Abstract
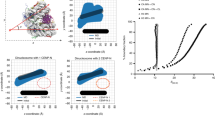
Post-translational histone modifications regulate epigenetic switching between different chromatin states. Distinct histone modifications, such as acetylation, methylation and phosphorylation, define different functional chromatin domains, and often do so in a combinatorial fashion. The centromere is a unique chromosomal locus that mediates multiple segregation functions, including kinetochore formation, spindle-mediated movements, sister cohesion and a mitotic checkpoint. Centromeric (CEN) chromatin is embedded in heterochromatin and contains blocks of histone H3 nucleosomes interspersed with blocks of CENP-A nucleosomes, the histone H3 variant that provides a structural and functional foundation for the kinetochore. Here, we demonstrate that the spectrum of histone modifications present in human and Drosophila melanogaster CEN chromatin is distinct from that of both euchromatin and flanking heterochromatin. We speculate that this distinct modification pattern contributes to the unique domain organization and three-dimensional structure of centromeric regions, and/or to the epigenetic information that determines centromere identity.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
Fischle, W., Wang, Y., Allis, C.D. Binary switches and modification cassettes in histone biology and beyond. Nature 425, 475–479 (2003).
Jenuwein, T. & Allis, C.D. Translating the histone code. Science 293, 1074–1080 (2001).
Bernstein, B.E. et al. Methylation of histone H3 Lys 4 in coding regions of active genes. Proc. Natl. Acad. Sci. USA 99, 8695–8700 (2002).
Santos-Rosa, H. et al. Active genes are tri-methylated at K4 of histone H3. Nature 419, 407–411 (2002).
Schneider, R. et al. Histone H3 Lys 4 methylation patterns in higher eukaryotic genes. Nat. Cell Biol. 6, 73–77 (2004).
Peters, A.H.F.M. et al. Partitioning and plasticity of repressive histone methylation states in mammalian chromatin. Mol. Cell 12, 1577–1589 (2003).
Rice, J.C. et al. Histone methyltransferases direct different degrees of methylation to define distinct chromatin domains. Mol. Cell 12, 1591–1599 (2003).
Nakayama, J.-I., Rice, J.C., Strahl, B.D., Allis, C.D. & Grewal, S.I.S. Role of histone H3 lysine 9 methylation in epigenetic control of heterochromatin assembly. Science 292, 110–113 (2001).
Schotta, G. et al. Central role of Drosophila SU(VAR)3-9 in histone H3-K9 methylation and heterochromatic gene silencing. EMBO J. 21, 1121–1131 (2002).
Muller, H.J. Types of visible variations induced by X-rays in Drosophila. J. Genet. 22, 299–334 (1930).
Allshire, R.C., Javerzat, J.P., Redhead, N.J. & Cranston, G. Position effect variegation at fission yeast centromeres. Cell 76, 157–169 (1994).
Sullivan, B.A., Blower, M.D. & Karpen, G.H. Determining centromere identity: cyclical stories and forking paths. Nat. Rev. Genet. 2, 584–596 (2001).
Cleveland, D.W., Mao, Y. & Sullivan, K.F. Centromere and kinetochores: from epigenetics to mitotic checkpoint signaling. Cell 112, 407–421 (2003).
Yoda, K. et al. Human centromere protein A (CENP-A) can replace histone H3 in nucleosomes reconstitution in vitro. Proc. Natl. Acad. Sci. USA 97, 7266–7271 (2000).
Blower, M.D., Sullivan, B.A. & Karpen, G.H. Conserved organization of centromeric chromatin in flies and humans. Dev. Cell 2, 319–330 (2002).
Mellone, B.G. & Allshire, R.C. Stretching it: putting the CEN(P-A) in centromere. Curr. Opin. Genet. Dev. 13, 191–198 (2003).
Nagaki, K. et al. Sequencing of a rice centromere uncovers active genes. Nat. Genet. 36, 138–145 (2004).
Partridge, J.F., Borgstrøm, B. & Allshire, R.C. Distinct protein interaction domains and protein spreading in a complex centromere. Genes Dev. 14, 783–791 (2000).
Blower, M.D. & Karpen, G.H. The role of Drosophila CID in kinetochore formation, cell-cycle progression and heterochromatin interactions. Nat. Cell Biol. 3, 730–739 (2001).
Taddei, A., Maison, C., Roche, D. & Almouzni, G. Reversible disruption of pericentric heterochromatin and centromere function by inhibiting deacetylases. Nat. Cell Biol. 3, 114–120 (2001).
O'Neill, L.P. & Turner, B.M. Histone H4 acetylation distinguishes coding regions of the human genome from heterochromatin in a differentiation-dependent but transcription-independent manner. EMBO J. 14, 3946–3957 (1995).
Boggs, B.A. et al. Differentially methylated forms of histone H3 show unique associations patterns with inactive human X chromosomes. Nat. Genet. 30, 73–76 (2002).
Maggert, K.A. & Karpen, G.H. The activation of a neocentromere in Drosophila requires proximity to an endogenous centromere. Genetics 158, 1615–1628 (2001).
Hoskins, R.A. et al. Heterochromatin sequences in a Drosophila whole-genome shotgun assembly. Genome Biol. 3, 0085.1–0085.16 (2002).
Mitchell, A.R., Jeppesen, P., Nicol, L., Morrision, H. & Kipling, D. Epigenetic control of mammalian centromere proteins binding: does DNA methylation have a role? J. Cell Sci. 109, 2199–2206 (1996).
Mitchell, A.R., Nicol, L., Malloy, P. & Kipling, D. Novel structural organization of a Mus musculus DBA/2 chromosome shows a fixed position for the centromere. J. Cell Sci. 106, 79–85 (1993).
Warburton, P.E. et al. Immunolocalization of CENP-A suggests a distinct nucleosomes structure at the inner kinetochore plate of active centromeres. Curr. Biol. 7, 901–904 (1997).
Ahmad, K. & Henikoff, S. Centromeres are specialized replication domains in heterochromatin. J. Cell Biol. 153, 101–110 (2001).
Sullivan, B. & Karpen, G. Centromere identity in Drosophila is not determined in vivo by replication timing. J. Cell Biol. 154, 683–690 (2001).
Kim, S.M., Dubey, D.D. & Huberman, J.A. Early-replicating heterochromatin. Genes Dev. 17, 330–335 (2003).
Tanaka, T., Cosma, M.P., Wirth, K. & Nasmyth, K. Identification of cohesin association sites at centromeres and along chromosome arms. Cell 98, 847–858 (1999).
Bernard, P. et al. Requirement of heterochromatin for cohesion at centromeres. Science 294, 2539–2542 (2001).
Ahmad, K. & Henikoff, S. The histone variant H3.3 marks active chromatin by replication-independent nucleosome assembly. Mol. Cell 9, 1191–1200 (2002).
Shelby, R.D., Monier, K. & Sullivan, K.F. Chromatin assembly at kinetochores is uncoupled from DNA replication. J. Cell Biol. 151, 1113–1118 (2000).
Ando, S., Yang, H., Nozaki, N., Okazaki, T. & Yoda, K. CENP-A, -B, and -C chromatin complex that contains the I-type alpha-satellite array constitutes the prekinetochore in HeLa cells. Mol. Cell. Biol. 22, 2229–2241 (2002).
Uhlmann, F. The mechanism of sister chromatid cohesion. Exp. Cell Res. 296, 80–85 (2004).
Acknowledgements
We thank A. Skora for technical assistance, and K. Scott, C. Vaziri, A. Dernburg, D. Allis, S. Erhardt and C. Yan for discussions and advice. We acknowledge K. Maggert for originally coining the term “centro-chromatin.” We are indebted to T. Jenuwein and A. Peters for use of their unique H3 Lys9-triMe antibodies. This work was supported by the following grants: US Department of Energy LDRD 366987 and US National Institutes of Health (NIH) R01 GM66272 (G.H.K.), and American Cancer Society IRG-72-001-29 and NIH R01 GM069514 (B.A.S.).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Sullivan, B., Karpen, G. Centromeric chromatin exhibits a histone modification pattern that is distinct from both euchromatin and heterochromatin. Nat Struct Mol Biol 11, 1076–1083 (2004). https://doi.org/10.1038/nsmb845
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb845
This article is cited by
-
CENP-A and CENP-B collaborate to create an open centromeric chromatin state
Nature Communications (2023)
-
CENP-N promotes the compaction of centromeric chromatin
Nature Structural & Molecular Biology (2022)
-
Human centromere repositioning activates transcription and opens chromatin fibre structure
Nature Communications (2022)
-
Incorporation of CENP-A/CID into centromeres during early Drosophila embryogenesis does not require RNA polymerase II–mediated transcription
Chromosoma (2022)
-
Mammalian SWI/SNF chromatin remodeler is essential for reductional meiosis in males
Nature Communications (2021)