Abstract
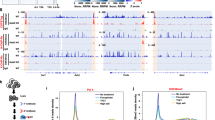
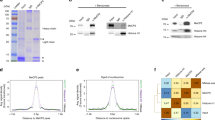
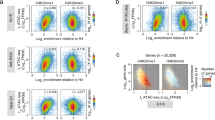
Dimethylation of histone H3 Arg2 (H3R2me2) maintains transcriptional silencing by inhibiting Set1 mediated trimethylation of H3K4. Here we demonstrate that Arg2 is also monomethylated (H3R2me1) in yeast but that its functional characteristics are distinct from H3R2me2: (i) H3R2me1 does not inhibit histone H3 Lys4 (H3K4) methylation; (ii) it is present throughout the coding region of genes; and (iii) it correlates with active transcription. Collectively, these results indicate that different H3R2 methylation states have defined roles in gene expression.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Accession codes
References
Kouzarides, T. Cell 128, 693–705 (2007).
Wysocka, J., Allis, C.D. & Coonrod, S. Front. Biosci. 11, 344–355 (2006).
Turner, B.M. Nat. Struct. Mol. Biol. 12, 110–112 (2005).
Bauer, U.M., Daujat, S., Nielsen, S.J., Nightingale, K. & Kouzarides, T. EMBO Rep. 3, 39–44 (2002).
Ma, H. et al. Curr. Biol. 11, 1981–1985 (2001).
Strahl, B.D. et al. Curr. Biol. 11, 996–1000 (2001).
Wang, H. et al. Science 293, 853–857 (2001).
Kirmizis, A. et al. Nature 449, 928–932 (2007).
Guccione, E. et al. Nature 449, 933–937 (2007).
Hyllus, D. et al. Genes Dev. 21, 3369–3380 (2007).
Iberg, A.N. et al. J. Biol. Chem. 283, 3006–3010 (2007).
Holstege, F.C. et al. Cell 95, 717–728 (1998).
Lakowski, T.M. & Frankel, A. J. Biol. Chem. 283, 10015–10025 (2008).
Ruthenburg, A.J., Allis, C.D. & Wysocka, J. Mol. Cell 25, 15–30 (2007).
Sims, R.J. III & Reinberg, D. Genes Dev. 20, 2779–2786 (2006).
Acknowledgements
We thank members of the T.K. laboratory for helpful discussions and M. Gilchrist for help with depositing genomic data. This work was supported by postdoctoral fellowship grants to A.K. from the European Molecular Biology Organization (EMBO) and Marie Curie. The T.K. laboratory is funded by grants from Cancer Research UK (CRUK) and the 6th Research Framework Program of the European Union (Epitron and Heroic).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
T.K. is a director of Abcam Plc. R.D.G. and M.A.S. are employees of NimbleGen Systems Inc.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1 and 2, Supplementary Table 1 and Supplementary Methods (PDF 1189 kb)
Rights and permissions
About this article
Cite this article
Kirmizis, A., Santos-Rosa, H., Penkett, C. et al. Distinct transcriptional outputs associated with mono- and dimethylated histone H3 arginine 2. Nat Struct Mol Biol 16, 449–451 (2009). https://doi.org/10.1038/nsmb.1569
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.1569
This article is cited by
-
Autophagy dictates sensitivity to PRMT5 inhibitor in breast cancer
Scientific Reports (2023)
-
Protein arginine methylation: a prominent modification and its demethylation
Cellular and Molecular Life Sciences (2017)
-
Readers of histone modifications
Cell Research (2011)
-
LSD1-mediated demethylation of histone H3 lysine 4 triggers Myc-induced transcription
Oncogene (2010)
-
Histone methyltransferases: regulation of transcription and contribution to human disease
Journal of Molecular Medicine (2010)