Abstract
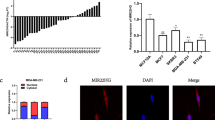
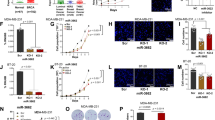
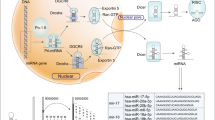
The tumor suppressor p53, encoded by the TP53 gene, is recognized as the guardian of the human genome because it regulates many downstream genes to exercise its function in cell cycle and cell death. Recent studies have revealed that several microRNAs (miRNAs) are important components of the p53 tumor suppressor network with miR-125b and miR-504 directly targeting TP53. In this study, we use a screening method to identify that two miRNAs (miR-25 and miR-30d) directly target the 3′UTR of TP53 to downregulate p53 protein levels and reduce the expression of genes that are transcriptionally activated by p53. Correspondingly, both miR-25 and miR-30d adversely affect apoptotic cell death, cell cycle arrest and cellular senescence. Inhibition of either miR-25 or miR-30d expression increases endogenous p53 expression and elevates cellular apoptosis in several cell lines, including one from multiple myeloma that has little TP53 mutations. Thus, beyond miR-125b and miR-504, the human TP53 gene is negatively regulated by two more miRNAs: miR-25 and miR-30d.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Bartel DP . (2004). MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 116: 281–297.
Bartel DP . (2009). MicroRNAs: target recognition and regulatory functions. Cell 136: 215–233.
Bommer GT, Gerin I, Feng Y, Kaczorowski AJ, Kuick R, Love RE et al. (2007). p53-mediated activation of miRNA34 candidate tumor-suppressor genes. Curr Biol 17: 1298–1307.
Bunz F, Dutriaux A, Lengauer C, Waldman T, Zhou S, Brown JP et al. (1998). Requirement for p53 and p21 to sustain G2 arrest after DNA damage. Science 282: 1497–1501.
Chang TC, Wentzel EA, Kent OA, Ramachandran K, Mullendore M, Lee KH et al. (2007). Transactivation of miR-34a by p53 broadly influences gene expression and promotes apoptosis. Mol Cell 26: 745–752.
Chng WJ, Price-Troska T, Gonzalez-Paz N, Van Wier S, Jacobus S, Blood E et al. (2007). Clinical significance of TP53 mutation in myeloma. Leukemia 21: 582–584.
Fonseca R, Barlogie B, Bataille R, Bastard C, Bergsagel PL, Chesi M et al. (2004). Genetics and cytogenetics of multiple myeloma: a workshop report. Cancer Res 64: 1546–1558.
Griffiths-Jones S, Grocock RJ, van Dongen S, Bateman A, Enright AJ . (2006). miRBase: microRNA sequences, targets and gene nomenclature. Nucleic Acids Res 34: D140–144.
Hanahan D, Weinberg RA . (2000). The Hallmarks of Cancer. Cell 100: 57–70.
Harris SL, Levine AJ . (2005). The p53 pathway: positive and negative feedback loops. Oncogene 24: 2899–2908.
He L, He X, Lim LP, de Stanchina E, Xuan Z, Liang Y et al. (2007). A microRNA component of the p53 tumour suppressor network. Nature 447: 1130–1134.
Hu W, Chan CS, Wu R, Zhang C, Sun Y, Song JS et al. (2010). Negative regulation of tumor suppressor p53 by microRNA miR-504. Mol Cell 38: 689–699.
John B, Enright AJ, Aravin A, Tuschl T, Sander C, Marks DS . (2004). Human microRNA targets. PLoS Biol 2: e363.
Johnson SM, Grosshans H, Shingara J, Byrom M, Jarvis R, Cheng A et al. (2005). RAS is regulated by the let-7 microRNA family. Cell 120: 635–647.
Kan T, Sato F, Ito T, Matsumura N, David S, Cheng Y et al. (2009). The miR-106b-25 polycistron, activated by genomic amplification, functions as an oncogene by suppressing p21 and Bim. Gastroenterology 136: 1689–1700.
Krek A, Grun D, Poy MN, Wolf R, Rosenberg L, Epstein EJ et al. (2005). Combinatorial microRNA target predictions. Nat Genet 37: 495.
Kruse J-P, Gu W . (2009). Modes of p53 Regulation. Cell 137: 609–622.
Le MTN, Teh C, Shyh-Chang N, Xie H, Zhou B, Korzh V et al. (2009). MicroRNA-125b is a novel negative regulator of p53. Genes & Dev 23: 862–876.
Lewis BP, Burge CB, Bartel DP . (2005). Conserved seed pairing, often flanked by adenosines, indicates that thousands of human genes are microRNA targets. Cell 120: 15–20.
Mayer B, Oberbauer R . (2003). Mitochondrial regulation of apoptosis. News Physiol Sci 18: 89–94.
Miranda KC, Huynh T, Tay Y, Ang YS, Tam WL, Thomson AM et al. (2006). A pattern-based method for the identification of microRNA binding sites and their corresponding heteroduplexes. Cell 126: 1203–1217.
Park SY, Lee JH, Ha M, Nam JW, Kim VN . (2009). miR-29 miRNAs activate p53 by targeting p85α and CDC42. Nat Struct Mol Biol 16: 23–29.
Petrocca F, Visone R, Onelli MR, Shah MH, Nicoloso MS, de Martino I et al. (2008). E2F1-regulated microRNAs impair TGFβ-dependent cell-cycle arrest and apoptosis in gastric cancer. Cancer Cell 13: 272–286.
Pichiorri F, Suh SS, Ladetto M, Kuehl M, Palumbo T, Drandi D et al. (2008). MicroRNAs regulate critical genes associated with multiple myeloma pathogenesis. Proc Natl Acad Sci USA 105: 12885–12890.
Prendergast NJ, Atkins MR, Schatte EC, Paulson DF, Walther PJ . (1996). p53 immunohistochemical and genetic alterations are associated at high incidence with post-irradiated locally persistent prostate carcinoma. J Urol 155: 1685–1692.
Raver-Shapira N, Marciano E, Meiri E, Spector Y, Rosenfeld N, Moskovits N et al. (2007). Transcriptional activation of miR-34a contributes to p53-mediated apoptosis. Mol Cell 26: 731–743.
Riley T, Sontag E, Chen P, Levine A . (2008). Transcriptional control of human p53-regulated genes. Nat Rev Mol Cell Biol 9: 402–412.
Suzuki S, Adachi A, Hiraiwa A, Ohashi M, Ishibashi M, Kiyono T . (1998). Cloning and characterization of human MCM7 promoter. Gene 216: 85–91.
Tarasov V, Jung P, Verdoodt B, Lodygin D, Epanchintsev A, Menssen A et al. (2007). Differential regulation of microRNAs by p53 revealed by massively parallel sequencing: miR-34a is a p53 target that induces apoptosis and G1-arrest. Cell Cycle 6: 1586–1593.
Toledo F, Wahl GM . (2006). Regulating the p53 pathway: in vitro hypotheses, in vivo veritas. Nat Rev Cancer 6: 909–923.
Ventura A, Kirsch DG, McLaughlin ME, Tuveson DA, Grimm J, Lintault L et al. (2007). Restoration of p53 function leads to tumour regression in vivo. Nature 445: 661–665.
Voeller HJ, Sugars LY, Pretlow T, Gelmann EP . (1994). p53 oncogene mutations in human prostate cancer specimens. J Urol 151: 492–495.
Vogelstein B, Lane D, Levine AJ . (2000). Surfing the p53 network. Nature 408: 307–310.
Voorhoeve PM, le Sage C, Schrier M, Gillis AJ, Stoop H, Nagel R et al. (2006). A genetic screen implicates miRNA-372 and miRNA-373 as oncogenes in testicular germ cell tumors. Cell 124: 1169–1181.
Vousden KH, Prives C . (2009). Blinded by the light: the growing complexity of p53. Cell 137: 413–431.
Wang XW, Zhan Q, Coursen JD, Khan MA, Kontny HU, Yu L et al. (1999). GADD45 induction of a G2/M cell cycle checkpoint. Proc Natl Acad Sci USA 96: 3706–3711.
Xiong W, Wu X, Starnes S, Johnson SK, Haessler J, Wang S et al. (2008). An analysis of the clinical and biologic significance of TP53 loss and the identification of potential novel transcriptional targets of TP53 in multiple myeloma. Blood 112: 4235–4246.
Xue W, Zender L, Miething C, Dickins RA, Hernando E, Krizhanovsky V et al. (2007). Senescence and tumour clearance is triggered by p53 restoration in murine liver carcinomas. Nature 445: 656–660.
Yanaihara N, Caplen N, Bowman E, Seike M, Kumamoto K, Yi M et al. (2006). Unique microRNA molecular profiles in lung cancer diagnosis and prognosis. Cancer Cell 9: 189–198.
Acknowledgements
Y L is supported by the Director's Biomarker Award from the Center for Environmental Genomics and Integrative Biology, funded by NIEHS/NIH (P30ES014443), Kentucky Lung Cancer Research Program, NCI/NIH (R01CA138688) and NCRR/NIH (P20 RR024489). K H Y is supported by the University of Wisconsin Paul Carbone Comprehensive Cancer Center Award, the Gundersen Medical Foundation Award, the University of Wisconsin Pathology R & D Fund and by NCI/NIH (P20CA103697 and 1RC1CA146299). We are grateful to the critical reviews from Drs Korise E Rasmusson and Nancy C Martin.Authorship contributions: MK, ZL, AT and WC performed research; NC and KY contributed human specimens; ZL, MK, KR, KY and YL designed research, analyzed data and wrote the paper; and all authors reviewed the paper.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on the Oncogene website
Supplementary information
Rights and permissions
About this article
Cite this article
Kumar, M., Lu, Z., Takwi, A. et al. Negative regulation of the tumor suppressor p53 gene by microRNAs. Oncogene 30, 843–853 (2011). https://doi.org/10.1038/onc.2010.457
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/onc.2010.457
Keywords
This article is cited by
-
miRNA-seq identification and clinical validation of CD138+ and circulating miR-25 in treatment response of multiple myeloma
Journal of Translational Medicine (2023)
-
Genome wide identification and characterization of fertility associated novel CircRNAs as ceRNA reveal their regulatory roles in sheep fecundity
Journal of Ovarian Research (2023)
-
Mesenchymal stem cells ameliorate cisplatin-induced acute kidney injury via let-7b-5p
Cell and Tissue Research (2023)
-
SUMOylation inhibition enhances dexamethasone sensitivity in multiple myeloma
Journal of Experimental & Clinical Cancer Research (2022)
-
Zinc Gluconate Induces Potentially Cancer Chemopreventive Activity in Barrett’s Esophagus: A Phase 1 Pilot Study
Digestive Diseases and Sciences (2021)