Abstract
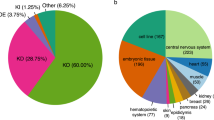
The elucidation of a code for regulated splicing has been a long-standing goal in understanding the control of post-transcriptional gene expression events that are crucial for cell survival, differentiation and development. We decoded functional RNA elements in vivo by constructing an RNA map for the cell type–specific splicing regulator FOX2 (also known as RBM9) via cross-linking immunoprecipitation coupled with high-throughput sequencing (CLIP-seq) in human embryonic stem cells. The map identified a large cohort of specific FOX2 targets, many of which are themselves splicing regulators, and comparison between the FOX2 binding profile and validated splicing events revealed a general rule for FOX2-regulated exon inclusion or skipping in a position-dependent manner. These findings suggest that FOX2 functions as a critical regulator of a splicing network, and we further show that FOX2 is important for the survival of human embryonic stem cells.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Black, D.L. Mechanisms of alternative pre-messenger RNA splicing. Annu. Rev. Biochem. 72, 291–336 (2003).
Thomson, J.A. et al. Embryonic stem cell lines derived from human blastocysts. Science 282, 1145–1147 (1998).
Keller, G. Embryonic stem cell differentiation: emergence of a new era in biology and medicine. Genes Dev. 19, 1129–1155 (2005).
Sonntag, K.C., Simantov, R. & Isacson, O. Stem cells may reshape the prospect of Parkinson's disease therapy. Brain Res. Mol. Brain Res. 134, 34–51 (2005).
Yeo, G.W. et al. Alternative splicing events identified in human embryonic stem cells and neural progenitors. PLOS Comput. Biol. 3, e196 (2007).
Jin, Y. et al. A vertebrate RNA-binding protein Fox-1 regulates tissue-specific splicing via the pentanucleotide GCAUG. EMBO J. 22, 905–912 (2003).
Underwood, J.G., Boutz, P.L., Dougherty, J.D., Stoilov, P. & Black, D.L. Homologues of the Caenorhabditis elegans Fox-1 protein are neuronal splicing regulators in mammals. Mol. Cell. Biol. 25, 10005–10016 (2005).
Ule, J. et al. CLIP identifies Nova-regulated RNA networks in the brain. Science 302, 1212–1215 (2003).
Robertson, G. et al. Genome-wide profiles of STAT1 DNA association using chromatin immunoprecipitation and massively parallel sequencing. Nat. Methods 4, 651–657 (2007).
Fairbrother, W.G., Yeh, R.F., Sharp, P.A. & Burge, C.B. Predictive identification of exonic splicing enhancers in human genes. Science 297, 1007–1013 (2002).
Auweter, S.D. et al. Molecular basis of RNA recognition by the human alternative splicing factor Fox-1. EMBO J. 25, 163–173 (2006).
Kabat, J.L. et al. Intronic alternative splicing regulators identified by comparative genomics in nematodes. PLOS Comput. Biol. 2, e86 (2006).
Goren, A. et al. Comparative analysis identifies exonic splicing regulatory sequences—the complex definition of enhancers and silencers. Mol. Cell 22, 769–781 (2006).
Yeo, G.W., Nostrand, E.L. & Liang, T.Y. Discovery and analysis of evolutionarily conserved intronic splicing regulatory elements. PLoS Genet. 3, e85 (2007).
Sorek, R. & Ast, G. Intronic sequences flanking alternatively spliced exons are conserved between human and mouse. Genome Res. 13, 1631–1637 (2003).
Yeo, G.W., Van Nostrand, E., Holste, D., Poggio, T. & Burge, C.B. Identification and analysis of alternative splicing events conserved in human and mouse. Proc. Natl. Acad. Sci. USA 102, 2850–2855 (2005).
Siepel, A. et al. Evolutionarily conserved elements in vertebrate, insect, worm, and yeast genomes. Genome Res. 15, 1034–1050 (2005).
Brudno, M. et al. Computational analysis of candidate intron regulatory elements for tissue-specific alternative pre-mRNA splicing. Nucleic Acids Res. 29, 2338–2348 (2001).
Baraniak, A.P., Chen, J.R. & Garcia-Blanco, M.A. Fox-2 mediates epithelial cell-specific fibroblast growth factor receptor 2 exon choice. Mol. Cell. Biol. 26, 1209–1222 (2006).
Makeyev, E.V., Zhang, J., Carrasco, M.A. & Maniatis, T. The microRNA miR-124 promotes neuronal differentiation by triggering brain-specific alternative pre-mRNA splicing. Mol. Cell 27, 435–448 (2007).
Boutz, P.L. et al. A post-transcriptional regulatory switch in polypyrimidine tract-binding proteins reprograms alternative splicing in developing neurons. Genes Dev. 21, 1636–1652 (2007).
Nakahata, S. & Kawamoto, S. Tissue-dependent isoforms of mammalian Fox-1 homologs are associated with tissue-specific splicing activities. Nucleic Acids Res. 33, 2078–2089 (2005).
Ule, J. et al. An RNA map predicting Nova-dependent splicing regulation. Nature 444, 580–586 (2006).
Uchida, N. et al. Direct isolation of human central nervous system stem cells. Proc. Natl. Acad. Sci. USA 97, 14720–14725 (2000).
Idziorek, T., Estaquier, J., De Bels, F. & Ameisen, J.C. YOPRO-1 permits cytofluorometric analysis of programmed cell death (apoptosis) without interfering with cell viability. J. Immunol. Methods 185, 249–258 (1995).
Halbeisen, R.E., Galgano, A., Scherrer, T. & Gerber, A.P. Post-transcriptional gene regulation: from genome-wide studies to principles. Cell. Mol. Life Sci. 65, 798–813 (2008).
Zhang, C. et al. Defining the regulatory network of the tissue-specific splicing factors Fox-1 and Fox-2. Genes Dev. 22, 2550–2563 (2008).
Ponthier, J.L. et al. Fox-2 splicing factor binds to a conserved intron motif to promote inclusion of protein 4.1R alternative exon 16. J. Biol. Chem. 281, 12468–12474 (2006).
Singer, O. et al. Targeting BACE1 with siRNAs ameliorates Alzheimer disease neuropathology in a transgenic model. Nat. Neurosci. 8, 1343–1349 (2005).
Crissman, H.A. & Steinkamp, J.A. Rapid, simultaneous measurement of DNA, protein, and cell volume in single cells from large mammalian cell populations. J. Cell Biol. 59, 766–771 (1973).
Krishan, A. Rapid flow cytofluorometric analysis of mammalian cell cycle by propidium iodide staining. J. Cell Biol. 66, 188–193 (1975).
Acknowledgements
The authors would like to acknowledge S. Aigner for technical advice, J. Simon for illustration assistance and R. Keithley and B. Miller for cell culture. G.W.Y. was funded by a Junior Fellowship from the Crick-Jacobs Center for Theoretical and Computational Biology, Salk Institute. F.H.G. is funded by the California Institute of Regenerative Medicine, The Picower Foundation and the Lookout Foundation. Part of this work was supported by US National Institutes of Health grants to X.-D.F. (GM049369 and HG004659) and G.W.Y. (HG004659).
Author information
Authors and Affiliations
Contributions
G.W.Y. directed the project; G.W.Y. and F.H.G. designed the project; G.W.Y., N.G.C. and X.-D.F. analyzed the data and wrote the manuscript; G.W.Y., N.G.C., T.Y.L. and G.E.P. performed the experiments; G.W.Y. and T.Y.L. carried out bioinformatics data analysis.
Corresponding authors
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–11, Supplementary Tables 1 and 2 and Supplementary Methods (PDF 5186 kb)
Rights and permissions
About this article
Cite this article
Yeo, G., Coufal, N., Liang, T. et al. An RNA code for the FOX2 splicing regulator revealed by mapping RNA-protein interactions in stem cells. Nat Struct Mol Biol 16, 130–137 (2009). https://doi.org/10.1038/nsmb.1545
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.1545
This article is cited by
-
RNA splicing analysis using heterogeneous and large RNA-seq datasets
Nature Communications (2023)
-
RBFOX2 deregulation promotes pancreatic cancer progression and metastasis through alternative splicing
Nature Communications (2023)
-
Nucleotide-amino acid π-stacking interactions initiate photo cross-linking in RNA-protein complexes
Nature Communications (2022)
-
Aberrant NOVA1 function disrupts alternative splicing in early stages of amyotrophic lateral sclerosis
Acta Neuropathologica (2022)
-
LncRNA MIAT Promotes Spinal Cord Injury Recovery in Rats by Regulating RBFOX2-Mediated Alternative Splicing of MCL-1
Molecular Neurobiology (2022)