Key Points
-
The Gram-negative outer membrane protein (OMP) family includes proteins that are associated with basic physiological functions, virulence and multidrug resistance, and therefore plays a fundamental part in the maintenance of cellular viability.
-
Understanding how these proteins are targeted and folded into this membrane is crucial, as it could offer important medical benefits. Compounds that inhibit key stages of this process would block key stages of OMP biogenesis, thereby inhibiting essential physiological, pathogenic and drug resistance functions, and could prove useful in combating diverse pathogens, including Pseudomonas aeruginosa, Neisseria meningitidis and Salmonella enterica.
-
OMP biogenesis in Gram-negative bacteria has, until recently, remained a largely unknown mechanism. However, over the past 3 years, a complex of proteins has been discovered that is known as the β-barrel assembly machinery (BAM) and is responsible for folding and inserting OMPs into the membrane.
-
Recent advances in our understanding of the molecular basis of OMP biogenesis in Gram-negative bacteria are discussed.
-
Emphasis is placed on analysis of the recently discovered component structures and accessory interactions, in particular with the periplasmic chaperones DegP, Skp and SurA, which are known to interact with OMPs.
-
The mechanisms that the BAM complex might use in the folding and insertion of OMPs into the membrane are also discussed.
Abstract
The folding of transmembrane proteins into the outer membrane presents formidable challenges to Gram-negative bacteria. These proteins must migrate from the cytoplasm, through the inner membrane and into the periplasm, before being recognized by the β-barrel assembly machinery, which mediates efficient insertion of folded β-barrels into the outer membrane. Recent discoveries of component structures and accessory interactions of this complex are yielding insights into how cells fold membrane proteins. Here, we discuss how these structures illuminate the mechanisms responsible for the biogenesis of outer membrane proteins.
Similar content being viewed by others
Main
Gram-negative bacteria are surrounded by two lipid bilayers: the inner membrane and the outer membrane. These membranes have important differences in their makeup. The inner membrane is composed of a symmetrical phospholipid bilayer and harbours predominantly α-helical proteins, whereas the outer membrane is asymmetrical and contains primarily β-barrel fold proteins. In the outer membrane, phospholipid is located in the inner leaflet and lipopolysaccharide (LPS) is located in the outer leaflet. The outer membrane protein (OMP) family includes proteins associated with basic physiological functions, virulence and multidrug resistance and therefore plays a fundamental part in the maintenance of cellular viability1. The understanding of how these proteins are targeted and folded into this membrane is therefore crucial, as it could offer medical benefits. Compounds that inhibit stages of this process would block key stages of OMP biogenesis, thereby inhibiting essential physiological, pathogenic and drug resistance functions, and could prove useful in combating diverse pathogens, including Pseudomonas aeruginosa , Neisseria meningitidis and Salmonella enterica .
The past 5 years have heralded the discovery of a network of proteins responsible for folding and inserting OMPs into the outer membrane. The core complex is now known as the β-barrel assembly machinery (BAM). The first structures of components of this complex have recently been solved and are shedding light on how β-barrels are built in vivo. This Review discusses the implications of these structures and interactions of BAM components, and focuses on the mechanisms responsible for trafficking and folding proteins into the outer membrane of Gram-negative bacteria.
The pathway of OMP biogenesis in Escherichia coli
All proteins in E. coli are synthesized in the cytoplasm. Those destined for the outer membrane must pass through both the inner membrane and the periplasmic space before even reaching the outer membrane, where folding and insertion takes place (Fig. 1). This process has multiple stages. The nascent OMP precursors are synthesized with an amino-terminal leader sequence and interact with cytoplasmic chaperones and the SecYEG complex to mediate translocation across the inner membrane in a process that is dependent on ATP and the proton motive force2,3,4,5,6,7. On entering the periplasm, the leader sequence is processed by a signal peptidase, and the nascent OMP associates with periplasmic chaperones, including SurA, Skp and DegP. These chaperones are thought to form two pathways, the SurA pathway and the Skp–DegP pathway, that transport nascent OMPs across the periplasmic space to the outer membrane8,9,10. Unlike the actively driven Sec translocon, the outer membrane uses the passive BAM complex. Almost all known OMPs require the BAM complex for folding11,12,13,14. One exception could be the secretin PulD, which has been shown to insert into the membrane in the absence of the BAM complex15,16. Interestingly, a subset of OMPs also require lipid synthesis for correct assembly, suggesting that the BAM complex mediates multiple folding pathways17. In E. coli this complex is composed of five proteins: YaeT, which is an integral membrane protein, and four accessory lipoproteins, YfgL, NlpB, YfiO and SmpA, which localize to the inner leaflet of the outer membrane9,12,18,19,20. These proteins have recently been renamed, although discrepancies exist in the literature concerning the correct nomenclature19,20,21,22. We will adhere to the early size-ordered designations: BamA (YaeT), BamB (YfgL), BamC (NlpB), BamD (YfiO) and BamE (SmpA).
Outer membrane proteins (OMPs) destined for the β-barrel assembly machinery (BAM) complex are first targeted to the SecYEG translocon. Following export through SecYEG, the nascent OMPs are recruited by two proposed chaperone pathways, the SurA and the Skp–DegP pathway, and are transported through the periplasm to the outer membrane. Excess levels of unfolded OMPs in the periplasm are targeted for breakdown by proteases, such as DegP, through an envelope stress response. Folding and insertion of nascent OMPs is thought to occur through the BAM complex (BamA–BamE)8. It is currently unclear how the BAM complex functions in OMP folding and insertion. However, a number of possible mechanisms exist. According to the pore-folding model, the β-barrel of BamA offers its pore for insertion of the nascent OMP into the membrane, and the POTRA (polypeptide transport-associated) domains and/or accessory components act to thread the OMP into the pore (1). In the complex pore-folding model, the central core is formed by a multimeric BAM complex that acts as the point of insertion into the membrane (2). Release of the OMP could then occur by dissociation of the multimeric BAM complexes. The barrel-folding model suggests that the β-barrel of BamA provides a template for barrel folding in the vicinity of the BAM complex (3). According to the chaperone-folding model, the periplasmic chaperones, and in particular DegP, act to fold the protein and protect it from degradation during passage through the periplasm (4). The BAM complex thus functions only to insert the protein into the membrane. Finally, in the accessory folding model the BAM complex functions to fold the nascent OMP but does not have a function in membrane insertion (5). The folded OMP is then released to DegP in a quality-control mechanism to remove incorrectly folded OMPs. The protein is then inserted into the membrane either by DegP or by some as-yet-unknown mechanism that could involve the BAM complex.
BamA, an essential Gram-negative protein
Voulhoux et al. showed that outer membrane protein 85 (omp85) is an essential gene in N. meningitidis and that depletion of its product BamA (also known as Omp85) resulted in the accumulation of unfolded OMP aggregates in the periplasm23. Initially, BamA was thought to be involved in LPS and phospholipid incorporation, rather than OMP assembly, as it is encoded in an operon that contains LPS biosynthetic genes, and LPS and phospholipid accumulate in the inner membrane when it is depleted24. However, from E. coli studies it is apparent that BamA plays a central part in OMP assembly11,12,13,23 and that its effects on LPS insertion are indirect owing to misfolding of LPS biosynthetic proteins, such as LptD (previously known as Imp)25.
Consistent with its essential role, BamA is found in all Gram-negative bacteria and contains two major components: a set of five POTRA (polypeptide transport-associated) domains oriented towards the periplasm and a carboxy-terminal β-barrel inserted into the outer membrane26. BamA bears striking sequence and structural similarity to the protein-translocating TpsB proteins of the Gram-negative bacterial two-partner secretion system and to homologues in the outer membranes of plastids and mitochondria, reflecting the bacterial origins of these latter organelles27,28 (Boxes 1, 2).
The structures of POTRA domains from E. coli BamA have recently been solved by NMR, small angle X-ray scattering (SAXS) and X-ray crystallography, revealing their novel folds and interactions29,30 (Fig. 2). Kim et al. reported the first crystal structure of POTRA domains 1–4 (POTRA1–4)29. Although these domains had only marginally similar sequences (<13% identity), they adopted a common fold that comprised a three-stranded β-sheet overlaid by a pair of antiparallel α-helices, albeit with distinct interdomain angles and interfaces. The dimeric form of the POTRA domains observed in the crystal was stabilized by β-strand pairing (β-augmentation) between an 'orphan' β-strand of the C-terminally truncated POTRA5 and the β-sheet of POTRA3. NMR and SAXS studies of POTRA1–2 and POTRA1–5 in solution revealed that they have similar domain structures but exist exclusively in monomeric states and possess different domain–domain interfaces and angles30. Whereas the crystal structure detected by Kim et al. exhibited highly ordered interdomain contacts29, NMR revealed flexible linkers and distinct interfaces between domains. Only the orientation revealed by NMR is consistent with the molecular envelope of POTRA1–5 that was identified using SAXS. This therefore suggests that the crystallized orientation of POTRA1–4 detected by Kim et al. does not occur in solution but results from non-physiological contacts between truncated BamA constructs in a crystalline environment30 that would not occur when the fifth POTRA domain is intact. Indeed this seems to be the case, as a more recent crystal structure of BamA POTRA1–4 solved by Gatzeva-Topalova and colleagues revealed that it is monomeric and exists in an extended conformation that is consistent with the SAXS conformer31. However, the observation that different interdomain orientations can be adopted suggests that POTRA domain linkers have a substantial amount of conformational freedom, and could reflect interdomain articulations that may have functional implications during the folding pathway.
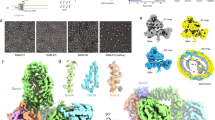
A | Ribbon models of the POTRA (polypeptide transport-associated) domains of BamA and its structurally related proteins. Aa | The Escherichia coli BamA POTRA domain fold, showing its characteristic three-stranded β-sheet (blue) overlaid by a pair of antiparallel α-helices (red). The structure is of POTRA 2 from E. coli BamA29 (protein data bank (PDB) code 2qcz). Ab | Solution structure of the BamE homologue OmlA from Xanthomonas axonopodis pathovar citri33 (PDB code 2pxg). Ac | Crystal structure of cytoplasmic dynein light chain from Drosophila melanogaster35 (PDB code 2pg1) (Z score = 3.8 and the root mean square deviation (RMSD) of Cα = 2.7 Å; the RMSD values were calculated against BamA POTRA1 using the program Dali). Ad | Crystal structure of a β-lactamase inhibitory protein domain from Streptomyces clavuligeris34 (PDB code 2g2u). Ae | Crystal structure of FtsQ residues 58–126 from E. coli36 (PDB code 2vh1) (Z score = 8.2; RMSD of Cα = 1.8 Å). B | Crystal structure of the E. coli BamA POTRA1–4 (Ref. 29) (PDB code 2qcz). For monomer 1, POTRA1 is coloured blue, POTRA2 is coloured red, POTRA3 is coloured green and POTRA4 is coloured yellow. For the carboxyl termini, POTRA5 truncation is coloured cyan and monomer 2 is coloured grey. C | NMR solution structure ensemble30 of E. coli BamA POTRA domains 1 and 2 (PDB code 2v9h). The ensemble of the 20 lowest energy structures is superimposed on the POTRA2 domain. D | A model of the POTRA1–5 construct from small angle X-ray scattering30. E | Crystal structure of FhaC32 (PDB code 2qdz) (Dali alignment of FhaC periplasmic domains to BamA POTRA1: Z score = 5.6 and 7.6 and RMSD of Cα = 2.5 and 1.4 Å, respectively). β-barrel, yellow; POTRA1, blue; POTRA2, red; amino-terminal α-helix, green.
Other proteins with similar folds to the POTRA domain have recently been identified, and could therefore have similar functional attributes. These include the filamentous haemagglutinin transporter protein FhaC from Bordetella pertussis , the cell division protein FtsQ from E. coli and dynein light chain from Drosophila melanogaster 32,33,34,35,36. Also similar to the POTRA domains are OmlA, a BamE homologue from Xanthomonas axonopodis pathovar citri, and beta-lactamase inhibitory protein (BLIP) from Streptomyces clavuligeris (Fig. 2). Although differences are apparent in the order and number of secondary structure elements, each domain adopts a similar topology with exposed β-strands. All domains have been shown, or are predicted, to bind to other protein ligands, suggesting they have common functionality. Studies of the BamA POTRA domain, BLIP and dynein light chain indicated that β-augmentation is probably the common mode of interaction. In the case of BLIP, β-augmentation occurs between the two 76-residue domains34, whereas for dynein light chain β-augmentation is associated with a dynein intermediate chain peptide35.
The structure of the BamA β-barrel has not yet been reported. However, the structure of the distantly related TpsB transporter FhaC has been determined32 (Fig. 2e). FhaC mediates translocation of the B. pertussis major adhesin filamentous haemagglutinin to the bacterial surface32. The FhaC structure contains a 16-stranded β-barrel with an N-terminal periplasmic extension composed of two POTRA domains and an α-helix that is embedded in the barrel pore. In addition to occlusion by the α-helix, the channel is further blocked by a large extracellular loop close to its C terminus that forms a hairpin in the barrel interior. Although BamA is predicted to possess a similar extracellular loop, no segment that corresponds to the N-terminal α-helix is present. Nonetheless, conductance studies indicate that the channel is closed, hinting that other elements may block its pore37.
Exactly how BamA functions in OMP assembly remains unclear. However, it is beginning to emerge that its POTRA domains might have a role in binding unfolded OMPs26,37. Robert et al. suggest that the BAM recognizes a specific recognition motif encoded in the C-terminal β-strand of OMPs37. Indeed, comparisons of the C-terminal β-strands of OMPs from different Gram-negative bacteria reveal a conserved amphipathic structure with hydrophobic residues at positions 1 (phenylalanine or tryptophan), 3 (preferentially tyrosine), 5, 7 and 9 from the C terminus; the terminal aromatic residue of the C terminus is necessary for efficient outer membrane insertion38. This targeting motif appears to differ between E. coli and N. meningitidis (the E. coli motif contains predominantly polar residues, whereas the N. meningitidis motif possesses residues at positions 2 and 4), suggesting that OMP sorting is species specific37. This could explain why E. coli BamA did not recognize the C-terminal motif of Neisseria PorA and why overexpression of Neisseria OMPs in E. coli is toxic37. Interestingly, the C-terminal motif does not appear to be essential, as PhoE mutants that lack the C-terminal phenylalanine can interact with BamA, albeit less efficiently37. Furthermore, in vivo, low-level expression can be tolerated, which leads to assembly of the mutant protein and suggests that other, currently unknown, motifs have a role in targeting and/or the kinetics of PhoE folding37.
Kahne and co-workers proposed that POTRA domains interact with folding substrates using β-augmentation29. They investigated this hypothesis by making mutations in POTRA3 to disrupt its propensity for β-pairing, and found that, although BamA function was maintained, its interactions with BamB were compromised. Knowles et al. tested the interactions of four peptides derived from the OMP porin PhoE, and found that individual BamA POTRA domains can function as discrete binding units by directly contacting OMP sequences through interactions with the alternating sides of the β-sheets of the first two POTRA domains30. This function seems to be conserved, as the POTRA domains of the mitochondrial homologue, Sam50, also bind β-barrel precursors39. The binding of various β-strand peptides other than the C-terminal strand suggests that the targeting motif is recognized elsewhere in the BAM complex and that the POTRA domain guides the nascent OMPs through the core complex by weak interactions that permit processive sliding30.
The functional importance of having five POTRA domains in all BamA proteins remains unclear, as deletion studies have yielded conflicting results. Tommassen and co-workers serially removed N. meningitidis BamA POTRA domains and found that only POTRA5 is essential; removal of the other domains resulted in only slight defects in OMP assembly40. By contrast, Kahne and colleagues removed individual POTRA domains from E. coli BamA and found that the three C-terminal POTRA domains are essential, whereas removal of POTRA1 or POTRA2 compromised growth, suggesting that all domains have important roles29. These differences may simply reflect the inability of accessory components to function in association with the BAM complex, as recent studies suggest that POTRA domains also act as a scaffold for the binding of accessory factors. In E. coli, BamB disengages when any POTRA domain other than POTRA1 is removed, whereas removal of POTRA5 leads to loss of all accessory factors29. In N. meningitidis, which lacks BamB, only deletion of POTRA5 leads to accessory protein loss40. However, this does not adequately explain why both E. coli and N. meningitidis, and all other Gram-negative bacteria, have precisely five POTRA domains. An explanation is offered by a study of N. meningitidis BamA in which the correct folding of larger OMPs correlated with the number of POTRA domains present, suggesting that more POTRA domains are needed to fold larger OMPs40.
The BAM accessory components
The four BAM accessory lipoproteins, BamB–E, form a tight complex with E. coli BamA based on their co-purification12,18. Although BamA–D exist in equal stoichiometry41, it is unclear whether this is also the case for BamE. Furthermore, it is possible that the native BAM complex is multimeric, as BamA and other BamA-like proteins have been shown to multimerize in vitro27,37,42,43. The interactions between BAM components are becoming apparent from by mutagenesis and binding studies. BamA interacts stably with the associated BamC, BamD and BamE components through an interaction that requires POTRA5 (Refs 14, 18, 29). BamB makes a direct interaction with BamA that can occur independently of the other components and is mediated by the Pro171–Pro181 and Glu221–Asp229 sequences and the four most C-terminal POTRA domains14,19,29. The participation of BamC in the complex requires the C terminus of BamD14, whereas BamD itself makes a direct contact with BamA, an interaction that is thought to be stabilized by BamE14,18.
All of the BAM lipoproteins have roles in OMP biogenesis, as their depletion leads to varying degrees of OMP assembly defects, but only BamD and BamA are crucial for cell viability and OMP biogenesis11,12,13,14. BamD is ubiquitous in Gram-negative bacteria14 and was previously described in Neisseria gonorrhoeae as the peptidoglycan-associated competence lipoprotein ComL44. Whereas BamD is essential, transposon insertion into the C termini of both the N. gonorrhoeae and E. coli bamD alleles yields organisms that are viable but have major cellular defects characterized by aberrant cellular morphology44 and decreased steady state levels of OMPs, respectively3,7. These partial losses of function suggest that the essential role of BamD is mediated by its N-terminal region. BamD has no obvious similarity to proteins of known structure, but homologues in Rickettsia spp. and other alphaproteobacteria have been predicted to contain up to six tetratricopeptide repeat motifs that form tandem helix–loop–helix structures and are associated with protein–protein interactions21,45,46. Such motifs contribute to other protein transport pathways, including that of the mitochondrial protein import receptor Tom70 (Refs 47, 48), which binds β-barrel substrate proteins en route to the mitochondrial homologue of the BAM47. It is possible that BamD performs a similarly important protein handling function that accounts for its obligatory requirement in the outer membrane.
In contrast to BamD, bamB-null strains are viable but are hypersensitive to antibiotics, such as vancomycin, which shows that the outer membrane permeability barrier is severely compromised49 and harbours defects in the correct assembly of various OMPs9,50. BamB is highly conserved among many Gram-negative bacteria, but is absent from some genomes, such as that of N. meningitidis and N. gonorrhoeae. Deletion of bamB attenuates some pathogenic bacteria51 and BamB has been linked to DNA-break repair and homologous recombination52. BamB is predicted to have a β-propeller fold with seven or eight blades based on homology to other proteins19,21. Using this information, Gatsos et al. have proposed a bioinformatic model of the BamB structure, and they suggest that its β-propeller fold could pair with the exposed β-strands of the POTRA domains of BamA or stabilize nascent β-strands of substrate proteins or both21.
The role of BamC in OMP biogenesis remains mysterious. E. coli strains in which bamC has been deleted show moderate outer membrane permeability defects, including sensitivity to rifampicin, yet retain viability and the ability to assemble OMPs9,12. Furthermore, BamC is not ubiquitous throughout Gram-negative bacteria, as no homologue has been found in any of the alphaproteobacteria for which genomes have been sequenced. BamC also lacks significant similarity to any protein of known structure, suggesting that it possesses novel folds or features.
BamE is present in all alpha-, beta- and gammaproteobacteria. Although BamE is not an essential BAM subunit, null mutants exhibit OMP folding defects and increased sensitivity to agents such as rifampicin and SDS, reflecting a compromised barrier function18. The structure of a BamE homologue, OmlA, from Xanthomonas axonopodis, has been determined33. As in E. coli, OmlA is required for outer membrane integrity and consists of a POTRA-like fold, although the order of secondary structural elements differs. There are currently no functional data on this protein except for the phenotype observed on its deletion. However, its similarity to the POTRA fold suggests it may be able to bind nascent OMPs or other BAM components using β-augmentation. The exposed β-strands in OmlE (BamE), like those in BamB, seem to be well suited for β-augmentation and could also assist in binding and folding nascent OMPs. The independent binding of these two accessory proteins to BamA might reflect the processing of different OMP subsets, with BamC–E functioning in one pathway, and BamB in another.
BAM interactions with periplasmic chaperones
The roles of the BAM accessory components remain enigmatic. However, they could either function as independent chaperones or as docking sites for periplasmic chaperones that carry nascent OMPs. Both putative functions would enable transfer of the OMPs to the BAM complex9. In the docking site model, release of the chaperone back into the periplasmic pool could then trigger the BAM complex to fold the protein into the outer membrane (Fig. 1). Although several periplasmic chaperones are present, DegP, SurA and Skp have been implicated as the major factors that transport and target OMPs from the Sec machinery to the BAM complex53. Paradoxically, DegP has both protease and chaperone activity and is regulated in a temperature-dependent manner54,55. SurA is a member of the peptidyl-prolyl isomerase family, but also has general chaperone activity and shows outer membrane permeability defects on depletion56. Skp is a general chaperone that has been shown to bind denatured OMPs, but not denatured periplasmic or cytosolic proteins57. Previous double knockout experiments have revealed functional redundancy among these chaperones, suggesting that Skp and DegP function in one pathway, whereas SurA acts in a separate, parallel pathway8,9,10. Both SurA and Skp have been shown to interact with OMPs as they leave the Sec translocon; for SurA this was revealed by kinetic analysis of LamB assembly, which suggested that SurA interacts before signal sequence cleavage58. By contrast, direct interaction of Skp with OmpA and PhoE was observed while the OMPs were still in complex with the Sec translocon59,60. However, only SurA has been shown to interact with the BAM complex, either directly or through a substrate protein, and does so in a manner that is independent of BamB8,19. No direct evidence for the binding of Skp or DegP has been presented, although deletion studies have shown that a ΔbamC mutant exhibited genetic interactions that were similar to those of a skp–degP double knockout, whereas a bamC–surA knockout produced a synthetic lethal phenotype, prompting the proposal that BamC is part of the Skp–DegP folding pathway9.
Sklar et al. suggest that the SurA pathway is primarily responsible for the assembly of most OMPs, whereas Skp and DegP rescue those that have fallen off the normal assembly route8. This would explain why in a surA, skp or degP depletion strain, OMP levels are reduced but not abolished8 and why depletion of surA produces a more drastic effect on OMP biogenesis than either skp or degP. However, this model does not explain why only Skp has been shown to cross link to OmpA and PhoE on entering the periplasm from the Sec machinery59,60. It is more plausible, although less elegant, that certain OMPs are specifically recruited to the SurA pathway, whereas others are designated for the Skp–DegP pathway.
A bamB–surA double knockout was previously shown to exhibit a synthetic lethal phenotype9, which suggested that BamB functions in concert with the Skp–DegP pathway, an observation that is consistent with the BamB-independent binding of SurA to the BAM complex19. However, a recent study by Typas et al. suggests that the bamB–surA double knockout is only lethal under high growth conditions61, indicating that its previous lethality is not the result of an absolute requirement for one of the gene products but rather is due to a kinetic effect on OMP processing. In the absence of both BamB and SurA, OMP folding and insertion is dramatically reduced but not completely abolished, implying that the observed phenotype is a cumulative effect of the loss of two non-redundant functions. Furthermore, a conditional lethal bamB–degP double knockout cannot be rescued by either Skp or SurA, suggesting that there is little or no functional overlap between BamB and either Skp or SurA50. Interestingly, a bamB double knockout with another periplasmic chaperone, FkpA, a heat shock peptidyl-prolyl cis–trans isomerase62, also produces a synthetic lethal phenotype9. FkpA is associated with the folding of soluble periplasmic proteins, such as the maltose-binding protein MalE, rather than with OMPs, although its substrate requirements are still not completely clear62. Taken together, these investigations suggest that BamB is involved in multiple periplasmic folding pathways9 and may function late in OMP biogenesis after an interaction of the chaperones with the BAM complex.
Proposed models of insertion
It is clear from the biochemical, genetic and structural studies discussed above that OMPs interact with periplasmic chaperones and the BAM complex before insertion into the outer membrane. However, the precise sequence of events and the route to the outer membrane remains puzzling. There are several mechanisms by which substrate OMPs might conceivably be inserted into the membrane (Fig. 1). In the pore-folding model, the BAM complex could function as a single monomeric complex and incorporate the nascent OMP into the β-barrel pore formed by BamA, while the periplasmic components of the complex act as targets for the periplasmic chaperones and thread the nascent OMP into the barrel core. In support of this model, BamA and its distant homologues exhibit pore activity that is responsive to substrate binding37,63,64. This activity seems to be modulated by the conserved extracellular loop close to the C terminus of the β-barrel, as demonstrated by structural studies of FhaC63. In the FhaC crystal structure, this loop, together with the N-terminal helix, is buried in the barrel pore, forming a 3 Å diameter channel. This contrasts with the 8–10 Å pore estimated from membrane conductivity experiments and suggests that the channel-blocking components are dynamic and can vacate the barrel32,63. Other studies of this loop have shown that similar topological rearrangements occur owing to co-expression of its substrate65. It therefore seems likely that BamA homologues involved in secretion, such as FhaC, expel components of the β-barrel core to clear a pathway for the translocation of substrate across the membrane. However, it is unlikely that the network of hydrogen bonds in the BamA β-barrel could rupture to allow lateral passage of the substrate OMP into the bilayer because of the structural destabilization required.
A derivative of this first model is the complex pore-folding model. If the BAM complex does oligomerize in vivo, as has been suggested from BamA in vitro studies (still under debate), a central pore could be formed that is lined by BamA β-barrels. Local distortions in the lipid population could then favour assembly and insertion of OMP structures into the membrane. Release of folded OMP into the membrane could then conceivably occur owing to the opening of the oligomeric assembly. In the complex pore-folding model, the periplasmic components of the BAM complex would help fold and deposit the protein directly into the membrane.
In a third model, the barrel-folding model, the BamA structure provides a surface onto which the interacting OMP folds in the vicinity of the membrane27. This model, together with the complex pore-folding model, is supported by conductance studies that detected a closed, low-conductance channel that is not widened by substrate binding.
The final models for BAM complex function have arisen following the recent publication of multimeric DegP structures detected by both electron microscopy and X-ray crystallography66,67,68. Previous studies of DegP have shown that it can assume both trimeric and hexameric structures and that it predominantly adopts its hexameric form when inactive; indeed, this was the structure observed when its crystal structure was first determined69. However, it has recently been found that when presented with a substrate protein, DegP can form large 12- or 24-mer cage-like structures66,67,68 (Fig. 3). Krojer et al. showed that both the 12-mer and 24-mer DegP cage-like structures can be co-purified with substrates such as OmpA, OmpC, OmpF and LamB, and that these proteins are folded and protected from degradation, suggesting that DegP is a chaperone or carrier for folded rather than unfolded OMPs67. Visualization of the 12-mer complex by electron microscopy reveals a folded OMP in the core of the DegP cage. Unfortunately, the bound OMP cargo in the 24-mer X-ray structure could not be resolved, perhaps owing to conformational and chemical heterogeneity. It is currently unclear whether the two cage forms adopt different functions in vivo, but in vitro the 24-mer cage that contained its OMP cargo was shown to have putative membrane attachment sites and indeed was shown to interact with lipid membranes.
a | A ribbon model of the asymmetric DegP12–outer membrane protein (OMP) complex viewed along its approximate three-fold axis that was calculated using electron microscopy (EM). DegP12 is in grey, and an OmpC monomer (red) that was modelled from the EM density is shown in the central pore. b | A ribbon model of DegP24 that shows its overall architecture; the trimeric units are coloured differently. The molecule is shown along its four-fold axis.
In light of the discovery that DegP functions as a chaperone for folded OMPs, DegP could conceivably operate at two points in the OMP biogenesis pathway. In the chaperone folding model, DegP acts immediately after release of the nascent OMP from the Sec machinery and before its interaction with the BAM complex. Alternatively, in the accessory folding model, DegP acts following an interaction with the BAM complex. We suggest that DegP, and potentially other periplasmic chaperones, could actively fold the nascent OMPs by completely encapsulating and sequestering them away from the periplasm following their release from the Sec machinery. Thus, on delivery to the outer membrane, the BAM complex would only function in the insertion of the protein into the membrane rather than in its folding70. Krojer et al.67 speculate that these cage structures might actually be large enough to span the periplasmic space and interact with both the inner and outer membranes, and therefore function as a macropore, allowing the protected diffusion of OMP precursors from the inner membrane to the outer membrane. However, in this model, the interaction of SurA and Skp with the OMPs is difficult to envisage, as presumably such a macropore would exclude such proteins from the nascent OMP. We propose that a more rational explanation for the currently available data is the accessory folding model. In this model, the periplasmic chaperones, such as SurA, deliver the nascent OMPs to the periplasmic components of the BAM complex, and then act in concert to fold the OMPs. Once folded, the OMPs could be guided into the outer membrane, possibly with the assistance of an additional periplasmic factor. The observation that unfolded OMPs are degraded owing to the protease activity of DegP, whereas folded OMPs are not, together with the observation of a folded OMP in the centre of the DegP cage, is compelling evidence for this model67. Furthermore, once loaded with its OMP cargo, DegP can interact directly with lipid67 and could potentially help insert the protein into the membrane. The use of DegP at this stage would be an ideal quality-control mechanism for the removal of misfolded proteins before their insertion into the membrane. In this model, the deletion of a single accessory factor or chaperone might not be lethal but would drastically impair the rate of OMP insertion into the outer membrane, an observation that is supported by the current literature. Deletion of more than one accessory factor would severely compromise the kinetics of OMP insertion and could lead to a synthetic lethal effect.
Assembly of OMP oligomers?
Although we can now rationalize the involvement of the BAM complex in the assembly of most OMPs, the assembly of trimeric OMPs, such as TolC71, presents an interesting conundrum. TolC trimers form a β-barrel, to which each monomer contributes four β-strands, in the outer membrane71. TolC is dependent on BamA for its assembly, but how the BAM complex contributes to the formation of such oligomers is perplexing11. Could the complex hold one monomer and wait until the second and third monomers bind before folding and releasing the trimeric β-barrel? This seems implausible, as the BAM complex would need to prevent different trimeric OMP monomers, and potentially other nascent OMPs, from binding. An alternative mechanism is that the BAM complex simply folds and inserts each monomer into the membrane, and that formation of the trimeric OMP then occurs through diffusion. However, this in itself seems somewhat stochastic: all monomers presumably have similar β-strand binding surfaces within the membrane, and therefore monomers from different trimeric OMPs could potentially associate. If such a process does occur, we suggest that the BAM complex and other accessory proteins are more likely to be involved in orchestrating the process.
Concluding remarks
Considerable advances in the field of OMP biogenesis have been made over the past year. However, many questions remain. What are the precise roles of the accessory factors? What is the process of OMP insertion: lateral passage through the BamA β-barrel, the use of annealing along the barrel surface or the use of accessory proteins? Where and how does SurA transfer bound nascent OMP to the BAM complex, and how can Skp–DegP take over the role of SurA in a ΔsurA mutant? In the coming years it is likely that the structures of more members of the complex will be solved, further progressing our understanding of the mechanisms involved. However, determining the functional interactions that occur during OMP assembly will be challenging owing to the complex protein and membrane interactions that pave the OMP assembly pathway and the diversity of the folding substrates that pass through them.
References
Bos, M. P., Robert, V. & Tommassen, J. Biogenesis of the Gram-negative bacterial outer membrane. Annu. Rev. Microbiol. 61, 191–214 (2007).
Breyton, C., Haase, W., Rapoport, T. A., Kuhlbrandt, W. & Collinson, I. Three-dimensional structure of the bacterial protein-translocation complex SecYEG. Nature 418, 662–665 (2002).
Van den Berg, B. et al. X-ray structure of a protein-conducting channel. Nature 427, 36–44 (2004).
Driessen, A. J. & Nouwen, N. Protein translocation across the bacterial cytoplasmic membrane. Annu. Rev. Biochem. 77, 643–667 (2008).
Rapoport, T. A. Protein translocation across the eukaryotic endoplasmic reticulum and bacterial plasma membranes. Nature 450, 663–669 (2007).
Qi, H. Y., Hyndman, J. B. & Bernstein, H. D. DnaK promotes the selective export of outer membrane protein precursors in SecA-deficient Escherichia coli. J. Biol. Chem. 277, 51077–51083 (2002).
Papanikou, E., Karamanou, S. & Economou, A. Bacterial protein secretion through the translocase nanomachine. Nature Rev. Microbiol. 5, 839–851 (2007).
Sklar, J. G., Wu, T., Kahne, D. & Silhavy, T. J. Defining the roles of the periplasmic chaperones SurA, Skp, and DegP in Escherichia coli. Genes Dev. 21, 2473–2484 (2007).
Onufryk, C., Crouch, M. L., Fang, F. C. & Gross, C. A. Characterization of six lipoproteins in the σE regulon. J. Bacteriol. 187, 4552–4561 (2005).
Rizzitello, A. E., Harper, J. R. & Silhavy, T. J. Genetic evidence for parallel pathways of chaperone activity in the periplasm of Escherichia coli. J. Bacteriol. 183, 6794–6800 (2001).
Werner, J. & Misra, R. YaeT (Omp85) affects the assembly of lipid-dependent and lipid-independent outer membrane proteins of Escherichia coli. Mol. Microbiol. 57, 1450–1459 (2005).
Wu, T. et al. Identification of a multicomponent complex required for outer membrane biogenesis in Escherichia coli. Cell 121, 235–245 (2005). First documented evidence that BamA forms a heterooligomeric structure; the BamB–D lipoproteins were identified as accessory components of the complex.
Doerrler, W. T. & Raetz, C. R. Loss of outer membrane proteins without inhibition of lipid export in an Escherichia coli YaeT mutant. J. Biol. Chem. 280, 27679–27687 (2005).
Malinverni, J. C. et al. YfiO stabilizes the YaeT complex and is essential for outer membrane protein assembly in Escherichia coli. Mol. Microbiol. 61, 151–164 (2006).
Collin, S., Guilvout, I., Chami, M. & Pugsley, A. P. YaeT-independent multimerization and outer membrane association of secretin PulD. Mol. Microbiol. 64, 1350–1357 (2007).
Guilvout, I. et al. In vitro multimerization and membrane insertion of bacterial outer membrane secretin PulD. J. Mol. Biol. 382, 13–23 (2008).
Bolla, J. M., Lazdunski, C. & Pages, J. M. The assembly of the major outer membrane protein OmpF of Escherichia coli depends on lipid synthesis. EMBO J. 7, 3595–3599 (1988).
Sklar, J. G. et al. Lipoprotein SmpA is a component of the YaeT complex that assembles outer membrane proteins in Escherichia coli. Proc. Natl Acad. Sci. USA 104, 6400–6405 (2007).
Vuong, P., Bennion, D., Mantei, J., Frost, D. & Misra, R. Analysis of YfgL and YaeT interactions through bioinformatics, mutagenesis, and biochemistry. J. Bacteriol. 190, 1507–1517 (2008).
Misra, R. First glimpse of the crystal structure of YaeT's POTRA domains. ACS Chem. Biol. 2, 649–651 (2007).
Gatsos, X. et al. Protein secretion and outer membrane assembly in Alphaproteobacteria. FEMS Microbiol. Rev. 32, 995–1009 (2008).
Aoki, S. K. et al. Contact-dependent growth inhibition requires the essential outer membrane protein BamA (YaeT) as the receptor and the inner membrane transport protein AcrB. Mol. Microbiol. 70, 323–340 (2008).
Voulhoux, R., Bos, M. P., Geurtsen, J., Mols, M. & Tommassen, J. Role of a highly conserved bacterial protein in outer membrane protein assembly. Science 299, 262–265 (2003). The authors of this paper described, for the first time, the essential nature of BamA and its role in the biogenesis of membrane proteins.
Genevrois, S., Steeghs, L., Roholl, P., Letesson, J. J. & van der Ley, P. The Omp85 protein of Neisseria meningitidis is required for lipid export to the outer membrane. EMBO J. 22, 1780–1789 (2003).
Bos, M. P., Tefsen, B., Geurtsen, J. & Tommassen, J. Identification of an outer membrane protein required for the transport of lipopolysaccharide to the bacterial cell surface. Proc. Natl Acad. Sci. USA 101, 9417–9422 (2004).
Sanchez-Pulido, L., Devos, D., Genevrois, S., Vicente, M. & Valencia, A. POTRA: a conserved domain in the FtsQ family and a class of β-barrel outer membrane proteins. Trends Biochem. Sci. 28, 523–526 (2003). The POTRA domains were identified in a wide range of proteins through in silico predictions.
Schleiff, E. & Soll, J. Membrane protein insertion: mixing eukaryotic and prokaryotic concepts. EMBO Rep. 6, 1023–1027 (2005).
Gentle, I. E., Burri, L. & Lithgow, T. Molecular architecture and function of the Omp85 family of proteins. Mol. Microbiol. 58, 1216–1225 (2005).
Kim, S. et al. Structure and function of an essential component of the outer membrane protein assembly machine. Science 317, 961–964 (2007). First identification of the crystal structure of POTRA 1–4 from E. coli BamA, which revealed the first structure of a POTRA fold.
Knowles, T. J. et al. Fold and function of polypeptide transport-associated domains responsible for delivering unfolded proteins to membranes. Mol. Microbiol. 68, 1216–1227 (2008). A description of the NMR structures of POTRA 1–2 from E. coli BamA; this study detected interdomain flexibility and evidence for direct binding of nascent barrel proteins.
Gatzeva-Topalova, P. Z., Walton, T. A. & Sousa, M. C. Crystal structure of YaeT: conformational flexibility and substrate recognition. Structure 16, 1873–1881 (2008).
Clantin, B. et al. Structure of the membrane protein FhaC: a member of the Omp85–TpsB transporter superfamily. Science 317, 957–961 (2007). This paper documents the crystal structure of FhaC, a two-partner secretion system OMP that is related to BamA. The authors found a tandem POTRA domain fold that was associated with an integral outer membrane barrel domain.
Vanini, M. M., Spisni, A., Sforca, M. L., Pertinhez, T. A. & Benedetti, C. E. The solution structure of the outer membrane lipoprotein OmlA from Xanthomonas axonopodis pv. citri reveals a protein fold implicated in protein–protein interaction. Proteins 71, 2051–2064 (2008).
Reynolds, K. A. et al. Structural and computational characterization of the SHV-1 β-lactamase-β-lactamase inhibitor protein interface. J. Biol. Chem. 281, 26745–26753 (2006).
Williams, J. C. et al. Structural and thermodynamic characterization of a cytoplasmic dynein light chain-intermediate chain complex. Proc. Natl Acad. Sci. USA 104, 10028–10033 (2007).
van den Ent, F. et al. Structural and mutational analysis of the cell division protein FtsQ. Mol. Microbiol. 68, 110–123 (2008).
Robert, V. et al. Assembly factor Omp85 recognizes its outer membrane protein substrates by a species-specific C-terminal motif. PLoS Biol. 4, e377 (2006).
Struyve, M., Moons, M. & Tommassen, J. Carboxy-terminal phenylalanine is essential for the correct assembly of a bacterial outer membrane protein. J. Mol. Biol. 218, 141–148 (1991).
Habib, S. J. et al. The N-terminal domain of Tob55 has a receptor-like function in the biogenesis of mitochondrial β-barrel proteins. J. Cell Biol. 176, 77–88 (2007).
Bos, M. P., Robert, V. & Tommassen, J. Functioning of outer membrane protein assembly factor Omp85 requires a single POTRA domain. EMBO Rep. 8, 1149–1154 (2007).
Stenberg, F. et al. Protein complexes of the Escherichia coli cell envelope. J. Biol. Chem. 280, 34409–34419 (2005).
Surana, N. K. et al. Evidence for conservation of architecture and physical properties of Omp85-like proteins throughout evolution. Proc. Natl Acad. Sci. USA 101, 14497–14502 (2004).
Li, H., Grass, S., Wang, T., Liu, T. & St Geme, J. W. 3rd. Structure of the Haemophilus influenzae HMW1B translocator protein: evidence for a twin pore. J. Bacteriol. 189, 7497–7502 (2007).
Fussenegger, M., Facius, D., Meier, J. & Meyer, T. F. A novel peptidoglycan-linked lipoprotein (ComL) that functions in natural transformation competence of Neisseria gonorrhoeae. Mol. Microbiol. 19, 1095–1105 (1996).
Blatch, G. L. & Lassle, M. The tetratricopeptide repeat: a structural motif mediating protein–protein interactions. Bioessays 21, 932–939 (1999).
D'Andrea, L. D. & Regan, L. TPR proteins: the versatile helix. Trends Biochem. Sci. 28, 655–662 (2003).
Chan, N. C., Likic, V. A., Waller, R. F., Mulhern, T. D. & Lithgow, T. The C-terminal TPR domain of Tom70 defines a family of mitochondrial protein import receptors found only in animals and fungi. J. Mol. Biol. 358, 1010–1022 (2006).
Wu, Y. & Sha, B. Crystal structure of yeast mitochondrial outer membrane translocon member Tom70p. Nature Struct. Mol. Biol. 13, 589–593 (2006).
Ruiz, N., Falcone, B., Kahne, D. & Silhavy, T. J. Chemical conditionality: a genetic strategy to probe organelle assembly. Cell 121, 307–317 (2005).
Charlson, E. S., Werner, J. N. & Misra, R. Differential effects of yfgL mutation on Escherichia coli outer membrane proteins and lipopolysaccharide. J. Bacteriol. 188, 7186–7194 (2006).
Rolhion, N., Barnich, N., Claret, L. & Darfeuille-Michaud, A. Strong decrease in invasive ability and outer membrane vesicle release in Crohn's disease-associated adherent-invasive Escherichia coli strain LF82 with the yfgL gene deleted. J. Bacteriol. 187, 2286–2296 (2005).
Khairnar, N. P., Kamble, V. A., Mangoli, S. H., Apte, S. K. & Misra, H. S. Involvement of a periplasmic protein kinase in DNA strand break repair and homologous recombination in Escherichia coli. Mol. Microbiol. 65, 294–304 (2007).
Ruiz, N., Kahne, D. & Silhavy, T. J. Advances in understanding bacterial outer-membrane biogenesis. Nature Rev. Microbiol. 4, 57–66 (2006).
Lipinska, B., Zylicz, M. & Georgopoulos, C. The HtrA (DegP) protein, essential for Escherichia coli survival at high temperatures, is an endopeptidase. J. Bacteriol. 172, 1791–1797 (1990).
Spiess, C., Beil, A. & Ehrmann, M. A temperature-dependent switch from chaperone to protease in a widely conserved heat shock protein. Cell 97, 339–347 (1999).
Behrens, S., Maier, R., de Cock, H., Schmid, F. X. & Gross, C. A. The SurA periplasmic PPIase lacking its parvulin domains functions in vivo and has chaperone activity. EMBO J. 20, 285–294 (2001).
Chen, R. & Henning, U. A periplasmic protein (Skp) of Escherichia coli selectively binds a class of outer membrane proteins. Mol. Microbiol. 19, 1287–1294 (1996).
Ureta, A. R., Endres, R. G., Wingreen, N. S. & Silhavy, T. J. Kinetic analysis of the assembly of the outer membrane protein LamB in Escherichia coli mutants each lacking a secretion or targeting factor in a different cellular compartment. J. Bacteriol. 189, 446–454 (2007).
Schafer, U., Beck, K. & Muller, M. Skp, a molecular chaperone of Gram-negative bacteria, is required for the formation of soluble periplasmic intermediates of outer membrane proteins. J. Biol. Chem. 274, 24567–24574 (1999).
Harms, N. et al. The early interaction of the outer membrane protein PhoE with the periplasmic chaperone Skp occurs at the cytoplasmic membrane. J. Biol. Chem. 276, 18804–18811 (2001).
Typas, A. et al. High-throughput, quantitative analyses of genetic interactions in E. coli. Nature Methods 5, 781–787 (2008).
Arie, J. P., Sassoon, N. & Betton, J. M. Chaperone function of FkpA, a heat shock prolyl isomerase, in the periplasm of Escherichia coli. Mol. Microbiol. 39, 199–210 (2001).
Meli, A. C. et al. Channel properties of TpsB transporter FhaC point to two functional domains with a C-terminal protein-conducting pore. J. Biol. Chem. 281, 158–166 (2006).
Ertel, F. et al. The evolutionarily related β-barrel polypeptide transporters from Pisum sativum and Nostoc PCC7120 contain two distinct functional domains. J. Biol. Chem. 280, 28281–28289 (2005).
Guedin, S. et al. Novel topological features of FhaC, the outer membrane transporter involved in the secretion of the Bordetella pertussis filamentous hemagglutinin. J. Biol. Chem. 275, 30202–30210 (2000).
Krojer, T. et al. Interplay of PDZ and protease domain of DegP ensures efficient elimination of misfolded proteins. Proc. Natl Acad. Sci. USA 105, 7702–7707 (2008).
Krojer, T. et al. Structural basis for the regulated protease and chaperone function of DegP. Nature 453, 885–890 (2008). This study indicated that DegP can form higher order homooligomers and is capable of interacting with folded outer membrane proteins and with lipid.
Jiang, J. et al. Activation of DegP chaperone-protease via formation of large cage-like oligomers upon binding to substrate proteins. Proc. Natl Acad. Sci. USA 105, 11939–11944 (2008).
Krojer, T., Garrido-Franco, M., Huber, R., Ehrmann, M. & Clausen, T. Crystal structure of DegP (HtrA) reveals a new protease-chaperone machine. Nature 416, 455–459 (2002).
Huber, D. & Bukau, B. DegP: a protein “death star”. Structure 16, 989–990 (2008).
Koronakis, V., Sharff, A., Koronakis, E., Luisi, B. & Hughes, C. Crystal structure of the bacterial membrane protein TolC central to multidrug efflux and protein export. Nature 405, 914–919 (2000).
Kozjak, V. et al. An essential role of Sam50 in the protein sorting and assembly machinery of the mitochondrial outer membrane. J. Biol. Chem. 278, 48520–48523 (2003).
Inoue, K. & Potter, D. The chloroplastic protein translocation channel Toc75 and its paralog OEP80 represent two distinct protein families and are targeted to the chloroplastic outer envelope by different mechanisms. Plant J. 39, 354–365 (2004).
Henderson, I. R., Navarro-Garcia, F., Desvaux, M., Fernandez, R. C. & Ala'Aldeen, D. Type V protein secretion pathway: the autotransporter story. Microbiol. Mol. Biol. Rev. 68, 692–744 (2004).
Wiedemann, N. et al. Biogenesis of the protein import channel Tom40 of the mitochondrial outer membrane: intermembrane space components are involved in an early stage of the assembly pathway. J. Biol. Chem. 279, 18188–18194 (2004).
Pfanner, N., Wiedemann, N., Meisinger, C. & Lithgow, T. Assembling the mitochondrial outer membrane. Nature Struct. Mol. Biol. 11, 1044–1048 (2004).
Kutik, S. et al. Dissecting membrane insertion of mitochondrial β-barrel proteins. Cell 132, 1011–1024 (2008).
Meisinger, C. et al. The mitochondrial morphology protein Mdm10 functions in assembly of the preprotein translocase of the outer membrane. Dev. Cell 7, 61–71 (2004).
Meisinger, C. et al. The morphology proteins Mdm12/Mmm1 function in the major β-barrel assembly pathway of mitochondria. EMBO J. 26, 2229–2239 (2007).
Author information
Authors and Affiliations
Corresponding author
Related links
Related links
DATABASES
Entrez Genome Project
Glossary
- Chaperone
-
A protein that facilitates the proper folding of other proteins.
- Amphipathic
-
A molecule with both polar and non-polar portions in its structure.
Rights and permissions
About this article
Cite this article
Knowles, T., Scott-Tucker, A., Overduin, M. et al. Membrane protein architects: the role of the BAM complex in outer membrane protein assembly. Nat Rev Microbiol 7, 206–214 (2009). https://doi.org/10.1038/nrmicro2069
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrmicro2069
This article is cited by
-
Aminolipids elicit functional trade-offs between competitiveness and bacteriophage attachment in Ruegeria pomeroyi
The ISME Journal (2023)
-
First report of whole genome sequence of septicemic Pasteurella multocida serovar B:2 ‘Soron’ strain isolated from swine
Brazilian Journal of Microbiology (2023)
-
Secretome analysis of an environmental isolate Enterobacter sp. S-33 identifies proteins related to pathogenicity
Archives of Microbiology (2022)
-
Structural basis for bacterial lipoprotein relocation by the transporter LolCDE
Nature Structural & Molecular Biology (2021)
-
Affinity of Skp to OmpC revealed by single-molecule detection
Scientific Reports (2020)