Key Points
-
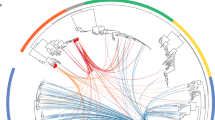
Genomic islands (GEIs) are characterized by their large size (>10 kb), their frequent association with tRNA-encoding genes and a different G+C content compared with the rest of the chromosome. Many genomic islands are flanked by repeat structures and carry fragments of other mobile and accessory genetic elements, such as bacteriophages, plasmids and insertion sequence (IS) elements.
-
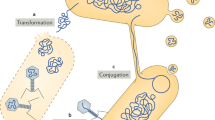
Some GEIs can excise themselves spontaneously from the chromosome and can be transferred to other suitable recipients. GEIs contribute to bacterial genome plasticity and, together with other mobile and accessory genetic elements, to the 'horizontal gene pool' of a given bacterial population.
-
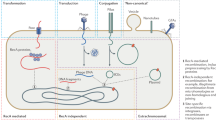
A hypothetical 'life cycle' of GEIs includes the insertion of mobile genetic elements into the bacterial chromosome. Through rearrangements and consecutive insertion and deletion events, the organization and gene content of the original element becomes modified and can lose the features of mobile elements. Owing to the action of bacteriophage integrases that are encoded on genomic islands, these genetic elements can be deleted from the chromosome and, upon transfer into a suitable host, can be chromosomally inserted by site-specific recombination.
-
GEIs contribute to fitness and adaptation. GEIs typically provide a gain-of-function to the host bacterium. As GEIs promote the transfer of multi-gene families, entire phenotypes can be changed in a single-step gene-transfer event.
-
GEIs are expected to have a role in ecological niches where microbial cell numbers and diversity are high and/or in environments that are constantly changing. The GEIs identified so far are relevant in the context of pathogenicity, symbiosis, antibiotic resistance, xenobiotic degradation, and primary and secondary metabolism. It is expected that the functional diversity of GEIs is even greater than is currently known.
-
As GEIs are widely distributed in pathogenic, non-pathogenic and environmental microorganisms, they represent a paradigm rather than a paradox for microbial evolution, underlining the importance of horizontal gene transfer in this process.
Abstract
Horizontal gene transfer is an important mechanism for the evolution of microbial genomes. Pathogenicity islands — mobile genetic elements that contribute to rapid changes in virulence potential — are known to have contributed to genome evolution by horizontal gene transfer in many bacterial pathogens. Increasing evidence indicates that equivalent elements in non-pathogenic species — genomic islands — are important in the evolution of these bacteria, influencing traits such as antibiotic resistance, symbiosis and fitness, and adaptation in general. This review discusses the recent lessons that have been learned from pathogenicity islands in pathogenic microorganisms and how they apply to the role of genomic islands in commensal, symbiotic and environmental bacteria.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Falkow, S. in Escherichia coli and Salmonella typhimurium: Cellular and Molecular Biology (eds Neidhardt, F. C. et al.) 2723–2729 (ASM Press, Washington DC, 1996).
Ochman, H., Lawrence, J. G. & Groisman, E. A. Lateral gene transfer and the nature of bacterial innovation. Nature 405, 299–304 (2000).
Boyd, E. F. & Brussow, H. Common themes among bacteriophage-encoded virulence factors and diversity among the bacteriophages involved. Trends Microbiol. 10, 521–529 (2002).
Shankar, N., Baghdayan, A. S. & Gilmore, M. S. Modulation of virulence within a pathogenicity island in vancomycin-resistant Enterococcus faecalis. Nature 417, 746–750 (2002).
Hacker, J. & Kaper, J. B. (eds) Pathogenicity Islands and the Evolution of Pathogenic Microbes (Springer, Berlin, 2002).
Schmidt, H. & Hensel, M. Pathogenicity islands in bacterial pathogenesis. Clin. Microbiol. Rev. 17, 14–56 (2004).
Hentschel, U. & Hacker, J. Pathogenicity islands: the tip of the iceberg. Microbes Infect. 3, 545–548 (2001).
Hacker, J. & Carniel, E. Ecological fitness, genomic islands and bacterial pathogenicity. A Darwinian view of the evolution of microbes. EMBO Rep. 2, 376–381 (2001). Describes the concept of genomic islands and their contribution to bacterial evolution.
Dobrindt, U. et al. Analysis of genome plasticity in pathogenic and commensal Escherichia coli isolates by use of DNA arrays. J. Bacteriol. 185, 1831–1840 (2003).
Preston, G. M., Haubold, B. & Rainey, P. B. Bacterial genomics and adaptation to life on plants: implications for the evolution of pathogenicity and symbiosis. Curr. Opin. Microbiol. 1, 589–597 (1998).
Lawrence, J. G. & Roth, J. R. Selfish operons: horizontal transfer may drive the evolution of gene clusters. Genetics 143, 1843–1860 (1996).
Maurelli, A. T., Fernandez, R. E., Bloch, C. A., Rode, C. K. & Fasano, A. 'Black holes' and bacterial pathogenicity: a large genomic deletion that enhances the virulence of Shigella spp. and enteroinvasive Escherichia coli. Proc. Natl. Acad. Sci. USA 95, 3943–3948 (1998). First detailed description of how specific loss of genetic information enhances bacterial virulence.
Gross, R., Hacker, J. & Goebel, W. The Leopoldina international symposium on parasitism, commensalism and symbiosis common themes, different outcome. Mol. Microbiol. 47, 1749–1758 (2003).
Moran, N. A. Microbial minimalism: genome reduction in bacterial pathogens. Cell 108, 583–586 (2002).
van Ham, R. C. et al. Reductive genome evolution in Buchnera aphidicola. Proc. Natl Acad. Sci. USA 100, 581–586 (2003).
Burrus, V., Pavlovic, G., Decaris, B. & Guedon, G. Conjugative transposons: the tip of the iceberg. Mol. Microbiol. 46, 601–610 (2002).
Schwartz, E. et al. Complete nucleotide sequence of pHG1: a Ralstonia eutropha H16 megaplasmid encoding key enzymes of H(2)-based ithoautotrophy and anaerobiosis. J. Mol. Biol. 332, 369–383 (2003).
Dimopoulou, I. D., Russell, J. E., Mohd-Zain, Z., Herbert, R. & Crook, D. W. Site-specific recombination with the chromosomal tRNALeu gene by the large conjugative Haemophilus resistance plasmid. Antimicrob. Agents Chemother. 46, 1602–1603 (2002).
Cheetham, B. F. & Katz, M. E. A role for bacteriophages in the evolution and transfer of bacterial virulence determinants. Mol. Microbiol. 18, 201–208 (1995).
Schleper, C., Holz, I., Janekovic, D., Murphy, J. & Zillig, W. A multicopy plasmid of the extremely thermophilic archaeon Sulfolobus effects its transfer to recipients by mating. J. Bacteriol. 177, 4417–4426 (1995).
She, Q. et al. Genetic profile of pNOB8 from Sulfolobus: the first conjugative plasmid from an archaeon. Extremophiles 2, 417–425 (1998).
Schleper, C., Kubo, K. & Zillig, W. The particle SSV1 from the extremely thermophilic archaeon Sulfolobus is a virus: demonstration of infectivity and of transfection with viral DNA. Proc. Natl Acad. Sci. USA 89, 7645–7649 (1992).
Peng, X., Holz, I., Zillig, W., Garrett, R. A. & She, Q. Evolution of the family of pRN plasmids and their integrase-mediated insertion into the chromosome of the crenarchaeon Sulfolobus solfataricus. J. Mol. Biol. 303, 449–454 (2000).
Kawarabayasi, Y. et al. Complete genome sequence of an aerobic thermoacidophilic crenarchaeon, Sulfolobus tokodaii strain7. DNA Res. 8, 123–140 (2001).
Dobrindt, U., Hentschel, U., Kaper, J. B. & Hacker, J. Genome plasticity in pathogenic and non-pathogenic enterobacteria. Curr. Top. Microbiol. Immunol. 264, 157–175 (2002).
Ochman, H. & Moran, N. A. Genes lost and genes found: evolution of bacterial pathogenesis and symbiosis. Science 292, 1096–1099 (2001).
Schmid, K., Ebner, R., Altenbuchner, J., Schmitt, R. & Lengeler, J. W. Plasmid-mediated sucrose metabolism in Escherichia coli K12: mapping of the scr genes of pUR400. Mol. Microbiol. 2, 1–8 (1988).
Sprenger, G. A. & Lengeler, J. W. Mapping of the sor genes for L-sorbose degradation in the chromosome of Klebsiella pneumoniae. Mol. Gen. Genet. 209, 352–359 (1987).
Hochhut, B., Jahreis, K., Lengeler, J. W. & Schmid, K. CTnscr94, a conjugative transposon found in enterobacteria. J. Bacteriol. 179, 2097–102 (1997).
Nesbø, C. L. & Doolittle, W. F. Targeting clusters of transferred genes in Thermotoga maritima. Environ. Microbiol. 5, 1144–1154 (2003). Describes the analysis of genome variability in the hyperthermophilic organism Thermotoga maritima by the identification of strain-specific genomic regions using genomic subtraction.
Hayashi, T. et al. Complete genome sequence of enterohemorrhagic Escherichia coli O157:H7 and genomic comparison with a laboratory strain K-12. DNA Res. 8, 11–22 (2001).
Perna, N. T. et al. Genome sequence of enterohaemorrhagic Escherichia coli O157:H7. Nature 409, 529–533 (2001).
Welch, R. A. et al. Extensive mosaic structure revealed by the complete genome sequence of uropathogenic Escherichia coli. Proc. Natl Acad. Sci. USA 99, 17020–17024 (2002).
Baar, C. et al. Complete genome sequence and analysis of Wolinella succinogenes. Proc. Natl Acad. Sci. USA 100, 11690–11695 (2003).
Fitzgerald, J. R., Sturdevant, D. E., Mackie, S. M., Gill, S. R. & Musser, J. M. Evolutionary genomics of Staphylococcus aureus: insights into the origin of methicillin-resistant strains and the toxic shock syndrome epidemic. Proc. Natl Acad. Sci. USA 98, 8821–8826 (2001).
Pridmore, R. D. et al. The genome sequence of the probiotic intestinal bacterium Lactobacillus johnsonii NCC 533. Proc. Natl Acad. Sci. USA 101, 2512–2517 (2004).
Copley, S. D. Evolution of a metabolic pathway for degradation of a toxic xenobiotic: the patchwork approach. Trends Biochem. Sci. 25, 261–265 (2000).
Top, E. M. & Springael, D. The role of mobile genetic elements in bacterial adaptation to xenobiotic organic compounds. Curr. Opin. Biotechnol. 14, 262–269 (2003).
Ravatn, R., Studer, S., Springael, D., Zehnder, A. J. & van der Meer, J. R. Chromosomal integration, tandem amplification, and deamplification in Pseudomonas putida F1 of a 105-kilobase genetic element containing the chlorocatechol degradative genes from Pseudomonas sp. strain B13. J. Bacteriol. 180, 4360–4369 (1998).
Toussaint, A. et al. The biphenyl- and 4-chlorobiphenyl-catabolic transposon Tn4371, a member of a new family of genomic islands related to IncP and Ti plasmids. Appl. Environ. Microbiol. 69, 4837–4845 (2003).
Weinel, C., Nelson, K. E. & Tümmler, B. Global features of the Pseudomonas putida KT2440 genome sequence. Environ. Microbiol. 4, 809–818 (2002).
Van der Meer, J. R. & Sentchilo, V. Genomic islands and the evolution of catabolic pathways in bacteria. Curr. Opin. Biotechnol. 14, 248–254 (2003).
Springael, D. et al. Community shifts in a seeded 3-chlorobenzoate degrading membrane biofilm reactor: indications for involvement of in situ horizontal transfer of the clc-element from inoculum to contaminant bacteria. Environ. Microbiol. 4, 70–80 (2002).
Müller, T. A., Werlen, C., Spain, J. & Van Der Meer, J. R. Evolution of a chlorobenzene degradative pathway among bacteria in a contaminated groundwater mediated by a genomic island in Ralstonia. Environ. Microbiol. 5, 163–173 (2003). Describes how metabolic pathway evolution can occur under natural conditions in a few steps through the activity of IS elements and formation of a genomic island.
Larbig, K. D. et al. Gene islands integrated into tRNAGly genes confer genome diversity on a Pseudomonas aeruginosa clone. J. Bacteriol. 184, 6665–6680 (2002).
Paulsen, I. T. et al. Role of mobile DNA in the evolution of vancomycin-resistant Enterococcus faecalis. Science 299, 2071–2074 (2003).
Rowe-Magnus, D. A., Guerout, A. M., Biskri, L., Bouige, P. & Mazel, D. Comparative analysis of superintegrons: engineering extensive genetic diversity in the Vibrionaceae. Genome Res. 13, 428–442 (2003).
Heidelberg, J. F. et al. DNA sequence of both chromosomes of the cholera pathogen Vibrio cholerae. Nature 406, 477–483 (2000).
Boyd, D. et al. Complete nucleotide sequence of a 43-kilobase genomic island associated with the multidrug resistance region of Salmonella enterica serovar Typhimurium DT104 and its identification in phage type DT120 and serovar Agona. J. Bacteriol. 183, 5725–5732 (2001).
Ito, T., Katayama, Y. & Hiramatsu, K. Cloning and nucleotide sequence determination of the entire mec DNA of pre-methicillin-resistant Staphylococcus aureus N315. Antimicrob. Agents Chemother. 43, 1449–1458 (1999).
Ito, T. et al. Structural comparison of three types of staphylococcal cassette chromosome mec integrated in the chromosome in methicillin-resistant Staphylococcus aureus. Antimicrob. Agents Chemother. 45, 1323–1336 (2001).
Kuroda, M. et al. Whole-genome sequencing of meticillin-resistant Staphylococcus aureus. Lancet 357, 1225–1240 (2001).
Luck, S. N., Turner, S. A., Rajakumar, K., Sakellaris, H. & Adler, B. Ferric dicitrate transport system (Fec) of Shigella flexneri 2a YSH6000 is encoded on a novel pathogenicity island carrying multiple antibiotic resistance genes. Infect. Immun. 69, 6012–6021 (2001).
Turner, S. A., Luck, S. N., Sakellaris, H., Rajakumar, K. & Adler, B. Molecular epidemiology of the SRL pathogenicity island. Antimicrob. Agents Chemother. 47, 727–734 (2003).
Hiramatsu, K., Katayama, Y., Yuzawa, H. & Ito, T. Molecular genetics of methicillin-resistant Staphylococcus aureus. Int. J. Med. Microbiol. 292, 67–74 (2002).
Beaber, J. W., Hochhut, B. & Waldor, M. K. Genomic and functional analyses of SXT, an integrating antibiotic resistance gene transfer element derived from Vibrio cholerae. J. Bacteriol. 184, 4259–4269 (2002).
Beaber, J. W., Hochhut, B. & Waldor, M. K. SOS response promotes horizontal dissemination of antibiotic resistance genes. Nature 427, 72–74 (2004).
Boltner, D., MacMahon, C., Pembroke, J. T., Strike, P. & Osborn, A. M. R391: a conjugative integrating mosaic comprised of phage, plasmid, and transposon elements. J. Bacteriol. 184, 5158–5169 (2002).
Osborn, A. M., Bruce, K. D., Ritchie, D. A. & Strike, P. The mercury resistance operon of the IncJ plasmid pMERH exhibits structural divergence from other Gram-negative mer operons. Microbiology 142, 337–345 (1996).
Egan, S., Wiener, P., Kallifidas, D. & Wellington, E. M. Transfer of streptomycin biosynthesis gene clusters within streptomycetes isolated from soil. Appl. Environ. Microbiol. 64, 5061–5063 (1998).
Egan, S., Wiener, P., Kallifidas, D. & Wellington, E. M. Phylogeny of Streptomyces species and evidence for horizontal transfer of entire and partial antibiotic gene clusters. Antonie Van Leeuwenhoek 79, 127–133 (2001).
Metsä-Ketelä, M. et al. Molecular evolution of aromatic polyketides and comparative sequence analysis of polyketide ketosynthase and 16S ribosomal DNA genes from various streptomyces species. Appl. Environ. Microbiol. 68, 4472–4479 (2002).
Bentley, S. D. et al. Complete genome sequence of the model actinomycete Streptomyces coelicolor A3(2). Nature 417, 141–147 (2002).
Ikeda, H. et al. Complete genome sequence and comparative analysis of the industrial microorganism Streptomyces avermitilis. Nature Biotechnol. 21, 526–531 (2003).
Lopez, J. V. Naturally mosaic operons for secondary metabolite biosynthesis: variability and putative horizontal transfer of discrete catalytic domains of the epothilone polyketide synthase locus. Mol. Genet. Genomics 270, 420–431 (2004).
Piel, J. A polyketide synthase–peptide synthetase gene cluster from an uncultured bacterial symbiont of Paederus beetles. Proc. Natl Acad. Sci. USA 99, 14002–14007 (2002).
Piel, J., Höfer, Y. & Hui, D. Evidence for a symbiosis Island involved in horizontal acquisition of pederin biosynthetic capabilities by the bacterial symbiont of Paederus fuscipes beetles. J. Bacteriol. 186, 1280–1286 (2004). Describes the characterization of a genomic island of a non-culturable symbiotic bacterium encoding genes that are involved in secondary metabolism.
Stinear, T. P. et al. Giant plasmid-encoded polyketide synthases produce the macrolide toxin of Mycobacterium ulcerans. Proc. Natl Acad. Sci. USA 101, 1345–1349 (2004).
Posey, J. E. & Gherardini, F. C. Lack of a role for iron in the Lyme disease pathogen. Science 288, 1651–1653 (2000).
Carniel, E., Guilvout, I. & Prentice, M. Characterization of a large chromosomal 'high-pathogenicity island' in biotype 1B Yersinia enterocolitica. J. Bacteriol. 178, 6743–6751 (1996).
Bach, S., de Almeida, A. & Carniel, E. The Yersinia high-pathogenicity island is present in different members of the family Enterobacteriaceae. FEMS Microbiol. Lett. 183, 289–294 (2000).
Ölschläger, T. A. et al. The high-pathogenicity island is absent in human pathogens of Salmonella enterica subspecies I but present in isolates of subspecies III and VI. J. Bacteriol. 185, 1107–1111 (2003).
Schubert, S., Rakin, A., Karch, H., Carniel, E. & Heesemann, J. Prevalence of the 'high-pathogenicity island' of Yersinia species among Escherichia coli strains that are pathogenic to humans. Infect. Immun. 66, 480–485 (1998).
Schubert, S., Dufke, S., Sorsa, J. & Heesemann, J. A novel integrative and conjugative element (ICE) of Escherichia coli: the putative progenitor of the Yersinia high-pathogenicity island. Mol. Microbiol. 51, 837–848 (2004).
Hueck, C. J. Type III protein secretion systems in bacterial pathogens of animals and plants. Microbiol. Mol. Biol. Rev. 62, 379–433 (1998).
Büttner, D. & Bonas, U. Getting across-bacterial type III effector proteins on their way to the plant cell. EMBO J. 21, 5313–5322 (2002).
Cornelis, G. R. The Yersinia Ysc–Yop virulence apparatus. Int. J. Med. Microbiol. 291, 455–462 (2002).
Galan, J. E. Salmonella interactions with host cells: type III secretion at work. Annu. Rev. Cell Dev. Biol. 17, 53–86 (2001).
Cascales, E. & Christie, P. J. The versatile bacterial type IV secretion systems. Nature Rev. Microbiol. 1, 137–149 (2003).
Capela, D. et al. Analysis of the chromosome sequence of the legume symbiont Sinorhizobium meliloti strain 1021. Proc. Natl Acad. Sci. USA 98, 9877–9882 (2001).
Barnett, M. J. et al. Nucleotide sequence and predicted functions of the entire Sinorhizobium meliloti pSymA megaplasmid. Proc. Natl Acad. Sci. USA 98, 9883–9888 (2001).
Galibert, F. et al. The composite genome of the legume symbiont Sinorhizobium meliloti. Science 293, 668–672 (2001).
Kaneko, T. et al. Complete genomic sequence of nitrogen-fixing symbiotic bacterium Bradyrhizobium japonicum USDA110. DNA Res. 9, 189–197 (2002).
Sullivan, J. T. & Ronson, C. W. Evolution of rhizobia by acquisition of a 500-kb symbiosis island that integrates into a phe-tRNA gene. Proc. Natl Acad. Sci. USA 95, 5145–5149 (1998).
Gonzalez, V. et al. The mosaic structure of the symbiotic plasmid of Rhizobium etli CFN42 and its relation to other symbiotic genome compartments. Genome Biol. 4, R36 (2003). Decribes the great variability of the genetic structure and gene content of symbiosis islands of rhizobia.
Kaneko, T. et al. Complete genome structure of the nitrogen-fixing symbiotic bacterium Mesorhizobium loti. DNA Res. 7, 331–338 (2000).
Freiberg, C. et al. Molecular basis of symbiosis between Rhizobium and legumes. Nature 387, 394–401 (1997).
Sullivan, J. T. et al. Comparative sequence analysis of the symbiosis island of Mesorhizobium loti strain R7A. J. Bacteriol. 184, 3086–3095 (2002).
Duchaud, E. et al. The genome sequence of the entomopathogenic bacterium Photorhabdus luminescens. Nature Biotechnol. 21, 1307–1313 (2003).
ffrench-Constant, R. H. et al. A genomic sample sequence of the entomopathogenic bacterium Photorhabdus luminescens W14: potential implications for virulence. Appl. Environ. Microbiol. 66, 3310–3329 (2000).
ffrench-Constant, R. et al. Photorhabdus: towards a functional genomic analysis of a symbiont and pathogen. FEMS Microbiol. Rev. 26, 433–456 (2003). First compilation of island-like DNA regions in the genome of Photorhabdus.
Zhang, R. & Zhang, C. T. Identification of genomic islands in the genome of Bacillus cereus by comparative analysis with Bacillus anthracis. Physiol. Genomics 16, 19–23 (2003).
Marie, C., Broughton, W. J. & Deakin, W. J. Rhizobium type III secretion systems: legume charmers or alarmers? Curr. Opin. Plant Biol. 4, 336–342 (2001).
Dale, C. & Welburn, S. C. The endosymbionts of tsetse flies: manipulating host-parasite interactions. Int. J. Parasitol. 31, 628–631 (2001).
Schubbe, S. et al. Characterization of a spontaneous nonmagnetic mutant of Magnetospirillum gryphiswaldense reveals a large deletion comprising a putative magnetosome island. J. Bacteriol. 185, 5779–5790 (2003).
Rajakumar, K. et al. Identification of a chromosomal Shigella flexneri multi-antibiotic resistance locus which shares sequence and organizational similarity with the resistance region of the plasmid NR1. Plasmid 37, 159–168 (1997).
Acknowledgements
We thank H. Merkert for excellent graphical assistance. Our own work related to the topic is supported by the Deutsche Forschungsgemeinschaft, the Bundesministerium für Bildung und Forschung, the Fonds der Chemischen Industrie and the Bavarian Research Foundation.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Related links
Related links
DATABASES
Entrez
Bradyrhizobium japonicum strain USDA110
Lactobacillus johnsonii strain NCC533
FURTHER INFORMATION
Pathogenic enterobacteria laboratory, University of Würzburg
Glossary
- SUBTRACTION HYBRIDIZATION
-
A technique that is used to specifically enrich the DNA species that are present in one sample but not in another by hybridizing the nucleic acids of the two samples and removing the associated double-stranded molecules.
- SYNTENY
-
The property of being located on the same chromosome. In bacteria, chromosomal regions with conserved synteny represent dna segments in which the gene order is conserved in different species.
- POLYKETIDE
-
A natural product that is assembled from malonyl CoA units through intermediates with many ketone groups (polyketonic) that allow for directed reactivity to product structures. Polyketides represent an important class of secondary metabolites that include many medicinal and antibiotic compounds.
- T-DNA
-
A DNA segment of the tumour-inducing (Ti) plasmid of A. tumefaciens. It is transferred into the nuclei of infected cells where it is stably integrated into the host genome and transcribed, causing crown gall disease. T-DNA contains two types of genes: oncogenes that encode enzymes involved in the synthesis of auxins and cytokinins, which are responsible for tumour formation; and genes coding for the synthesis of opines.
- OPINE FOOD SUBSTRATES
-
Compounds produced by condensation between amino acids and sugars. They are synthesized and excreted by crown-gall cells and consumed by A. tumefaciens as carbon and nitrogen sources. Opines are not easily catabolized by other bacterial species and allow A. tumefaciens to benefit from the production of plant opines.
- NOD FACTOR
-
Bacterially produced substituted lipooligosaccharides that affect plant development in a host-specific way. They switch on a developmental pathway within the plant that triggers early features of nodule organogenesis in the rhizobium–legume symbiosis.
- COLICINS
-
Polypeptide toxins produced by and active against Escherichia coli and closely related bacteria.
- METAGENOMICS
-
The genomic analysis of all microorganisms present in a specific habitat.
Rights and permissions
About this article
Cite this article
Dobrindt, U., Hochhut, B., Hentschel, U. et al. Genomic islands in pathogenic and environmental microorganisms. Nat Rev Microbiol 2, 414–424 (2004). https://doi.org/10.1038/nrmicro884
Issue Date:
DOI: https://doi.org/10.1038/nrmicro884
This article is cited by
-
Comparative genomics of two Vietnamese Helicobacter pylori strains, CHC155 from a non-cardia gastric cancer patient and VN1291 from a duodenal ulcer patient
Scientific Reports (2023)
-
Silent gene clusters encode magnetic organelle biosynthesis in a non-magnetotactic phototrophic bacterium
The ISME Journal (2023)