Abstract
Myotonic dystrophy is an RNA gain-of-function disease caused by expanded CUG or CCUG repeats, which sequester the RNA binding protein MBNL1. Here we describe a newly discovered function for MBNL1 as a regulator of pre-miR-1 biogenesis and find that miR-1 processing is altered in heart samples from people with myotonic dystrophy. MBNL1 binds to a UGC motif located within the loop of pre-miR-1 and competes for the binding of LIN28, which promotes pre-miR-1 uridylation by ZCCHC11 (TUT4) and blocks Dicer processing. As a consequence of miR-1 loss, expression of GJA1 (connexin 43) and CACNA1C (Cav1.2), which are targets of miR-1, is increased in both DM1- and DM2-affected hearts. CACNA1C and GJA1 encode the main calcium- and gap-junction channels in heart, respectively, and we propose that their misregulation may contribute to the cardiac dysfunctions observed in affected persons.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Brook, J.D. et al. Molecular basis of myotonic dystrophy: expansion of a trinucleotide (CTG) repeat at the 3′ end of a transcript encoding a protein kinase family member. Cell 68, 799–808 (1992).
Fu, Y.H. et al. An unstable triplet repeat in a gene related to myotonic muscular dystrophy. Science 255, 1256–1258 (1992).
Mahadevan, M. et al. Myotonic dystrophy mutation: an unstable CTG repeat in the 3′ untranslated region of the gene. Science 255, 1253–1255 (1992).
Liquori, C.L. et al. Myotonic dystrophy type 2 caused by a CCTG expansion in intron 1 of ZNF9. Science 293, 864–867 (2001).
Miller, J.W. et al. Recruitment of human muscleblind proteins to (CUG)n expansions associated with myotonic dystrophy. EMBO J. 19, 4439–4448 (2000).
Kanadia, R.N. et al. A muscleblind knockout model for myotonic dystrophy. Science 302, 1978–1980 (2003).
Ho, T.H. et al. Muscleblind proteins regulate alternative splicing. EMBO J. 23, 3103–3112 (2004).
Kanadia, R.N. et al. Reversal of RNA missplicing and myotonia after muscleblind overexpression in a mouse poly(CUG) model for myotonic dystrophy. Proc. Natl. Acad. Sci. USA 103, 11748–11753 (2006).
Philips, A.V., Timchenko, L.T. & Cooper, T.A. Disruption of splicing regulated by a CUG-binding protein in myotonic dystrophy. Science 280, 737–741 (1998).
Kuyumcu-Martinez, N.M. et al. Increased steady-state levels of CUGBP1 in myotonic dystrophy 1 are due to PKC-mediated hyperphosphorylation. Mol. Cell 28, 68–78 (2007).
Mankodi, A. et al. Myotonic dystrophy in transgenic mice expressing an expanded CUG repeat. Science 289, 1769–1773 (2000).
Lin, X. et al. Failure of MBNL1-dependent post-natal splicing transitions in myotonic dystrophy. Hum. Mol. Genet. 15, 2087–2097 (2006).
Kalsotra, A. et al. A postnatal switch of CELF and MBNL proteins reprograms alternative splicing in the developing heart. Proc. Natl. Acad. Sci. USA 105, 20333–20338 (2008).
Savkur, R.S. et al. Aberrant regulation of insulin receptor alternative splicing is associated with insulin resistance in myotonic dystrophy. Nat. Genet. 29, 40–47 (2001).
Mankodi, A. et al. Expanded CUG repeats trigger aberrant splicing of ClC-1 chloride channel pre-mRNA and hyperexcitability of skeletal muscle in myotonic dystrophy. Mol. Cell 10, 35–44 (2002).
Charlet-B., N. et al. Loss of the muscle-specific chloride channel in type 1 myotonic dystrophy due to misregulated alternative splicing. Mol. Cell 10, 45–53 (2002).
Wheeler, T.M. et al. Correction of ClC-1 splicing eliminates chloride channelopathy and myotonia in mouse models of myotonic dystrophy. J. Clin. Invest. 117, 3952–3957 (2007).
Groh, W.J. et al. Electrocardiographic abnormalities and sudden death in myotonic dystrophy type 1. N. Engl. J. Med. 358, 2688–2697 (2008).
Pelargonio, G. et al. Myotonic dystrophy and the heart. Heart 88, 665–670 (2002).
Phillips, M.F. & Harper, P.S. Cardiac disease in myotonic dystrophy. Cardiovasc. Res. 33, 13–22 (1997).
Lazarus, A. et al. Long-term follow-up of arrhythmias in patients with myotonic dystrophy treated by pacing: a multicenter diagnostic pacemaker study. J. Am. Coll. Cardiol. 40, 1645–1652 (2002).
Wang, G.S. et al. Elevation of RNA-binding protein CUGBP1 is an early event in an inducible heart-specific mouse model of myotonic dystrophy. J. Clin. Invest. 117, 2802–2811 (2007).
Mahadevan, M.S. et al. Reversible model of RNA toxicity and cardiac conduction defects in myotonic dystrophy. Nat. Genet. 38, 1066–1070 (2006).
Sayed, D. et al. MicroRNAs play an essential role in the development of cardiac hypertrophy. Circ. Res. 100, 416–424 (2007).
Carè, A. et al. MicroRNA-133 controls cardiac hypertrophy. Nat. Med. 13, 613–618 (2007).
Yang, B. et al. The muscle-specific microRNA miR-1 regulates cardiac arrhythmogenic potential by targeting GJA1 and KCNJ2. Nat. Med. 13, 486–491 (2007).
Zhao, Y. et al. Dysregulation of cardiogenesis, cardiac conduction, and cell cycle in mice lacking miRNA-1–2. Cell 129, 303–317 (2007).
Schoser, B.G. et al. Sudden cardiac death in myotonic dystrophy type 2. Neurology 63, 2402–2404 (2004).
Lee, Y. et al. The nuclear RNase III Drosha initiates microRNA processing. Nature 425, 415–419 (2003).
Denli, A.M. et al. Processing of primary microRNAs by the Microprocessor complex. Nature 432, 231–235 (2004).
Han, J. et al. The Drosha-DGCR8 complex in primary microRNA processing. Genes Dev. 18, 3016–3027 (2004).
Lund, E. et al. Nuclear export of microRNA precursors. Science 303, 95–98 (2004).
Yi, R. et al. Exportin-5 mediates the nuclear export of pre-microRNAs and short hairpin RNAs. Genes Dev. 17, 3011–3016 (2003).
Zhang, H. et al. Single processing center models for human Dicer and bacterial RNase III. Cell 118, 57–68 (2004).
Chendrimada, T.P. et al. TRBP recruits the Dicer complex to Ago2 for microRNA processing and gene silencing. Nature 436, 740–744 (2005).
Goers, E.S. et al. MBNL1 binds GC motifs embedded in pyrimidines to regulate alternative splicing. Nucleic Acids Res. 38, 2467–2484 (2010).
Yuan, Y. et al. Muscleblind-like 1 interacts with RNA hairpins in splicing target and pathogenic RNAs. Nucleic Acids Res. 35, 5474–5486 (2007).
Heo, I. et al. Lin28 mediates the terminal uridylation of let-7 precursor MicroRNA. Mol. Cell 32, 276–284 (2008).
Rybak, A. et al. A feedback loop comprising lin-28 and let-7 controls pre-let-7 maturation during neural stem-cell commitment. Nat. Cell Biol. 10, 987–993 (2008).
Hagan, J.P. et al. Lin28 recruits the TUTase Zcchc11 to inhibit let-7 maturation in mouse embryonic stem cells. Nat. Struct. Mol. Biol. 16, 1021–1025 (2009).
Heo, I. et al. TUT4 in concert with Lin28 suppresses microRNA biogenesis through pre-microRNA uridylation. Cell 138, 696–708 (2009).
Yang, D.H. & Moss, E.G. Temporally regulated expression of Lin-28 in diverse tissues of the developing mouse. Gene Expr. Patterns 3, 719–726 (2003).
Kalsotra, A. et al. A postnatal switch of CELF and MBNL proteins reprograms alternative splicing in the developing heart. Proc. Natl. Acad. Sci. USA 105, 20333–20338 (2008).
Ikeda, S. et al. MicroRNA-1 negatively regulates expression of the hypertrophy-associated calmodulin and Mef2a genes. Mol. Cell. Biol. 8, 2193–2204 (2009).
Terentyev, D. et al. miR-1 overexpression enhances Ca2+ release and promotes cardiac arrhythmogenesis by targeting PP2A regulatory subunit B56α and causing CaMKII-dependent hyperphosphorylation of RyR2. Circ. Res. 104, 514–521 (2009).
Elia, L. et al. Reciprocal regulation of microRNA-1 and insulin-like growth factor-1 signal transduction cascade in cardiac and skeletal muscle in physiological and pathological conditions. Circulation 120, 2377–2385 (2009).
Li, Q. et al. Attenuation of microRNA-1 derepresses the cytoskeleton regulatory protein twinfilin-1 to provoke cardiac hypertrophy. J. Cell Sci. 123, 2444–2452 (2010).
Splawski, I. et al. CaV1.2 calcium channel dysfunction causes a multisystem disorder including arrhythmia and autism. Cell 119, 19–31 (2004).
Hirai, H. et al. MyoD regulates apoptosis of myoblasts through microRNA-mediated down-regulation of Pax3. J. Cell Biol. 191, 347–365 (2010).
Chen, J.F. et al. microRNA-1 and microRNA-206 regulate skeletal muscle satellite cell proliferation and differentiation by repressing Pax7. J. Cell Biol. 190, 867–879 (2010).
Cacchiarelli, D. et al. MicroRNAs involved in molecular circuitries relevant for the Duchenne muscular dystrophy pathogenesis are controlled by the dystrophin / nNOS pathway. Cell Metab. 12, 341–351 (2010).
Chen, J.F. et al. The role of microRNA-1 and microRNA-133 in skeletal muscle proliferation and differentiation. Nat. Genet. 38, 228–233 (2006).
Yan, D. et al. MicroRNA-1/206 targets c-Met and inhibits rhabdomyosarcoma development. J. Biol. Chem. 284, 29596–29604 (2009).
Furling, D., Lemieux, D., Taneja, K. & Puymirat, J. Decreased levels of myotonic dystrophy protein kinase (DMPK) and delayed differentiation in human myotonic dystrophy myoblasts. Neuromuscul. Disord. 8, 728–735 (2001).
Furling, D. et al. Defective satellite cells in congenital myotonic dystrophy. Hum. Mol. Genet. 10, 2079–2087 (2001).
Guil, S. & Cáceres, J.F. The multifunctional RNA-binding protein hnRNP A1 is required for processing of miR-18a. Nat. Struct. Mol. Biol. 14, 591–596 (2007).
Davis, B.N. et al. Smad proteins bind a conserved RNA sequence to promote microRNA maturation by Drosha. Mol. Cell 39, 373–384 (2010).
Trabucchi, M. et al. The RNA-binding protein KSRP promotes the biogenesis of a subset of microRNAs. Nature 459, 1010–1014 (2009).
Yang, W. et al. Modulation of microRNA processing and expression through RNA editing by ADAR deaminases. Nat. Struct. Mol. Biol. 13, 13–21 (2006).
Nakamori, M. et al. Aberrantly spliced α-dystrobrevin alters α-syntrophin binding in myotonic dystrophy type 1. Neurology 70, 677–685 (2008).
Acknowledgements
We thank T. Cooper (Baylor College of Medicine) for the gift of the DT960 and tgCUGBP1 plasmids, C. Branlant (National Center of Scientific Research) for the pGEX-MBNL1-Δ101 (MBNL1ΔCter) vector, M. Swanson (University of Florida) for the pGEX-6P-MBNL1-His vector, C. Thornton (University of Rochester) for the polyclonal antibody against MBNL1, N. Kim (University of Seoul) for the pri-miR-16, Flag-TUT4, His-LIN28 and Flag-LIN28 plasmids, P. Provost (Université Laval) for the gift of the Myc-Dicer plasmid and all members of the French Myotonic Dystrophy Network for fruitful discussion. This work was supported by Institut National de la Santé et de la Recherche Médicale (INSERM) AVENIR (N.C.-B.), Agence nationale de la recherche (ANR) GENOPAT P007942 (N.C.B.), Association Française contre les Myopathies (AFM) MNM1-12982 (N.C.B.), the UPMC-emergence program (D.F), US National Institutes of Health (NIH) P30 AR057220 (J.W.D.), Muscular Dystrophy Center Core Laboratories (J.W.D.), the Ministère de l'enseignement supérieur et de la recherche and AFM (F.R., F.F.) and the University of Strasbourg (F.R., F.F.).
Author information
Authors and Affiliations
Contributions
Experiments were conducted by F.R., F.F., C.F., J.-P.V., D.D., N.D., M.-C.F., A.N., D.A. and B.J. Clinical samples and patient data were obtained from J.W.D., D.D., K.W., D.F., G.G., H.F., D.D., M.P.T. and from the Research Resource Network supported by the Research Grant for Nervous and Mental Disorders from the Ministry of Health, Labour and Welfare, Japan. The study was designed and coordinated by N.D., D.F. and N.C.-B. The paper was written by N.C.-B.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–3, Supplementary Table 1 and Supplementary Methods (PDF 979 kb)
Supplementary Data
Microarray data. (a) Analysis using the Agilent GeneSpring GX Software of miRNA microarray performed on RNA extracted from three independent cultures of primary muscle cultures (differentiated 10 days) derived from controls and age-matched DM1 patients. Among the various miRNA with a mis-regulation of fold change >2 only, miR-1, miR-126, miR-138, miR-146, miR-199 and miR-499 had a corrected p value < 0.05. (b) Raw data and crude analysis using excel. Data are accessible through NCBI GEO GSE24109. (XLS 3656 kb)
Rights and permissions
About this article
Cite this article
Rau, F., Freyermuth, F., Fugier, C. et al. Misregulation of miR-1 processing is associated with heart defects in myotonic dystrophy. Nat Struct Mol Biol 18, 840–845 (2011). https://doi.org/10.1038/nsmb.2067
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.2067
This article is cited by
-
RNA modification in cardiovascular disease: implications for therapeutic interventions
Signal Transduction and Targeted Therapy (2023)
-
MiR-199-3p enhances muscle regeneration and ameliorates aged muscle and muscular dystrophy
Communications Biology (2021)
-
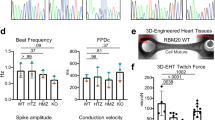
iPSC-derived cardiomyocytes from patients with myotonic dystrophy type 1 have abnormal ion channel functions and slower conduction velocities
Scientific Reports (2021)
-
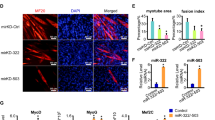
miR-322/-503 rescues myoblast defects in myotonic dystrophy type 1 cell model by targeting CUG repeats
Cell Death & Disease (2020)
-
Myocardial fibrosis by late gadolinium enhancement cardiovascular magnetic resonance in myotonic muscular dystrophy type 1: highly prevalent but not associated with surface conduction abnormality
Journal of Cardiovascular Magnetic Resonance (2019)