-oncogene, as identified by gene expression profiling, is essential for the transformation of endothelial cells. J. Virol. 76, 8383–8399 (2002). This paper represents the first study to identify a novel drug target for an infectious disease using microarrays, and to test the efficacy of a pharmacological agent against the protein in a disease model.
Casadevall, A. & Pirofski, L. A. The damage-response framework of microbial pathogenesis. Nature Rev. Microbiol. 1, 17–24 (2003).
Schnappinger, D. et al. Transcriptional adaptation of Mycobacterium tuberculosis within macrophages: insights into the phagosomal environment. J. Exp. Med. 198, 693–704 (2003).
Ren, B. et al. Genome-wide location and function of DNA binding proteins. Science 290, 2306–2309 (2000).
Odom, D. T. et al. Control of pancreas and liver gene expression by HNF transcription factors. Science 303, 1378–1381 (2004).
Rodriguez, N. E., Chang, H. K. & Wilson, M. E. Novel program of macrophage gene expression induced by phagocytosis of Leishmania chagasi. Infect. Immun. 72, 2111–2122 (2004).
Dietz, A. B., Bulur, P. A., Knutson, G. J., Matasic, R. & Vuk-Pavlovic, S. Maturation of human monocyte-derived dendritic cells studied by microarray hybridization. Biochem. Biophys. Res. Commun. 275, 731–738 (2000).
Feezor, R. J. et al. Molecular characterization of the acute inflammatory response to infections with gram-negative versus gram-positive bacteria. Infect. Immun. 71, 5803–5813 (2003).
Jones, J. O. & Arvin, A. M. Microarray analysis of host cell gene transcription in response to varicella-zoster virus infection of human T cells and fibroblasts in vitro and SCIDhu skin xenografts in vivo. J. Virol. 77, 1268–1280 (2003).
Iizuka, N. et al. Oligonucleotide microarray for prediction of early intrahepatic recurrence of hepatocellular carcinoma after curative resection. Lancet 361, 923–929 (2003).
Kramer, M. F. et al. Latent herpes simplex virus infection of sensory neurons alters neuronal gene expression. J. Virol. 77, 9533–9541 (2003).
Radhakrishnan, S. et al. JC virus-induced changes in cellular gene expression in primary human astrocytes. J. Virol. 77, 10638–10644 (2003).
Warke, R. V. et al. Dengue virus induces novel changes in gene expression of human umbilical vein endothelial cells. J. Virol. 77, 11822–11832 (2003).
Belcher, C. E. et al. The transcriptional responses of respiratory epithelial cells to Bordetella pertussis reveal host defensive and pathogen counter-defensive strategies. Proc. Natl Acad. Sci. USA 97, 13847–13852 (2000).
Zhang, Y. et al. Expression of respiratory syncytial virus-induced chemokine gene networks in lower airway epithelial cells revealed by cDNA microarrays. J. Virol. 75, 9044–9058 (2001).
Tian, B. et al. Identification of NF-κB-dependent gene networks in respiratory syncytial virus-infected cells. J. Virol. 76, 6800–6814 (2002).
Zhang, Y. et al. Ribavirin treatment up-regulates antiviral gene expression via the interferon-stimulated response element in respiratory syncytial virus-infected epithelial cells. J. Virol. 77, 5933–5947 (2003).
Cuadras, M. A., Feigelstock, D. A., An, S. & Greenberg, H. B. Gene expression pattern in Caco-2 cells following rotavirus infection. J. Virol. 76, 4467–4482 (2002).
Wells, S. I. et al. Transcriptome signature of irreversible senescence in human papillomavirus-positive cervical cancer cells. Proc. Natl Acad. Sci. USA 100, 7093–7098 (2003).
Guillemin, K., Salama, N. R., Tompkins, L. S. & Falkow, S. Cag pathogenicity island-specific responses of gastric epithelial cells to Helicobacter pylori infection. Proc. Natl Acad. Sci. USA 99, 15136–15141 (2002).
Mueller, A. et al. Distinct gene expression profiles characterize the histopathological stages of disease in Helicobacter–induced mucosa–associated lymphoid tissue lymphoma. Proc. Natl Acad. Sci. USA 100, 1292–1297 (2003).
Acknowledgements
The authors thank E. Herbolsheimer and G. Bell for help with data processing and Gene Ontology annotation. We apologize to any authors whose papers we inadvertently missed when compiling Supplementary information S1,S2,S3 (tables and references). This work was funded by the National Institute of Allergy and Infectious Diseases.
Author information
Authors and Affiliations
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Related links
Related links
DATABASES
Swiss-Prot
FURTHER INFORMATION
Glossary
- DNA MICROARRAY
-
An ordered array of DNA probes of known sequence on a solid support, fabricated on a very small scale. The probes are used to interrogate the composition of DNA mixtures through hybridization.
- INTERFERON
-
(IFN). Cytokines that can induce cells to resist infection. Type 1 IFNs (α and β) are produced by several cell types; type 2 IFN(γ) is produced by T cells and NK cells.
- TRANSCRIPTOME
-
The set of mRNA transcripts that are present in any one cell at a given time.
- CLUSTER ANALYSIS
-
A multivariate statistical classification technique that partitions variables (which for microarray data are genes or samples) into groups (clusters) based on shared characteristics (in this case, gene expression patterns).
- LATENT INFECTION (LATENCY)
-
The stage of some virus life-cycles in which no progeny virions are produced. Viral gene expression is restricted.
- APOPTOSIS
-
Programmed cell death, a controlled physiological process by which a cell terminates its own existence.
- ADAPTOR PROTEINS
-
Linker proteins that connect cell-surface receptors to downstream members of intracellular signalling pathways.
- T-HELPER (TH) CELLS
-
T cells that help in the activation of other cell types; TH1 cells are mainly involved in macrophage activation, TH2 cells in B-cell activation.
- FORWARD GENETICS
-
The experimental process of identifying the genes responsible for a defined function or phenotype.
- REVERSE GENETICS
-
The experimental process of assigning functions to a gene, usually through mutation or inhibition of its expression.
- GRAM-POSITIVE BACTERIA
-
A group of bacteria that retain the stain of Gram's solution after rinsing. They have a single cell wall that does not contain LPS. Gram-negative bacteria have a double cell wall that contains LPS, which prevents the retention of the stain.
- DOUBLE-STRANDED (ds)RNA
-
Double-stranded RNA, which is produced during the life-cycle of many viruses but which is otherwise not normally found in the cell (except for very short RNA species).
- GLIOMA CELL LINE
-
A self-propagating cell population that is derived from a cancer of glial cells.
- ELISA
-
Enzyme-linked immunosorbent assay — an assay in which antigen is detected by an antibody linked to an enzyme that produces a coloured product.
- VIRULENCE FACTOR
-
A pathogen protein that is essential for causing disease in a host.
- RNA INTERFERENCE
-
(RNAI) Post-transcriptional gene silencing (PTGS) induced by the direct introduction of dsRNA.
Rights and permissions
About this article
Cite this article
Jenner, R., Young, R. Insights into host responses against pathogens from transcriptional profiling. Nat Rev Microbiol 3, 281–294 (2005). https://doi.org/10.1038/nrmicro1126
Issue Date:
DOI: https://doi.org/10.1038/nrmicro1126
This article is cited by
-
The Mycobacterium tuberculosis methyltransferase Rv2067c manipulates host epigenetic programming to promote its own survival
Nature Communications (2023)
-
Network meta-analysis of transcriptome expression changes in different manifestations of dengue virus infection
BMC Genomics (2022)
-
Dynamic Host Immune and Transcriptomic Responses to Respiratory Syncytial Virus Infection in a Vaccination-Challenge Mouse Model
Virologica Sinica (2021)
-
Mycoplasma gallisepticum triggers immune damage in the chicken thymus by activating the TLR-2/MyD88/NF-κB signaling pathway and NLRP3 inflammasome
Veterinary Research (2020)
-
The DOT1L inhibitor Pinometostat decreases the host-response against infections: Considerations about its use in human therapy
Scientific Reports (2019)