Abstract
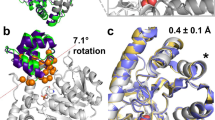
Traditional approaches for increasing the affinity of a protein for its ligand focus on constructing improved surface complementarity in the complex by altering the protein binding site to better fit the ligand. Here we present a novel strategy that leaves the binding site intact, while residues that allosterically affect binding are mutated. This method takes advantage of conformationally distinct states, each with different ligand-binding affinities, and manipulates the equilibria between these conformations. We demonstrate this approach in the Escherichia coli maltose binding protein by introducing mutations, located at some distance from the ligand binding pocket, that sterically affect the equilibrium between an open, apo-state and a closed, ligand-bound state. A family of 20 variants was generated with affinities ranging from a ∼100-fold improvement (7.4 nM) to a ∼two-fold weakening (1.8 mM) relative to the wild type protein (800 nM).
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Lowman, H.B., Bass, S.H., Simpson, N. & Wells, J.A. Biochemistry 30, 10832–10838 (1991).
Lowman, H.B. & Wells, J.A. J. Mol. Biol. 234, 564–578 (1993).
Short, M.K., Jeffrey, P.D., Kwong, R.F. & Margolies, M.N. J. Biol. Chem. 270, 28541–28550 (1995).
Saggio, I., Gloaguen, I., Poiana, G. & Laufer, R. EMBO J. 14, 3045–3054 (1995).
Loladze, V.V., Ibarra-Molero, B., Sanchez-Ruiz, J.M. & Makhatadze, G.I. Biochemistry 38, 16419–16423. (1999).
Gerstein, M., Lesk, A. & Chothia, C. Biochemistry 33, 6739–6749 (1994).
Tam, R. & Saier, M.H. Microbiol. Rev. 57, 320–346 (1993).
Quiocho, F.A., Spurlino, J.C. & Rodseth, L.E. Structure 5, 997–1015 (1997).
Sharff, A.J., Rodseth, L.E., Spurlino, J.C. & Quiocho, F.A. Biochemistry 31, 10657–10663 (1992).
Marvin, J.S. et al. Proc. Natl. Acad. Sci. USA 94, 4366–4371 (1997).
Wiseman, T., Williston, S., Brandts, J.F. & Lin, L.N. Anal. Biochem. 179, 131–137 (1989).
Miller, D.M., Olson, J.S., Pflugrath, J.W. & Quiocho, F.A. J. Biol. Chem. 258, 13665–13672 (1983).
Donner, J., Caruthers, M.H. & Gill, S.J. J. Biol. Chem. 257, 14826–14829 (1982).
Sturtevant, J.M. Proc. Natl. Acad. Sci. USA 74, 2236–2240 (1977).
Chervenak, M.C. & Toone, E.J. J. Am. Chem. Soc. 116, 10533–10539 (1994).
Spolar, R.S. & Record, M.T. Science 263, 777–784 (1994).
Johnson, K.A. In The enzymes, Vol 20. (ed. Boyer, P.D.)1–61 (Academic Press, Inc., San Diego; 1992).
Voss, E.W. J. Mol. Recogn. 6, 51–58 (1993).
Kelley, R.L. & Yanofsky, C. Proc. Natl. Acad. Sci. USA 82, 483–487 (1985).
Zhang, R.G. et al. Nature 327, 591–597 (1987).
Vermersch, P.S., Tesmer, J.J., Lemon, D.D. & Quiocho, F.A. A J. Biol. Chem. 265, 16592–16603 (1990).
Kundrot, C.E. & Evans, P.R. Biochemistry 30, 1478–1484 (1991).
Shimaoka, M. et al. Nature Struct. Biol. 7, 674–678 (2000).
Richardson, D.C. & Richardson, J.S. Protein Sci. 1, 3–9 (1992).
Shrake, A. & Rupley, J.A. J. Mol. Biol. 79, 351–371. (1973).
Hellinga, H.W. & Richards, F.M. J. Mol. Biol. 222, 763–785 (1991).
Thomson, J., Liu, Y., Sturtevant, J.M. & Quiocho, F.A.A. Biophys. Chem. 70, 101–108 (1998).
Acknowledgements
The authors would like to thank E.J. Toone for helpful discussions regarding the thermodynamics of maltose binding. This work was funded by grants from the National Institutes of Health and the Office of Naval Research.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Marvin, J., Hellinga, H. Manipulation of ligand binding affinity by exploitation of conformational coupling. Nat Struct Mol Biol 8, 795–798 (2001). https://doi.org/10.1038/nsb0901-795
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nsb0901-795
This article is cited by
-
Computational remodeling of an enzyme conformational landscape for altered substrate selectivity
Nature Communications (2023)
-
A Mathematical Model for the Kinetics of the MalFGK\(_2\) Maltose Transporter
Bulletin of Mathematical Biology (2020)
-
A genetically encoded fluorescent sensor for in vivo imaging of GABA
Nature Methods (2019)
-
Evolution of cyclohexadienyl dehydratase from an ancestral solute-binding protein
Nature Chemical Biology (2018)
-
Stability, affinity, and chromatic variants of the glutamate sensor iGluSnFR
Nature Methods (2018)