Abstract
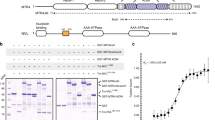
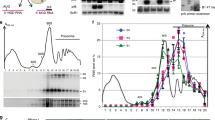
The eukaryotic exosome is a ten-subunit 3′ exoribonucleolytic complex responsible for many RNA-processing and RNA-degradation reactions. How the exosome accomplishes this is unknown. Rrp44 (also known as Dis3), a member of the RNase II family of enzymes, is the catalytic subunit of the exosome. We show that the PIN domain of Rrp44 has endoribonucleolytic activity. The PIN domain is preferentially active toward RNA with a 5′ phosphate, suggesting coordination of 5′ and 3′ processing. We also show that the endonuclease activity is important in vivo. Furthermore, the essential exosome subunit Csl4 does not contain any domains that are required for viability, but its zinc-ribbon domain is required for exosome-mediated mRNA decay. These results suggest that specific exosome domains contribute to specific functions, and that different RNAs probably interact with the exosome differently. The combination of an endoRNase and an exoRNase activity seems to be a widespread feature of RNA-degrading machines.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Moser, M.J., Holley, W.R., Chatterjee, A. & Mian, I.S. The proofreading domain of Escherichia coli DNA polymerase I and other DNA and/or RNA exonuclease domains. Nucleic Acids Res. 25, 5110–5118 (1997).
Mian, I.S. Comparative sequence analysis of ribonucleases HII, III, II PH and D. Nucleic Acids Res. 25, 3187–3195 (1997).
Deutscher, M.P. & Li, Z. Exoribonucleases and their multiple roles in RNA metabolism. Prog. Nucleic Acid Res. Mol. Biol. 66, 67–105 (2001).
Zuo, Y. & Deutscher, M.P. Exoribonuclease superfamilies: structural analysis and phylogenetic distribution. Nucleic Acids Res. 29, 1017–1026 (2001).
van Hoof, A., Lennertz, P. & Parker, R. Three conserved members of the RNase D family have unique and overlapping functions in the processing of 5S, 5.8S, U4, U5, RNase MRP and RNase P RNAs in yeast. EMBO J. 19, 1357–1365 (2000).
Muhlrad, D., Decker, C.J. & Parker, R. Turnover mechanisms of the stable yeast PGK1 mRNA. Mol. Cell. Biol. 15, 2145–2156 (1995).
Jacobs Anderson, J.S. & Parker, R. The 3′ to 5′ degradation of yeast mRNAs is a general mechanism for mRNA turnover that requires the SKI2 DEVH box protein and 3′ to 5′ exonucleases of the exosome complex. EMBO J. 17, 1497–1506 (1998).
van Hoof, A., Frischmeyer, P.A., Dietz, H.C. & Parker, R. Exosome-mediated recognition and degradation of mRNAs lacking a termination codon. Science 295, 2262–2264 (2002).
Meaux, S. & Van Hoof, A. Yeast transcripts cleaved by an internal ribozyme provide new insight into the role of the cap and poly(A) tail in translation and mRNA decay. RNA 12, 1323–1337 (2006).
Mitchell, P., Petfalski, E., Shevchenko, A., Mann, M. & Tollervey, D. The exosome: a conserved eukaryotic RNA processing complex containing multiple 3′ → 5′ exoribonucleases. Cell 91, 457–466 (1997).
Allmang, C. et al. The yeast exosome and human PM-Scl are related complexes of 3′ → 5′ exonucleases. Genes Dev. 13, 2148–2158 (1999).
Allmang, C. et al. Functions of the exosome in rRNA, snoRNA and snRNA synthesis. EMBO J. 18, 5399–5410 (1999).
van Hoof, A., Lennertz, P. & Parker, R. Yeast exosome mutants accumulate 3′-extended polyadenylated forms of U4 small nuclear RNA and small nucleolar RNAs. Mol. Cell. Biol. 20, 441–452 (2000).
de la Cruz, J., Kressler, D., Tollervey, D. & Linder, P. Dob1p (Mtr4p) is a putative ATP-dependent RNA helicase required for the 3′ end formation of 5.8S rRNA in Saccharomyces cerevisiae. EMBO J. 17, 1128–1140 (1998).
Liu, Q., Greimann, J.C. & Lima, C.D. Reconstitution, activities, and structure of the eukaryotic RNA exosome. Cell 127, 1223–1237 (2006).
Lorentzen, E. et al. The archaeal exosome core is a hexameric ring structure with three catalytic subunits. Nat. Struct. Mol. Biol. 12, 575–581 (2005).
Symmons, M.F., Jones, G.H. & Luisi, B.F. A duplicated fold is the structural basis for polynucleotide phosphorylase catalytic activity, processivity, and regulation. Structure 8, 1215–1226 (2000).
Navarro, M.V., Oliveira, C.C., Zanchin, N.I. & Guimaraes, B.G. Insights into the mechanism of progressive RNA degradation by the archaeal exosome. J. Biol. Chem. 283, 14120–14131 (2008).
Lorentzen, E. & Conti, E. Structural basis of 3′ end RNA recognition and exoribonucleolytic cleavage by an exosome RNase PH core. Mol. Cell 20, 473–481 (2005).
Buttner, K., Wenig, K. & Hopfner, K.P. Structural framework for the mechanism of archaeal exosomes in RNA processing. Mol. Cell 20, 461–471 (2005).
Dziembowski, A., Lorentzen, E., Conti, E. & Seraphin, B. A single subunit, Dis3, is essentially responsible for yeast exosome core activity. Nat. Struct. Mol. Biol. 14, 15–22 (2007).
Chekanova, J.A., Dutko, J.A., Mian, I.S. & Belostotsky, D.A. Arabidopsis thaliana exosome subunit AtRrp4p is a hydrolytic 3′ → 5′ exonuclease containing S1 and KH RNA-binding domains. Nucleic Acids Res. 30, 695–700 (2002).
Lorentzen, E., Basquin, J., Tomecki, R., Dziembowski, A. & Conti, E. Structure of the active subunit of the yeast exosome core, Rrp44: diverse modes of substrate recruitment in the RNase II nuclease family. Mol. Cell 29, 717–728 (2008).
Schneider, C., Anderson, J.T. & Tollervey, D. The exosome subunit Rrp44 plays a direct role in RNA substrate recognition. Mol. Cell 27, 324–331 (2007).
Wang, H.W. et al. Architecture of the yeast Rrp44 exosome complex suggests routes of RNA recruitment for 3′ end processing. Proc. Natl. Acad. Sci. USA 104, 16844–16849 (2007).
Frazão, C. et al. Unravelling the dynamics of RNA degradation by ribonuclease II and its RNA-bound complex. Nature 443, 110–114 (2006).
Barbas, A. et al. New insights into the mechanism of RNA degradation by ribonuclease II: identification of the residue responsible for setting the RNase II end product. J. Biol. Chem. 283, 13070–13076 (2008).
Arcus, V.L., Backbro, K., Roos, A., Daniel, E.L. & Baker, E.N. Distant structural homology leads to the functional characterization of an archaeal PIN domain as an exonuclease. J. Biol. Chem. 279, 16471–16478 (2004).
Levin, I. et al. Crystal structure of a PIN (PilT N-terminus) domain (AF0591) from Archaeoglobus fulgidus at 1.90 resolution. Proteins 56, 404–408 (2004).
Glavan, F., Behm-Ansmant, I., Izaurralde, E. & Conti, E. Structures of the PIN domains of SMG6 and SMG5 reveal a nuclease within the mRNA surveillance complex. EMBO J. 25, 5117–5125 (2006).
Daines, D.A., Wu, M.H. & Yuan, S.Y. VapC-1 of nontypeable Haemophilus influenzae is a ribonuclease. J. Bacteriol. 189, 5041–5048 (2007).
Bunker, R.D., McKenzie, J.L., Baker, E.N. & Arcus, V.L. Crystal structure of PAE0151 from Pyrobaculum aerophilum, a PIN-domain (VapC) protein from a toxin-antitoxin operon. Proteins 72, 510–518 (2008).
Eberle, A.B., Lykke-Andersen, S., Muhlemann, O. & Jensen, T.H. SMG6 promoted endonucleoytic cleavage of nonsense mRNA in human cells. Nat. Struct. Mol. Biol. advance online publication, doi:10.1038/nsmb.1530 (07 December 2008).
Fatica, A., Tollervey, D. & Dlakic, M. PIN domain of Nob1p is required for D-site cleavage in 20S pre-rRNA. RNA 10, 1698–1701 (2004).
Bleichert, F., Granneman, S., Osheim, Y.N., Beyer, A.L. & Baserga, S.J. The PINc domain protein Utp24, a putative nuclease, is required for the early cleavage steps in 18S rRNA maturation. Proc. Natl. Acad. Sci. USA 103, 9464–9469 (2006).
Amblar, M. & Arraiano, C.M. A single mutation in Escherichia coli ribonuclease II inactivates the enzyme without affecting RNA binding. FEBS J. 272, 363–374 (2005).
Amblar, M., Barbas, A., Fialho, A.M. & Arraiano, C.M. Characterization of the functional domains of Escherichia coli RNase II. J. Mol. Biol. 360, 921–933 (2006).
Barbas, A. et al. New insights into the mechanism of RNA degradation by ribonuclease II: identification of the residue responsible for setting the RNase II end product. J. Biol. Chem. 283, 13070–13076 (2008).
Cairrao, F., Arraiano, C. & Newbury, S. Drosophila gene tazman, an orthologue of the yeast exosome component Rrp44p/Dis3, is differentially expressed during development. Dev. Dyn. 232, 733–737 (2005).
Johnson, A.W. & Kolodner, R.D. Synthetic lethality of sep1 (xrn1) ski2 and sep1 (xrn1) ski3 mutants of Saccharomyces cerevisiae is independent of killer virus and suggests a general role for these genes in translation control. Mol. Cell. Biol. 15, 2719–2727 (1995).
Andrade, J.M., Pobre, V., Silva, I.J., Domingues, S. & Arraiano, C.M. The role of 3′ to 5′ exonucleases in RNA degradation. Prog. Nucleic Acid Res. Mol. Biol. (in the press).
Mackie, G.A. Ribonuclease E is a 5′-end-dependent endonuclease. Nature 395, 720–723 (1998).
Yang, W. An equivalent metal ion in one- and two-metal-ion catalysis. Nat. Struct. Mol. Biol. 15, 1228–1231 (2008).
Ross-Macdonald, P. et al. Large-scale analysis of the yeast genome by transposon tagging and gene disruption. Nature 402, 413–418 (1999).
Chekanova, J.A. et al. Genome-wide high-resolution mapping of exosome substrates reveals hidden features in the Arabidopsis transcriptome. Cell 131, 1340–1353 (2007).
Koonin, E.V., Wolf, Y.I. & Aravind, L. Prediction of the archaeal exosome and its connections with the proteasome and the translation and transcription machineries by a comparative-genomic approach. Genome Res. 11, 240–252 (2001).
Carpousis, A.J., Van Houwe, G., Ehretsmann, C. & Krisch, H.M. Copurification of E. coli RNAase E and PNPase: evidence for a specific association between two enzymes important in RNA processing and degradation. Cell 76, 889–900 (1994).
Py, B., Causton, H., Mudd, E.A. & Higgins, C.F. A protein complex mediating mRNA degradation in Escherichia coli. Mol. Microbiol. 14, 717–729 (1994).
Liu, J., Valencia-Sanchez, M.A., Hannon, G.J. & Parker, R. MicroRNA-dependent localization of targeted mRNAs to mammalian P-bodies. Nat. Cell Biol. 7, 719–723 (2005).
Ingelfinger, D., Arndt-Jovin, D.J., Luhrmann, R. & Achsel, T. The human LSm1–7 proteins colocalize with the mRNA-degrading enzymes Dcp1/2 and Xrnl in distinct cytoplasmic foci. RNA 8, 1489–1501 (2002).
Bashkirov, V.I., Scherthan, H., Solinger, J.A., Buerstedde, J.M. & Heyer, W.D. A mouse cytoplasmic exoribonuclease (mXRN1p) with preference for G4 tetraplex substrates. J. Cell Biol. 136, 761–773 (1997).
Cougot, N., Babajko, S. & Seraphin, B. Cytoplasmic foci are sites of mRNA decay in human cells. J. Cell Biol. 165, 31–40 (2004).
Zheng, D. et al. Deadenylation is prerequisite for P-body formation and mRNA decay in mammalian cells. J. Cell Biol. 182, 89–101 (2008).
Andrei, M.A. et al. A role for eIF4E and eIF4E-transporter in targeting mRNPs to mammalian processing bodies. RNA 11, 717–727 (2005).
Giaever, G. et al. Functional profiling of the Saccharomyces cerevisiae genome. Nature 418, 387–391 (2002).
Rockmill, B., Lambie, E.J. & Roeder, G.S. Spore enrichment. Methods Enzymol. 194, 146–149 (1991).
Sikorski, R.S. & Hieter, P. A system of shuttle vectors and yeast host strains designed for efficient manipulation of DNA in Saccharomyces cerevisiae. Genetics 122, 19–27 (1989).
Wilson, M.A., Meaux, S. & van Hoof, A. A genomic screen in yeast reveals novel aspects of nonstop mRNA metabolism. Genetics 177, 773–784 (2007).
van Hoof, A., Staples, R.R., Baker, R.E. & Parker, R. Function of the ski4p (Csl4p) and Ski7p proteins in 3′-to-5′ degradation of mRNA. Mol. Cell. Biol. 20, 8230–8243 (2000).
Lebreton, A., Tomecki, R., Dziembowski, A. & Séraphin, B. Endonucleolytic RNA cleavage by a eukaryotic exosome. Nature advance online publication, doi:10.1038/nature07480 (7 December 2008).
Acknowledgements
We are grateful to T. Link and R. Brennan for help with calculating the buried surfaces between the different domains of the cap proteins and the PH ring, and A. Klauer for technical assistance. GAL::rrp44, GAL::csl4, GAL::rrp4 and GAL::rrp40 strains were kindly provided by P. Mitchell (University of Sheffield) and D. Tollervey (University of Edinburgh). R. Parker, M. Wilkinson, M. Steiger and members of the van Hoof and Arraiano laboratories gave insightful comments on the manuscript. This research was supported by the Pew Scholarship Program in the Biomedical Sciences and by the National Institutes of Health (GM069900) to A.v.H. E.G.D and M.S.-R. were supported by The University of Texas at Houston Medical School-Summer Research Program. The work at the Instituto de Tecnologia Química e Biológica was supported by Fundação para a Ciência e a Tecnologia (FCT), Portugal. A.B. was a recipient of a post-doctoral fellowship from FCT, Portugal.
Author information
Authors and Affiliations
Corresponding author
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–7 and Supplementary Tables 1–3 (PDF 4057 kb)
Rights and permissions
About this article
Cite this article
Schaeffer, D., Tsanova, B., Barbas, A. et al. The exosome contains domains with specific endoribonuclease, exoribonuclease and cytoplasmic mRNA decay activities. Nat Struct Mol Biol 16, 56–62 (2009). https://doi.org/10.1038/nsmb.1528
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.1528
This article is cited by
-
DIS3L2 knockdown impairs key oncogenic properties of colorectal cancer cells via the mTOR signaling pathway
Cellular and Molecular Life Sciences (2023)
-
piRNAs Interact with Cold-Shock Domain-Containing RNA Binding Proteins and Regulate Neuronal Gene Expression During Differentiation
Molecular Neurobiology (2022)
-
A comprehensive insight into the biology of Rhizoctonia solani AG1-IA Kühn, the causal organism of the sheath blight disease of rice
Journal of Plant Pathology (2022)
-
Circular RNAs and their role in renal cell carcinoma: a current perspective
Cancer Cell International (2021)
-
LISTERIN E3 Ubiquitin Ligase and Ribosome-Associated Quality Control (RQC) Mechanism
Molecular Neurobiology (2021)