Abstract
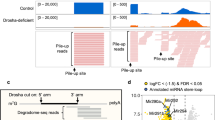
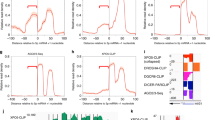
The Drosha–DGCR8 complex (Microprocessor) is required for microRNA (miRNA) biogenesis. DGCR8 recognizes the RNA substrate, whereas Drosha functions as the endonuclease. Using high-throughput sequencing and cross-linking immunoprecipitation (HITS-CLIP) we identified RNA targets of DGCR8 in human cells. Unexpectedly, miRNAs were not the most abundant targets. DGCR8-bound RNAs also comprised several hundred mRNAs as well as small nucleolar RNAs (snoRNAs) and long noncoding RNAs. We found that the Microprocessor controlled the abundance of several mRNAs as well as of MALAT1. By contrast, DGCR8-mediated cleavage of snoRNAs was independent of Drosha, suggesting the involvement of DGCR8 in cellular complexes with other endonucleases. Binding of DGCR8 to cassette exons is a new mechanism for regulation of the relative abundance of alternatively spliced isoforms. These data provide insights in the complex role of DGCR8 in controlling the fate of several classes of RNAs.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Accession codes
References
Bartel, D.P. MicroRNAs: target recognition and regulatory functions. Cell 136, 215–233 (2009).
Han, J. et al. The Drosha-DGCR8 complex in primary microRNA processing. Genes Dev. 18, 3016–3027 (2004).
Zeng, Y., Yi, R. & Cullen, B.R. Recognition and cleavage of primary microRNA precursors by the nuclear processing enzyme Drosha. EMBO J. 24, 138–148 (2005).
Morlando, M. et al. Primary microRNA transcripts are processed co-transcriptionally. Nat. Struct. Mol. Biol. 15, 902–909 (2008).
Pawlicki, J.M. & Steitz, J.A. Primary microRNA transcript retention at sites of transcription leads to enhanced microRNA production. J. Cell Biol. 182, 61–76 (2008).
Yi, R., Qin, Y., Macara, I.G. & Cullen, B.R. Exportin-5 mediates the nuclear export of pre-microRNAs and short hairpin RNAs. Genes Dev. 17, 3011–3016 (2003).
Lund, E., Guttinger, S., Calado, A., Dahlberg, J.E. & Kutay, U. Nuclear export of microRNA precursors. Science 303, 95–98 (2004).
Bohnsack, M.T., Czaplinski, K. & Gorlich, D. Exportin 5 is a RanGTP-dependent dsRNA-binding protein that mediates nuclear export of pre-miRNAs. RNA 10, 185–191 (2004).
Bernstein, E., Caudy, A.A., Hammond, S.M. & Hannon, G.J. Role for a bidentate ribonuclease in the initiation step of RNA interference. Nature 409, 363–366 (2001).
Hutvagner, G. et al. A cellular function for the RNA-interference enzyme Dicer in the maturation of the let-7 small temporal RNA. Science 293, 834–838 (2001).
Shiohama, A., Sasaki, T., Noda, S., Minoshima, S. & Shimizu, N. Molecular cloning and expression analysis of a novel gene DGCR8 located in the DiGeorge syndrome chromosomal region. Biochem. Biophys. Res. Commun. 304, 184–190 (2003).
Landthaler, M., Yalcin, A. & Tuschl, T. The human DiGeorge syndrome critical region gene 8 and Its D. melanogaster homolog are required for miRNA biogenesis. Curr. Biol. 14, 2162–2167 (2004).
Denli, A.M., Tops, B.B., Plasterk, R.H., Ketting, R.F. & Hannon, G.J. Processing of primary microRNAs by the Microprocessor complex. Nature 432, 231–235 (2004).
Gregory, R.I. et al. The Microprocessor complex mediates the genesis of microRNAs. Nature 432, 235–240 (2004).
Davis, B.N. & Hata, A. Regulation of microRNA biogenesis: a miRiad of mechanisms. Cell Commun. Signal. 7, 18 (2009).
Winter, J., Jung, S., Keller, S., Gregory, R.I. & Diederichs, S. Many roads to maturity: microRNA biogenesis pathways and their regulation. Nat. Cell Biol. 11, 228–234 (2009).
Krol, J., Loedige, I. & Filipowicz, W. The widespread regulation of microRNA biogenesis, function and decay. Nat. Rev. Genet. 11, 597–610 (2010).
Han, J. et al. Molecular basis for the recognition of primary microRNAs by the Drosha-DGCR8 complex. Cell 125, 887–901 (2006).
Faller, M., Matsunaga, M., Yin, S., Loo, J.A. & Guo, F. Heme is involved in microRNA processing. Nat. Struct. Mol. Biol. 14, 23–29 (2007).
Faller, M. et al. DGCR8 recognizes primary transcripts of microRNAs through highly cooperative binding and formation of higher-order structures. RNA 16, 1570–1583 (2010).
Licatalosi, D.D. et al. HITS-CLIP yields genome-wide insights into brain alternative RNA processing. Nature 456, 464–469 (2008).
Sanford, J.R. et al. Splicing factor SFRS1 recognizes a functionally diverse landscape of RNA transcripts. Genome Res. 19, 381–394 (2009).
Chi, S.W., Zang, J.B., Mele, A. & Darnell, R.B. Argonaute HITS-CLIP decodes microRNA-mRNA interaction maps. Nature 460, 479–486 (2009).
Zisoulis, D.G. et al. Comprehensive discovery of endogenous Argonaute binding sites in Caenorhabditis elegans. Nat. Struct. Mol. Biol. 17, 173–179 (2010).
Konig, J. et al. iCLIP reveals the function of hnRNP particles in splicing at individual nucleotide resolution. Nat. Struct. Mol. Biol. 17, 909–915 (2010).
Hafner, M. et al. Transcriptome-wide identification of RNA-binding protein and microRNA target sites by PAR-CLIP. Cell 141, 129–141 (2010).
Han, J. et al. Posttranscriptional crossregulation between Drosha and DGCR8. Cell 136, 75–84 (2009).
Kadener, S. et al. Genome-wide identification of targets of the drosha-pasha/DGCR8 complex. RNA 15, 537–545 (2009).
Triboulet, R., Chang, H.M., LaPierre, R.J. & Gregory, R.I. Post-transcriptional control of DGCR8 expression by the Microprocessor. RNA 15, 1005–1011 (2009).
Ule, J., Jensen, K., Mele, A. & Darnell, R.B. CLIP: A method for identifying protein-RNA interaction sites in living cells. Methods 37, 376–386 (2005).
Licatalosi, D.D. & Darnell, R.B. RNA processing and its regulation: global insights into biological networks. Nat. Rev. Genet. 11, 75–87 (2010).
Karginov, F.V. et al. Diverse endonucleolytic cleavage sites in the mammalian transcriptome depend upon microRNAs, Drosha, and additional nucleases. Mol. Cell 38, 781–788 (2010).
Ji, P. et al. MALAT-1, a novel noncoding RNA, and thymosin beta4 predict metastasis and survival in early-stage non-small cell lung cancer. Oncogene 22, 8031–8041 (2003).
Tollervey, D. & Kiss, T. Function and synthesis of small nucleolar RNAs. Curr. Opin. Cell Biol. 9, 337–342 (1997).
Kiss, T. SnoRNP biogenesis meets pre-mRNA splicing. Mol. Cell 23, 775–776 (2006).
Taft, R.J. et al. Small RNAs derived from snoRNAs. RNA 15, 1233–1240 (2009).
Ender, C. et al. A human snoRNA with microRNA-like functions. Mol. Cell 32, 519–528 (2008).
Bernstein, E. et al. Dicer is essential for mouse development. Nat. Genet. 35, 215–217 (2003).
Wang, Y., Medvid, R., Melton, C., Jaenisch, R. & Blelloch, R. DGCR8 is essential for microRNA biogenesis and silencing of embryonic stem cell self-renewal. Nat. Genet. 39, 380–385 (2007).
Shenoy, A. & Blelloch, R. Genomic analysis suggests that mRNA destabilization by the microprocessor is specialized for the auto-regulation of Dgcr8. PLoS ONE 4, e6971 (2009).
Chong, M.M. et al. Canonical and alternate functions of the microRNA biogenesis machinery. Genes Dev. 24, 1951–1960 (2010).
Lin, Y.T. & Sullivan, C.S. Expanding the role of Drosha to the regulation of viral gene expression. Proc. Natl. Acad. Sci. USA 108, 11229–11234 (2011).
Wu, H., Xu, H., Miraaglia, L.J. & Crooke, S.T. Human RNase III is a 160-kDa protein involved in preribosomal RNA processing. J. Biol. Chem. 275, 36957–36965 (2000).
Liang, T.J. & Qin, C.Y. The emerging role of microRNAs in immune cell development and differentiation. APMIS 117, 635–643 (2009).
Scott, M.S., Avolio, F., Ono, M., Lamond, A.I. & Barton, G.J. Human miRNA precursors with box H/ACA snoRNA features. PLoS Comput. Biol. 5, e1000507 (2009).
Shiohama, A., Sasaki, T., Noda, S., Minoshima, S. & Shimizu, N. Nucleolar localization of DGCR8 and identification of eleven DGCR8-associated proteins. Exp. Cell Res. 313, 4196–4207 (2007).
Stark, K.L. et al. Altered brain microRNA biogenesis contributes to phenotypic deficits in a 22q11-deletion mouse model. Nat. Genet. 40, 751–760 (2008).
Fenelon, K. et al. Deficiency of Dgcr8, a gene disrupted by the 22q11.2 microdeletion, results in altered short-term plasticity in the prefrontal cortex. Proc. Natl. Acad. Sci. USA 108, 4447–4452 (2011).
Michlewski, G. & Caceres, J.F. Antagonistic role of hnRNP A1 and KSRP in the regulation of let-7a biogenesis. Nat. Struct. Mol. Biol. 17, 1011–1018 (2010).
Caceres, J.F., Misteli, T., Screaton, G.R., Spector, D.L. & Krainer, A.R. Role of the modular domains of SR proteins in subnuclear localization and alternative splicing specificity. J. Cell Biol. 138, 225–238 (1997).
Guil, S. & Caceres, J.F. The multifunctional RNA-binding protein hnRNP A1 is required for processing of miR-18a. Nat. Struct. Mol. Biol. 14, 591–596 (2007).
Flicek, P. et al. Ensembl's 10th year. Nucleic Acids Res. 38, D557–D562 (2010).
Slater, G.S. & Birney, E. Automated generation of heuristics for biological sequence comparison. BMC Bioinformatics 6, 31 (2005).
Fujita, P.A. et al. The UCSC Genome Browser database: update 2011. Nucleic Acids Res. 39, D876–D882 (2011).
Acknowledgements
We are grateful to S. Heras for discussions and critical reading of the manuscript; N. Kim (Seoul National University) for Flag-Drosha, dominant negative DGCR8 and Drosha expression vectors; B. Seraphin (Institut de Génétique et de Biologie Moléculaire et Cellulaire, Strasbourg) for the Dcp1 antibody and R. Blelloch (University of California, San Francisco) for Dicer knockout and Dicer flox/flox cell lines. This work was supported by the Medical Research Council and by the Wellcome Trust (084057/Z/07/Z to J.F.C.; A.S., G.M. and S.M. were partially funded by this). E.E. and M.P. were supported by grants from the Spanish Ministry of Science and by the Sandra Ibarra Foundation (BIO2008-01091, BIO2011-23920 and CSD2009-00080). S.M. was the recipient of a European Molecular Biology Organization long-term postdoctoral fellowship. M.P. is supported by the Novo Nordisk Foundation. J.F.C is a recipient of a Wellcome Trust Senior Investigator Award (grant 095518/Z/11/Z).
Author information
Authors and Affiliations
Contributions
S.M. and J.F.C. conceived, designed and interpreted the experiments. S.M., G.M. and A.S. performed experiments and analyzed data. M.P. and E.E. performed all bioinformatics analysis, including mapping of the CLIP tags to the genome and statistical analysis. J.F.C. supervised the project. All authors wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–5, Supplementary Tables 1,3 and Supplementary Note (PDF 3023 kb)
Supplementary Table 2
List of alternative splicing changes in Dgcr8 knockout mouse embryonic stem cells. (XLS 1187 kb)
Rights and permissions
About this article
Cite this article
Macias, S., Plass, M., Stajuda, A. et al. DGCR8 HITS-CLIP reveals novel functions for the Microprocessor. Nat Struct Mol Biol 19, 760–766 (2012). https://doi.org/10.1038/nsmb.2344
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.2344
This article is cited by
-
LncRNA DANCR represses Doxorubicin-induced apoptosis through stabilizing MALAT1 expression in colorectal cancer cells
Cell Death & Disease (2021)
-
Complement C3 activation regulates the production of tRNA-derived fragments Gly-tRFs and promotes alcohol-induced liver injury and steatosis
Cell Research (2019)
-
Dgcr8 knockout approaches to understand microRNA functions in vitro and in vivo
Cellular and Molecular Life Sciences (2019)
-
Deletion of DGCR8 in VSMCs of adult mice results in loss of vascular reactivity, reduced blood pressure and neointima formation
Scientific Reports (2018)
-
RNA fate determination through cotranscriptional adenosine methylation and microprocessor binding
Nature Structural & Molecular Biology (2017)