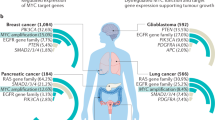
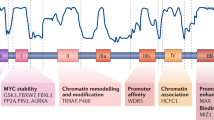
Abstract
The tyrosine kinase c-Abl (or Abl) and the prolyl-isomerase Pin1 cooperatively activate the transcription factor p73 by enhancing recruitment of the acetyltransferase p300. As the transcription factor c-Myc (or Myc) is a known target of Pin1 and p300, we hypothesized that it might be regulated in a similar manner. Consistent with this hypothesis, overexpression of Pin1 augmented the interaction of Myc with p300 and transcriptional activity. The action of Abl, however, was more complex than predicted. On one hand, Abl indirectly enhanced phosphorylation of Myc on Ser 62 and Thr 58, its association with Pin1 and p300 and its acetylation by p300. These effects of Abl were exerted through phosphorylation of substrate(s) other than Myc itself. On the other hand, Abl interacted with the C-terminal domain of Myc and phosphorylated up to five tyrosine residues in its N-terminus, the principal of which was Y74. Indirect immunofluorescence or immunohistochemical staining suggested that the Y74-phosphorylated form of Myc (Myc-pY74) localized to the cytoplasm and coexisted either with active Abl in a subset of mammary carcinomas or with Bcr-Abl in chronic myeloid leukemia. In all instances, Myc-pY74 constituted a minor fraction of the cellular Myc protein. Thus, our data unravel two potential effects of Abl on Myc: first, Abl signaling can indirectly augment acetylation of Myc by p300, and most likely also its transcriptional activity in the nucleus; second, Abl can directly phosphorylate Myc on tyrosine: the resulting form of Myc appears to be cytoplasmic, and its presence correlates with Abl activation in cancer.
Similar content being viewed by others
Introduction
The prolyl-isomerase Pin1 selectively isomerizes Proline residues immediately preceded by phosphorylated serine or threonine, generating conformational changes that modulate the activities of its substrates.1 Pin1 positively regulates a variety of transcription factors, including p53, p73, c-Jun and c-Fos.2, 3, 4, 5 These transcription factors recruit a diversity of cofactors, including the histone acetyltransferase p300.2, 3, 6 Pin1 has been shown to enhance recruitment of p300 by p53 and p73, leading to augmented acetylation of the transcription factors themselves.2, 3 In the case of p73, this effect of Pin1 is modulated by the tyrosine kinase c-Abl (hereafter Abl); in response to DNA damage, Abl directly phosphorylated p73 on tyrosine and, through activation of the p38 mitogen-activated protein kinase pathway, also favored phosphorylation of p73 on serine and threonine. This led to a chain of events including enhanced Pin1 binding, p300 recruitment and acetylation and stabilization of p73, culminating in enhanced transcriptional activity.3
Myc is a target of Pin1,7 as well as of p300,8, 9, 10, 11 but opposite effects have been reported. Pin1 targets phospho-T58-P59 in Myc, enhancing recruitment of protein phosphatase 2A, thereby facilitating dephosphorylation of the adjacent phospho-S62-P63 site and subsequent degradation of Myc by the ubiquitin–proteasome pathway.7 On the other hand, p300 stabilizes Myc, and does so independently from its histone acetyltransferase activity.9 Like other histone acetyltransferases,12 p300 also acetylates Myc8, 9, 11 and augments its transcriptional activity. Myc can also recruit various histone acetyltransferase complexes to chromatin, thereby enhancing histone acetylation and transcription.11, 13, 14
None of the above studies addressed whether Pin1 and p300 might act in concert to regulate Myc activity. Furthermore, by analogy with the regulation of p73, it remains to be investigated whether Myc may also be targeted by Abl, and whether Pin1 and Abl may synergize to augment p300 recruitment (Figure 1a). In this study, we addressed the concerted action of Pin1, p300 and Abl on Myc. Our data reveal a more complex regulation of Myc than initially hypothesized (Figure 1b); whereas overexpression of Pin1 and p300 augments Myc acetylation and transcriptional activity, Abl acts indirectly in this setting, via the enhancement of Myc T58/S62 phosphorylation. On the other hand, direct phosphorylation of Myc on several tyrosines (in particular Y74), appears to give rise to a cytoplasmic form of Myc, the action of which remains to be unraveled.
Schematic summary. (a) Working hypothesis: based on the concerted action of Abl and Ser/Thr Kinases on the function of p73, we tested whether a similar scheme might apply to Myc. (b) Model: the dual action of Abl (or Bcr-Abl) on Myc, based on the data presented in this work. The horizontal arrows depict relative intensities of transcription of Myc/Max-target genes. The positive effect of Abl on Myc transcriptional activity is not directly demonstrated by our data, but is inferred here, based on (1) the enhancement of Myc transcriptional activity by Pin1 and p300 and (2) the positive action of Abl on p300 binding and Myc acetylation (Figure 2). See text for further detail.
Results
Abl indirectly stimulates Myc phosphorylation on S62/T58, binding to Pin1 and p300, and acetylation
Transient transfection with a Luciferase reporter driven by the Myc-responsive Nucleolin promoter (Ncl-Luc)15 showed that Pin1 and p300 additively enhanced the transcriptional activity of Myc (Figure 2a). Abl could not be tested in this assay, because it deregulated the activity of all reporters used (whether Ncl-Luc, other Luc reporters or the Renilla Luciferase used as a normalizer). Hence, we addressed whether Abl and Pin1 might influence acetylation of Myc by p300; 293T cells were transfected with vectors expressing Myc, Pin1, p300 or Abl, each with the indicated epitope tags. Immunoblot analysis of Flag-Myc immunoprecipitates with an anti-acetyl-lysine antibody confirmed that p300 acetylates Myc;9, 12 coexpression of Pin1 and Abl together with p300 further increased Myc acetylation, whereas adding only Abl or Pin1 had no consistent effect (Figures 2b and c). This cooperative effect was lost when using catalytically inactive mutants of either Pin1 (Y23A; Figure 2b)16 or Abl (kinase-defective mutant (KD); Figure 2d),17 or when inhibiting Abl activity with STI571 (Figure 2e). Co-immunoprecipitation analysis of the transfected cells showed that Pin1 and Abl markedly enhanced binding of p300 to Myc (Figure 2c). In the above experiments, Myc was substantially acetylated when coexpressed with p300 alone, which might conceivably have depended upon endogenous Pin1 in 293T cells. In mouse embryo fibroblasts homozygous for a Pin1 knockout allele,3 coexpression of p300 also caused Myc acetylation, showing that Pin1 is not obligatory for this event; however, Myc acetylation was now markedly increased by coexpression of Pin1 and Abl (Figure 2f). Altogether, our data demonstrate that Pin1 and Abl cooperate to enhance the interaction of Myc with p300 and its resulting acetylation.
Abl enhances Myc transcriptional activity through Pin1 and p300. (a) U2OS cells were transfected with the reporter plasmid pNuc-Luc either alone or together with expression plasmids for Flag-Myc, HA-Pin1 or HA-p300, as indicated. The transcriptional activity of Myc was assayed measuring the activity of the luciferase gene under the control of the nucleolin promoter. pRL-TK (8 ng) were co-transfected to allow normalization for transfection efficiency. The histograms represent the means and s.d. of three independent experiments. The total levels of the indicated proteins were assessed by immunoblot. Asterisk: background band cross-reacting with the anti-Flag antibody. (b–e) 293T cells were transfected with plasmids expressing the indicated proteins (top). Lysates were immunoprecipitated (IP) with anti-Flag beads, and the precipitates subsequently analyzed by immunoblotting with the indicated antibodies. Total: direct immunoblot analysis of cell lysates without IP. Ac-Lys: anti acetyl-lysine. STI571: cells were treated with 1 μM STI571 for 2h prior to lysis. (f) Pin1 KO MEFs were transfected with plasmids expressing the indicated proteins and processed as in (b).
The association of Pin1 with Myc depends upon phosphorylation of Myc on T58 by glycogen synthase kinase 3, which in turn depends upon a priming phosphorylation event at S62.7, 18 Immunoblotting with phospho-specific antibodies revealed that Abl significantly enhanced phosphorylation of wild-type (WT) Myc on both S62 and T58 in co-transfected cells (Figure 3a). Mutation of T58 and/or S62 to alanine (T58A, S62A) confirmed the specificity of the antibodies used, as well as the dependency of T58 phosphorylation upon prior S62 phosphorylation. Consistent with the increase in T58 phosphorylation, Abl also enhanced the interaction between Pin1 and Myc, as assayed by either pull-down with a recombinant GST-Pin1 protein (Figures 3a and b) or co-immunoprecipitation (Figure 3c). As expected, mutation of Myc S62, T58 or both significantly reduced (although did not abrogate) the interaction with Pin1 (Figure 3a), and Myc did not co-precipitate with the Pin1Y23A mutant, which shows impaired binding to Pin1 substrates (Figure 3c).16
Pin1 binds P-T58 and P-S62 enhancing Myc acetylation independently of tyrosine phosphorylation. 293T cells were transfected with plasmids encoding the indicated proteins (top) in wild-type or mutant form (see text). Lysates were analyzed by GST-Pin1 pull-down (a, b) or immunoprecipitation (c, d) as indicated, followed by immunoblotting with the indicated antibodies. pT58, pS62 and pY74: antibodies directed against the corresponding phospho-residues in Myc. P-Tyr: anti phospho-Tyrosine antibody (4G10). IP, Total, Ac-Lys: as in Figures 2b-e. Long and short indicate relative times of exposure to film of the same blot.
In the above experiments, coexpression of Abl with Myc caused phosphorylation of Myc on tyrosine, as revealed by immunoblot analysis of Myc immunoprecipitates with antibodies raised either against generic phospho-tyrosine (Figures 2d and 3b and d) or against phospho-tyrosine 74 on Myc (pY74, Figure 3a). As will be described in detail below, the major phospho-tyrosines on Myc were Y74 and Y32, whereas five substitutions altogether (mutant Myc-5YF) abrogated phosphorylation of Myc by Abl. Most relevant here, Abl still enhanced T58/S62 phosphorylation in the Myc Y32F/Y74F mutant (Figure 3a), as well as acetylation of the Myc-5YF mutant in concert with Pin1 and p300 (Figure 3d), implying that these effects did not require direct phosphorylation of Myc by Abl.
Altogether, the data presented so far showed that Abl triggers a series of effects on Myc, including enhanced T58/S62 phosphorylation, Pin1 and p300 binding and acetylation, resulting most likely in elevated Myc transcriptional activity (Figure 1b). These effects are indirect in the sense that they do not depend upon direct phosphorylation of Myc by Abl, and must therefore involve the regulation of additional signaling pathways (Discussion). Most importantly, however, our data revealed that Abl also phosphorylates Myc on tyrosine, a new connection that we decided to characterize in the remainder of this work.
Abl directly binds and phosphorylates Myc
Coexpression of Abl and Flag-Myc, followed by anti-Flag immunoprecipitation and immunoblotting with an anti-phospho-tyrosine antibody confirmed tyrosine phosphorylation on Myc (Figure 4a and Supplementary Figure S1a). This was lost either with the inactive Abl KD mutant or upon treatment of cells with the Abl inhibitor STI571 (Supplementary Figures S1a and b). In the same experiments, Abl was co-immunoprecipitated with Flag-Myc (Figure 4a). To map the region of Myc targeted by Abl, we used a series of Flag- or hemagglutinin (HA)-tagged Myc deletion mutants (Figure 4b). Notably, only full-length Myc was both bound and phosphorylated by Abl; Myc proteins lacking the C-terminal region (mutants B, C and D) were unable to co-immunoprecipitate Abl, but were still phosphorylated on tyrosine upon co-transfection with Abl, although with lower efficiency than full-length Myc (Figure 4c, black arrowheads). Conversely, proteins lacking the N-terminal domain (mutants A, E and F) associated with Abl, but contained no detectable phospho-tyrosine (Figures 4c and d white arrowheads). Thus, as summarized in Figure 4b, Abl bound the C-terminus of Myc (as defined by mutant A), but phosphorylated residues within its N-terminal 110 residues (deleted in mutant E). In human Myc, this region included six tyrosines (Y12, 16, 22, 24, 32 and 74; Figure 4e).
Abl interacts with and phosphorylates Myc. (a) 293T cells were transfected with Myc- and Abl-expressing plasmids. At 36 h post transfection, cell lysates were immunoprecipitated using anti-Flag beads followed by immunoblotting with anti-phosphotyrosine 4G10 (P-Tyr) and anti-Abl. (b) Schematic representation of the Myc mutants used in this work (see list in Materials and methods) and summary of the results for phosphorylation and binding by Abl. (c, d) 293T cells were transfected with plasmids encoding Myc (wild-type or deletion mutants, labeled as in (b)) and Abl, as indicated. Cell lysates were subject to immunoprecipitation with anti-HA or anti-Flag antibodies as indicated, and immunoblotting for tyrosine phosphorylation as in (a). (e) The Myc N-termini from the indicated species and human N-Myc are aligned. The relevant tyrosines (see text), T58 and S62 are indicated. The various Y-F mutant variants of (f) Myc Δ1-24 (labeled as in (b)), or (g) full-length Myc (all Flag-tagged) were expressed in 293T cells together with Abl as indicated, and processed as in (a). The Total levels of the Myc-HA variants in (c) and of the Myc-Flag variants in (d) were assessed by re-probing the same blots with anti-HA and anti-Flag, respectively. Other total levels were all directly determined on whole cell lysates. Asterisks indicate cross-reacting Ig chains present in the blots.
In order to identify the tyrosine(s) phosphorylated by Abl, we mutated candidate residues. We first analyzed a mutant deleted of residues 1–24 that retained only two of the six tyrosines (Y32, Y74: mutant G, Figure 4b), and mutated these to phenylalanine; in this context, substitution of Y32 (mutant H) decreased phosphorylation by Abl, whereas that of Y74 alone, or Y32 and Y74 together (mutants I and J) abrogated all detectable phosphorylation (Figure 4f). In the context of full-length Myc, we targeted Y32, Y74 or a cluster of three residues (Y12/16/22); each of these mutations was introduced alone or in combination with the others. The sixth tyrosine (Y24) was excluded from our analysis, as it was the only one to be (1) conserved in N-Myc (which was not phosphorylated by Abl: Supplementary Figure S1c) and (2) not conserved in mouse c-Myc (Figure 4e). Substitution of Y74 alone in full-length Myc substantially decreased phosphorylation, whereas that of either Y32 or Y12/16/22 had little effect (Figure 4g). The phospho-tyrosine signal was essentially lost when Y74 was substituted together with either Y32, the Y12/16/22 group or all together (the mutant referred hereafter as Myc 5YF). Instead, a mutant retaining only Y74 (with substitution of Y12/16/22 and Y32) showed residual phosphorylation (Figure 4g). Altogether, these data imply that Abl phosphorylates Y74, Y32 and one or more residues among Y12/16/22 in a hierarchical manner, phosphorylation of Y74 being important (although not strictly essential) for that of the others, and retention of either Y32 or Y12/16/22 alone being insufficient to support significant phosphorylation by Abl.
Because Y74 was the main target of Abl, a rabbit antiserum was raised against a Y74-phosphorylated peptide (anti-pY74). Immunoblot and immunoprecipitation analysis on WT or mutant Myc in co-transfected 293T cells showed that the anti-pY74 antibody recognized Myc in a Y74- and Abl-dependent manner (Figure 3a and Supplementary Figures S1d and e). These experiments confirmed that Myc Y74 is phosphorylated by Abl, and provided us with a reagent to detect this form of Myc in cells (see below).
Finally, two experiments were performed to confirm that Abl, rather than a secondary kinase, was directly responsible for Myc phosphorylation. First, we immunoprecipitated Flag-Myc from transfected 293T cells, incubated the precipitates with recombinant Abl and ATP, resolved the reactions by sodium dodecyl sulfate–polyacrylamide gel electrophoresis, and immunoblotted them with the anti-phospho-tyrosine antibody. Wild-type Flag-Myc was effectively phosphorylated in this assay, above the background observed with the 5YF mutant (Supplementary Figure S2a). Second, a bacterially expressed GST-Myc (1–262) protein was incubated in the presence of recombinant Abl and ATP, and analyzed as above, demonstrating direct phosphorylation of the Myc N-terminal domain by Abl (Supplementary Figure S2b).
Staining for Y74-phosphorylated Myc indicates cytoplasmic localization
To test the specificity of anti-pY74 in immunofluorescence (IF) and immunohistochemistry (IHC), we expressed Abl and simultaneously knocked down Myc in HeLa cells by co-transfection with vectors expressing Abl and either a small hairpin RNA (shRNA) directed against the c-myc mRNA (shMyc) or a control scramble shRNA. As expected, a Myc-specific antibody yielded a prevalently nuclear signal; this was largely reduced by shMyc, whereas Abl had no significant effect (Supplementary Figures S3a and S4a). A total of 30–35% of the cells co-transfected with Abl and the control shRNA scored positive for Myc-pY74 but, unlike total Myc, the Myc-pY74 signal was cytoplasmic (Supplementary Figures S3b and S4b). In both assays, co-transfection with shMyc reduced to similar extent the percentages of cells positive for either Myc-pY74 or total Myc, indicating that anti-pY74 was truly recognizing a form of Myc located in the cytoplasm, and not a different crossreacting protein. In IHC, an antibody recognizing Abl auto-phosphorylated on Y412 (P-Abl) stained the same number of cells as Myc-pY74 and, as expected, the P-Abl signal was insensitive to shMyc (compare Supplementary Figures S4b and c; note that the IF signal for P-Abl was very weak).
To visualize Myc and Abl in individual cells, we transfected HeLa cells with a plasmid expressing a GFP-Abl fusion protein, and stained by IF for Myc (either total or pY74); Myc-pY74 was detected exclusively in the cytoplasm of green fluorescent protein (GFP)-positive cells and, as above, co-transfection with shMyc reduced Myc-pY74 and total Myc to the same extent (Figure 5). This result was confirmed in the same experiment by a second shRNA (shMyc2, Supplementary Figure S5).
Myc-pY74 localizes to the cytoplasm. HeLa cells were transfected with GFP-Abl and shMyc or shScramble (Control) vectors. DAPI (4,6-diamidino-2-phenylindole) staining, GFP fluorescence and IF for Myc (total or pY74) were imaged on the confocal microscope. The percentages of Myc- and Myc pY74-positive cells are expressed relative to the total number of GFP-Abl-positive cells (that is, of all transfected cells). White bar, 10 μm.
Altogether, our IF and IHC assays indicate that the form of Myc phosphorylated on Y74 (Myc-pY74) is strictly dependent upon Abl and is localized to the cytoplasm. This form must constitute a minor fraction of the total Myc protein in Abl-expressing cells, explaining the predominantly nuclear localization of the total Myc protein in all conditions, as well as our inability to detect Myc-pY74 above background in biochemical assays (western or immunoprecipitation/western: data not shown). Although we cannot formally rule out the crossreaction of anti-pY74 with a different Abl substrate, loss of the signal upon Myc knockdown by two different c-myc-directed shRNAs constitutes a very strong indication—albeit not formal proof—that this corresponds to cytoplasmic Myc-pY74.
Mutation of Y74 decreases Myc activity in a Bcr-Abl/Myc co-transformation assay
To investigate the potential role of Y74 phosphorylation in Myc-transforming activity, we monitored the co-transformation of Rat1 cells by Myc and Bcr-Abl.19, 20 The cells were sequentially infected with retroviruses encoding Bcr-Abl (WT or KD) and Myc (WT or Y74F; the 5YF mutant was not efficiently expressed in this assay and was not studied further). The expression of active Bcr-Abl was confirmed by immunoblotting whole-cell lysates with an anti-phospho-tyrosine antibody (Supplementary Figure S6a). WT and Y74F Myc were expressed at similar levels and independently from Bcr-Abl activity (Supplementary Figure S6b). Double-infected cells were plated in soft agar, and colonies counted after 2 weeks. As expected, cells coexpressing Bcr-Abl WT and Myc formed colonies in soft agar (Supplementary Figure S6c, left). Expression of either protein alone, or of Bcr-Abl KD with or without Myc, did not allow colony formation (data not shown). The Y74F mutation did not abolish Myc-transforming activity together with Bcr-Abl WT, but caused a 2- to 3-fold reduction in colony numbers, as shown in two independent experiments (Supplementary Figure S6c). This result implies that Y74 is critical for maximal Myc activity in cellular transformation, whether due to phosphorylation of Y74 by Abl or another effect of this residue.
Cytoplasmic Myc-pY74 is detectable in mammary carcinomas with active Abl
A fraction of breast cancer cell lines (5 out of 9 tested) were reported to express active Abl.21, 22 However, it remained to be addressed whether Abl activation—and in the context of this work also Myc Y74 phosphorylation—occurs in tumors. We thus performed an IHC screen for phospho-Abl, Myc-pY74 and total Myc on a series of 70 breast cancer samples spotted on a tissue microarray. A minor fraction of all tumors (6/70 or 8.6%) scored positive for P-Abl, all of which also scored with Myc-pY74. Among the 64 remaining samples, only 4 (6.3%) scored with Myc-pY74 but not P-Abl, most likely because of phosphorylation of Y74 by another tyrosine kinase. Thus, although rare, Abl activation in mammary carcinoma appears to be systematically associated with Myc Y74 phosphorylation (P=1.6E–06). Of the 6 P-Abl/Myc-pY74 double-positive tumors, 5 (83%) were metastatic, compared with 31/60 (51.7%) of the double-negative cases; this difference, however, did not reach statistical significance (P=0.12). Of the four Myc-pY74-only tumors, only one was metastatic. Larger number will be required to address whether Abl activation correlates with metastasis, as reported in melanoma.23 Similarly, no clear correlation could be established between phospho-Abl/Myc-pY74 and other clinical features such as estrogen receptor or progesterone receptor, p53 or HER2 status.
Close examination of our IHC sections revealed that the Myc-pY74 signal was mainly cytoplasmic whereas total Myc was predominantly nuclear (Figure 6a and Supplementary Figure S7). Thus, in agreement with our data on transfected HeLa cells, Abl activation in mammary carcinoma is accompanied by the appearance of the Myc-pY74 signal in the cytoplasm.
Cytoplasmic Myc-pY74 is detectable by IHC in p-Abl positive carcinomas. (a) Representative images from our breast cancer tissue microarray stained with the indicated antibodies. (b) Tissue sections of the positive ovarian carcinoma identified in the second tissue microarray (Supplementary Figure S8).
In a parallel experiment, we used a tissue microarray including a total of 416 cases of 15 different types of solid tumors. IHC analysis revealed seven cases positive for P-Abl, and four for Myc-pY74: of these, two scored as double positives, including a cervical and an ovarian carcinoma (Supplementary Figure S8). The latter was also analyzed in whole sections, showing prevalent nuclear staining for total Myc and cytoplasmic staining for Myc-pY74 (Figure 6b).
Myc-Y74 phosphorylation by Bcr-Abl in chronic myeloid leukemia
The Bcr-Abl fusion is the driving oncogene in chronic myeloid leukemia (CML).24 As shown above for Abl, Bcr-Abl induced Myc Y74 phosphorylation when coexpressed with Myc in 293T cells (Supplementary Figures S2c and d). IF with anti-pY74 yielded a clear cytoplasmic staining in three CML cell lines (MOLM, KYO and K562) but not in HL60, a Bcr-Abl-negative promyelocytic leukemia cell line (Figure 7a). Knockdown of Myc in K562 cells by infection with the shMyc vector led to a proportional reduction in cells reactive with total Myc and pY74 antibodies, whereas only the Myc-pY74 signal was eliminated by treatment of the cells with STI571 (Supplementary Figure S9). The cytoplasmic Myc-pY74 signal was also detectable in a substantial fraction (15 to 25%) of precursors and more mature myeloid cells in bone marrow samples from CML patients (Figure 7b), whereas non-neoplastic bone marrow specimens were negative (Figure 7c). Altogether, our data indicate that cytoplasmic Myc-pY74 occurs in CML as a consequence of Bcr-Abl activity.
Myc-pY74 is located in the cytoplasm of CML cells. Representative images of immunofluorescence using the Myc-pY74 antibody, performed on (a) three CML cell lines (MOLM, KYO and K562) and a promyelocytic leukemia cell line (HL60), (b) bone marrow samples from CML patients and (c) non-neoplastic bone marrow.
Discussion
Abl indirectly regulates Myc T58/S62 phosphorylation, interaction with Pin1 and p300, and acetylation
In the first part of our work, we have described a series of effects of Abl on Myc, which are likely to be intimately interconnected: (1) enhancement of Myc S62 and T58 phosphorylation, (2) increased binding of Pin1 and p300 and (3) hyper-acetylation of Myc by p300. These effects are likely to result in augmented transcriptional activity of Myc (Figure 1b). Although the latter could not be tested directly because of the interference by Abl with reporter constructs, Pin1 and p300 together showed additive positive effects on Myc transcriptional activity. Most importantly, this chain of events—as demonstrated for hyper-phosphorylation of S62/T58 and hyper-acetylation of Myc by p300—does not require direct tyrosine phosphorylation of Myc, and must therefore depend on the phosphorylation of different Abl substrates.
Positive effects of Abl on p300-mediated acetylation have been described in different settings.17, 25, 26 Besides possible effects on p300 recruitment or activity, Abl may be boosting signaling pathways that can promote Myc-S62 phosphorylation, such as the extracellular signal-regulated kinase (ERK), c-Jun N-terminal (JNK) kinase or p38 mitogen-activated protein kinase pathways.27, 28, 29, 30 Phosphorylated S62 will in turn prime T58 phosphorylation and Pin1 binding.7 How Pin1 would contribute to enhancing p300 interaction and Myc acetylation remains to be addressed. Most importantly in this context, our data reveal a new facet of Pin1 activity on Myc; previous data indicated that Pin1 is required upon phosphorylation of T58 for recruitment of protein phosphatase 2A to Myc, S62 de-phosphorylation and Myc turnover.7 In our experiments, instead, both S62 and T58 phosphorylation were enhanced by coexpression of Pin1 and Abl, accompanied by enhanced p300 binding and Myc acetylation. In preliminary experiments, we have observed no consistent effects of Abl on the Myc/protein phosphatase 2A interaction or bulk Myc levels (see, for example, Figures 2b and c). We surmise that the promotion of S62-T58 phosphorylation and p300 recruitment may be distinct effects of Pin1 and Abl.
Our data also indicate that Pin1 and p300, in the absence of Abl activity, augment transcriptional activation by Myc. This is consistent with previous work showing that Pin1 and p300 together modulate the activity of other transcription factors. Examples include signal transducer and activator of transcription 3 (STAT3) in the regulation of epithelial-to-mesenchymal transition,31 or c-Jun and c-Fos in response to growth factors.4, 5 A similar mechanism has been described for the co-activator SRC-3, which binds Pin1 and enhances the transcriptional activity of the estrogen receptor through p300.32 The transcriptional activities of p53 and p73 can be regulated in a similar way.3, 33 In the case of p73, distinct effects coexist, one direct and the other not (Figure 1a); first, phosphorylation of p73 on Y99 by Abl is responsible for stabilization of the transcription factor;17, 34, 35 second, Abl also activates p38 mitogen-activated protein kinase, which phosphorylates p73, generating binding sites for Pin1 that enhance p73 acetylation through p300 recruitment.3 These observations provide a strong mechanistic rationale for our results on Myc, including (1) the cooperative effect of Pin1 and p300 in enhancing Myc transcriptional activity and (2) the enhancement of Pin1/p300 recruitment and Myc acetylation by Abl, although independently from direct phosphorylation of Myc on tyrosine. Indeed, as will be discussed below, direct phosphorylation of Myc by Abl appears to have a wholly different consequence (Figure 1b).
The notion that Abl signaling promotes Myc acetylation and—as we infer—transcriptional activity (Figure 1b) lends new significance to previous reports indicating that Myc is essential for Abl-transforming activity36 and that Myc overexpression in a CML cell line can antagonize induction of differentiation by Abl inhibitors.37 Altogether, these observations indicate that Abl indirectly enhances Myc activity (Figure 1b), which in turn is an essential effector of Abl in cellular transformation.
Abl phosphorylates Myc on tyrosine: cytoplasmic localization of Myc-pY74
Our data show that Abl phosphorylates Myc in vivo and in vitro. We mapped the residues phosphorylated by Abl to a group of five tyrosines located in the N-terminal part of the protein, flanking the first conserved Myc box (MBI). These residues include Y74—which is the main target of Abl—as well as Y32, and one or more among Y12, Y16 or Y22. Our data indicate that Y74-phosphorylated Myc (Myc-pY74) constitutes a minor part of the Myc protein and is localized to the cytoplasm (Figure 1b).
In IF and IHC assays, the anti-pY74 antibody allowed us to detect endogenous Myc-Y74 in the cytoplasm, in a manner dependent upon Abl activity. With the reagents available to date, we have not been able to provide a biochemical detection of endogenous Myc-pY74 above background (whether in the nucleus or cytoplasm), most likely owing to the low amounts of this form of Myc in cells. Thus, a plausible criticism to our data is that the cytoplasmic signal seen by IF and IHC might be because of a crossreacting phospho-epitope on an unrelated Abl substrate(s). To address this issue, we knocked down Myc expression through RNA interference with two independent shRNA constructs; in all our experiments, the percentage of cells scoring positive for cytoplasmic Myc-pY74 and nuclear total Myc decreased to similar extents. Although still short of definitive biochemical confirmation, these data provide strong evidence for the presence of Myc-pY74 in the cytoplasm as a consequence of Abl activation.
Our co-transfection and co-immunoprecipitation data showed that a distinct C-terminal region of Myc mediates stable interaction with Abl. Most importantly, in cells, this interaction is rate limiting for phosphorylation of the N-terminal tyrosines. The Abl-binding domain in the Myc C-terminal region overlaps with the basic helix-loop-helix leucine zipper (bHLH-LZ) domain, responsible for its dimerization with Max and DNA binding.38 As the bulk of Myc in cells is associated with Max and is localized to the nucleus, it remains unclear whether Abl initially interacts with free Myc or with Myc/Max dimers, and whether this occurs in the nucleus or in the cytoplasm. In an analogous manner, we presently do not know whether Y74 phosphorylation directly instructs cytoplasmic retention and/or nuclear export of Myc, or whether it occurs in the cytoplasm as a consequence of an independent localization mechanism.
Myc-Nick is a cytoplasmic form of Myc resulting from cleavage by calpain and loss of the C-terminal domain.39 Myc-Nick interacts with α-tubulin, promotes its acetylation and, unlike full-length Myc, favors cellular differentiation. Thus, conceivably, Myc-Nick might be the form of Myc phosphorylated by Abl in vivo. We deem this possibility unlikely, based on our finding that binding of Abl to the Myc C-terminus is rate limiting for phosphorylation of the N-terminal tyrosines. Nonetheless, Abl was capable of phosphorylating N-terminal fragments of Myc in our transfected cells; conceivably, other mechanisms may bring Myc-Nick and Abl in close proximity under physiological settings, perhaps in association with tubulins, as Abl interacts with the tubulin-associated protein Rack1.40 As an alternative possibility, tyrosine phosphorylation of Myc may promote its cleavage by calpain, resulting in the accumulation of tyrosine-phosphorylated Myc-Nick in the cytoplasm. Our transfection and immunoprecipitation/western experiments clearly showed that Abl can phosphorylate full-length Myc (Figure 4), but did not allow us to conclude about the effect of Abl on the processing of Myc. Moreover, most likely because of the low fraction of Myc phosphorylated on Y74 combined with the poor sensitivity of the anti-pY74 antibody, we have not been able to detect endogenous Myc-pY74 biochemically in cells, whether in the full-length or the truncated form. Hence, the existence of tyrosine-phosphorylated Myc-Nick remains a formal possibility.
In additional experiments based on cleavage of in vitro-translated Myc,39 we have observed that K562 cytoplasmic lysates show very high calpain activity; this was unaffected by treatment of the cells with imatinib before preparation of the lysate, suggesting that Bcr-Abl activity does not directly affect calpain activity.
Cytoplasmic Myc-pY74 is detectable in tumors with active Abl or Bcr-Abl
IHC analysis of tissue microarrays with a phospho-Abl antibody revealed Abl activation in a small minority of mammary carcinomas, associated with the presence of the cytoplasmic Myc-pY74 signal. Consistent with previous studies associating Abl activity to invasiveness in breast cancer cell lines,21, 22 the majority of the P-Abl/Myc-pY74-positive cases in our data set were metastatic. Whether Myc-pY74 or other Abl substrates contribute to the metastatic phenotype remains to be addressed.
Our co-transfection experiments showed that the CML-associated oncoprotein Bcr-Abl was as effective as Abl in phosphorylating Myc. IF analysis of CML cell lines revealed a cytoplasmic Myc-pY74 signal that was sensitive to either Myc knockdown or treatment with STI571. Altogether, these data indicate that the activation of Bcr-Abl in CML leads to the appearance of cytoplasmic Myc-pY74.
Several studies reported the existence of a cytoplasmic Myc signal in tumors,41, 42, 43 although those observations had remained unexplained. We surmise that this may be due to Y74 phosphorylation, to the cleaved form Myc-Nick,39 or both. Our findings warrant future analysis of the biological function of Myc-pY74 in the cytoplasm, and of its contribution to the pathogenesis of CML and other Abl-associated tumors.
Materials and methods
Plasmids
pEBB-Abl WT and pSG5-Abl KD (expressing the kinase-dead mutant K290R) were supplied by Ricardo Sanchez-Prieto (Albacete, Spain); pCMV-N-Myc, pCMV-HA-p300, pCDNA3-HA-Pin1 WT and Y23A were supplied by Gianni del Sal (Trieste, Italy); the Bcr-Abl plasmids pKI-210 WT and pKI 210 KD (expressing the kinase-dead mutant) were supplied by Warren Pear (Philadelphia, PA, USA); and pCEFL-GFP-Abl by Silvio Gutkind (Bethesda, MD, USA). pcDNA-210 was supplied by Pier-Giuseppe Pelicci (Milan, Italy). For the mapping of interaction and phosphorylation sites on Myc, we used plasmids pCβF-Myc Flag, pCβF-MycΔ1–110 Flag and pCβF-MycΔ1–180 Flag, kindly provided by Michael Cole (Lebanon, NH, USA), as well as pCDNA3-Myc(1–262)-HA (mutant C in Figure 2b; our reference number BA1594), pCDNA3-Myc(1–347)-HA (mutant B; BA1595), pCDNA3-Myc(1–145)-HA (mutant D; BA1593), pCDNA3-Myc(263–439)-HA (mutant A; BA1518), pCMV-Myc-T58A-Flag (BA1987), pCMV-Myc-S62A-Flag (BA1986), pCMV-Myc-T58A-S62A-Flag (BA2235) and pGEX-Myc 1–262 (BA91). The Myc deletion mutant Δ1–24 and point mutants thereof were generated using site-directed mutagenesis by PCR, subcloning in a pCMV Myc-Flag vector and confirmation by DNA sequencing. The constructs were pCMV-Δ1–24 Myc Flag (mutant G; BA2029), pCMV-Δ1–24 Y32F Myc Flag (mutant H; BA2030), pCMV-Δ1–24 Y74F Myc Flag (mutant I; BA2088), pCMV-Δ1–24 Y32-74F Myc Flag (mutant J; BA2089), pCMV-Myc Flag (BA2046), pCMV-Y12,16,22,32,74F Myc Flag (mutant 5YF; BA2095), pCMV-Y12,16,22,74F Myc Flag (BA2094), pCMV-Y12,16,22,32F Myc Flag (BA2093), pCMV-Y12,16,22F Myc Flag (BA2023), pCMV-Y32F Myc Flag (BA2090), pCMV-Y74F Myc Flag (BA2091). To generate these mutant constructs we performed PCR with the following primers:
Myc Y12,16,22F:
PCR1: 5′primer (AP5435): GAAGAAGGATCCCCGGGCGAGC; 3′primer (AP5438): CGGCTGCACCGAGTCGWAGTCAAGGTCGWAGTTCCTGTTGGT
PCR2: 5′primer (AP5437): GACTCGGTGCAGCCGTWCTTCTACTGCGAC
3′primer (AP5436 after the ClaI site in c-myc): AGCAGAAGGTGATCCAGACTCTGAC
PCR3: Used the combined PCR1 and PCR2 products as template, with primers AP5435 and AP5436.
Myc Δ1-24 Y32F:
PCR1: 5′primer (AP5441): GAAGAAGGATCCAACATGTGCGACGAGGAGGAGAACTTCTWCCAGCA; 3′primer (AP5436).
Myc Y32F:
PCR1: 5′primer (AP5435): GAAGAAGGATCCCCGGGCGAGC; 3′primer (AP5568): GCTGCTGGAAGAAGTTCTCCTC
PCR2: 5′primer (AP5567): GAACTTCTTCCAGCAGCAGCA; 3′primer (AP5436 after the ClaI site in c-myc): AGCAGAAGGTGATCCAGACTCTGAC
PCR3: 5′primer (AP5435); 3′primer (AP5436)
Myc Y74F:
PCR1: 5′primer (AP5435) GAAGAAGGATCCCCGGGCGAGC; 3′primer (AP5570): GCAACGAAGGAGGGCGA
PCR2: 5′ primer (AP5569): CCTCCTTCGTTGCGGTCA; 3′primer (AP5436 after the ClaI site in c-myc): AGCAGAAGGTGATCCAGACTCTGAC
PCR3: 5′primer (AP5435); 3′primer (AP5436)
Fusion of 12/16/22 mutations and 32/74 mutations:
PCR1: 5′primer (AP5435): GAAGAAGGATCCCCGGGCGAGC; 3′primer (AP6008): GAAGTTCTCCTCCTCGTCGCA
PCR2: 5′ primer (AP6007): TTCTACTGCGACGAGGAGGAGAAC; 3′primer (AP5436 after the ClaI site in c-myc): AGCAGAAGGTGATCCAGACTCTGAC
PCR3: 5′primer (AP5435); 3′primer (AP5436)
For Rat1 infection we used pBH2-HA-Flag-Myc (BA2313) and pBH2-HA-Flag-Myc Y74F (BA2315)
Antibodies
The following antibodies were used: anti-Abl (K-19), anti-Myc (C-33) anti-Myc (N-262) and anti-N-Myc (H-50) from Santa Cruz (Santa Cruz, CA, USA); Myc (Y69) from Abcam (Cambridge, UK); anti-HA from Covance (San Diego, CA, USA) (MMS-101P); anti-Flag (F3165), anti-Flag beads (A2220) and anti-Vinculin (V9264) from Sigma (St Louis, MO, USA); anti-phosphotyrosine (4G10) and anti-Pin1 (PC270) from Upstate (Billerica, MA, USA); anti-acetyl lysine (no. 9441), anti-P-Abl (Y412) and anti-Myc phospho-Thr 58/Ser 62 (no. 9401) from Cell Signaling (Danvers, MA, USA).
The antibody directed against phospho-Tyr 74 in Myc (here anti Myc-pY74) was generated by Abcam upon our request and is commercially available (ab46848; Abcam).
In order to selectively investigate the phosphorylation status of Myc Ser 62, rabbit polyclonal antisera were raised against a phosphorylated peptide containing phospho-Ser 62. For the immunization and subsequent affinity purification steps the following peptides were synthesized:
Myc ELL pS62: ELLPTPPL(p)SPSRRSGLC
Myc ELL pT58: ELLP(p)TPPLSPSRRSGLC
Myc ELL pT58-S62: ELLP(p)TPPL(p)SPSRRSGLC
Myc ELL: ELLPTPPLSPSRRSGLC
Crude bleeds were first immunodepleted with Myc ELL pT58, Myc ELL pT58-S62 and non-phosphorylated Myc ELL, and then affinity purified against the immunogenic peptide (Myc ELL p-S62). For characterization, the purified antibodies were used to immunoblot lysates from 293T cells transfected with expression vectors encoding either Flag-tagged Myc WT or T58A/S62A mutants. As shown in Supplementary Figure S1, purified anti-Myc p62 efficiently recognized both Myc WT and T58A, whereas, as expected, no signal was detected for the S62A mutant (Supplementary Figure S10).
Cell culture, transfections and reagents
K562, MOLM, KYO, 293T, U2OS and HeLa cells were grown under standard conditions. Pin1 KO mouse embryonic fibroblasts were cultivated in Dulbecco’s modified Eagle’s medium supplemented with glutamine, 10% fetal bovine serum, 100 units/ml penicillin, 100 mg/ml streptomycin, non-essential amino acids and β-mercaptoethanol. Cells were transfected using lipofectamine (Invitrogen, Grand Island, NY, USA) following the protocol supplied by the vendor. The Abl inhibitor STI571 was kindly provided by Novartis (Basel, Switzerland).
Immunoprecipitation, co-immunoprecipitation and in vitro binding
GST-Pin1 pull-down assays were performed as described (Zacchi el al.33). Immunoprecipitation was carried out essentially as described in Haupt et al. (1996)44.
IF and IHC
For IF, adherent cells were grown on glass slides and fixed for 10 min in 4% paraformaldehyde. Cells growing in suspension were subjected to a cytospin (20 g for 5 min) and fixed as above. Cells were permeabilized with 0.2% Triton in phosphate-buffered saline (10 min) and blocked in 5% bovine serum albumin (30 min at room temperature), followed by incubation with the primary antibody in 5% bovine serum albumin (1 h 30 min at room temperature) and with the secondary antibody (cy3 anti-rabbit) in 5% bovine serum albumin (60 min at room temperature). The slides were finally stained with 4,6-diamidino-2-phenylindole and mounted with Mowiol Mounting Media Germany Polysciences, Eppelheim, Germany.
IHC staining was performed with the avidin–biotin–peroxidase technique. Sections, 5 μm thick, were cut from the tissue specimens and placed on poly-L-lysine-coated glass slides. Sections were deparaffined by xylene and rehydrated in graded alcohol. Endogenous peroxidase was blocked by immersing the sections in 0.1% hydrogen peroxidase in absolute methanol for 20 min. For antigen retrieval, the tissue sections were heated in a pressure cooker in citric acid monohydrate 10 mM, pH 9.0, for 5 min, and then incubated with the primary antibody at room temperature. IHC was performed with Benchmark XT (Ventana Medical Systems, Inc., Tucson, AZ, USA). The primary antibodies and dilutions used were: anti-p-Abl (1:50, p-Y412; Cell Signaling Technology), anti-Myc-pY74 (1:50, ab46848; Abcam) and anti-Myc (1:100, ab32072, Y69 rabbit monoclonal; Abcam ). The incubation time for the antibodies was 60, 120 and 320 min, respectively. All slides were hematoxylin counterstained, dehydrated and mounted. Negative controls were performed by omitting the primary antibody.
Kinase assay
293T cells were transfected with Myc WT or 5YF mutant. Myc was immunoprecipitated using anti-Flag beads, followed by three washes in IP buffer (50 mM Tris pH 8, 5 mM EDTA, 150–300 mM NaCl and 0.5% NP-40) and one wash in kinase buffer. Immunoprecipitates were resuspended in kinase buffer (60 mM HEPES, 5 mM MgCl2, 5 mM MnCl2, 3 mM Na2VO4, 1.25 mM DTT and 200 mM ATP) adding 50 ng Abl1 Kinase (Cell Signaling). The reaction was incubated at room temperature for 30 min. Immunoprecipitates were analyzed by immunoblot with the anti-phosphotyrosine 4G10 antibody.
Dual luciferase assay
U2OS cells were transfected with Fugene with 200 ng of pNuc-Luc reporter, 200 ng of pCMV-Flag-Myc, 100 ng pcDNA-HA-Pin1 and 50 ng pCMV-HA-p300. For normalization and transfection efficiency control, we used 8 ng of pRL-TK reporter (Promega, Fitchburg, WI, USA) that constitutively expresses the Renilla luciferase. After 36 h, cells were lysed and assayed for luciferase activity using the Dual Luciferase kit (Promega).
Rat1 co-transformation assay
Co-transformation of Rat1 cells by Myc and Bcr-Abl was as previously described.19, 20 To produce recombinant retroviruses, Phoenix Eco cells were transfected with retroviral constructs expressing Bcr-Abl (pKI-Bcr-Abl WT or KD) or Myc (pBabe-Hygro-Myc-Flag, WT or Y74F). Rat1 cells were infected with either form of the Bcr-Abl viruses, selected with Puromycin, superinfected with either form of the Myc viruses and selected with Hygromycin. Infected cells were directly plated in triplicate in medium containing 0.3% Bacto-agar laid on top of a layer of 0.6% Bacto-agar, and cultured for 2 weeks. Fresh media were changed every 2 days. Colonies were visualized by 0.1% p-iodonitrotetrazolium violet (INT, Sigma) staining.
References
Lu KP, Finn G, Lee TH, Nicholson LK . Prolyl cis-trans isomerization as a molecular timer. Nat Chem Biol 2007; 3: 619–629.
Mantovani F, Tocco F, Girardini J, Smith P, Gasco M, Lu X et al. The prolyl isomerase Pin1 orchestrates p53 acetylation and dissociation from the apoptosis inhibitor iASPP. Nat Struct Mol Biol 2007; 14: 912–920.
Mantovani F, Piazza S, Gostissa M, Strano S, Zacchi P, Mantovani R et al. Pin1 links the activities of c-Abl and p300 in regulating p73 function. Mol Cell 2004; 14: 625–636.
Wulf GM, Ryo A, Wulf GG, Lee SW, Niu T, Petkova V et al. Pin1 is overexpressed in breast cancer and cooperates with Ras signaling in increasing the transcriptional activity of c-Jun towards cyclin D1. EMBO J 2001; 20: 3459–3472.
Monje P, Hernández-Losa J, Lyons RJ, Castellone MD, Gutkind JS . Regulation of the transcriptional activity of c-Fos by ERK. A novel role for the prolyl isomerase PIN1. J Biol Chem 2005; 280: 35081–35084.
Bitomsky N, Böhm M, Klempnauer K-H . Transformation suppressor protein Pdcd4 interferes with JNK-mediated phosphorylation of c-Jun and recruitment of the coactivator p300 by c-Jun. Oncogene 2004; 23: 7484–7493.
Yeh E, Cunningham M, Arnold H, Chasse D, Monteith T, Ivaldi G et al. A signalling pathway controlling c-Myc degradation that impacts oncogenic transformation of human cells. Nat Cell Biol 2004; 6: 308–318.
Adhikary S, Marinoni F, Hock A, Hulleman E, Popov N, Beier R et al. The ubiquitin ligase HectH9 regulates transcriptional activation by Myc and is essential for tumor cell proliferation. Cell 2005; 123: 409–421.
Faiola F, Liu X, Lo S, Pan S, Zhang K, Lymar E et al. Dual regulation of c-Myc by p300 via acetylation-dependent control of Myc protein turnover and coactivation of Myc-induced transcription. Mol Cell Biol 2005; 25: 10220–10234.
Zhang K, Faiola F, Martinez E . Six lysine residues on c-Myc are direct substrates for acetylation by p300. Biochem Biophys Res Commun 2005; 336: 274–280.
Vervoorts J, Luscher-Firzlaff JM, Rottmann S, Lilischkis R, Walsemann G, Dohmann K et al. Stimulation of c-MYC transcriptional activity and acetylation by recruitment of the cofactor CBP. EMBO Rep 2003; 4: 484–490.
Patel JH, Du Y, Ard PG, Phillips C, Carella B, Chen C-J et al. The c-MYC oncoprotein is a substrate of the acetyltransferases hGCN5/PCAF and TIP60. Mol Cell Biol 2004; 24: 10826–10834.
Frank SR, Schroeder M, Fernandez P, Taubert S, Amati B . Binding of c-Myc to chromatin mediates mitogen-induced acetylation of histone H4 and gene activation. Genes Dev 2001; 15: 2069–2082.
Frank SR, Parisi T, Taubert S, Fernandez P, Fuchs M, Chan H-M et al. MYC recruits the TIP60 histone acetyltransferase complex to chromatin. EMBO Rep 2003; 4: 575–580.
Greasley PJ, Bonnard C, Amati B . Myc induces the nucleolin and BN51 genes: possible implications in ribosome biogenesis. Nucleic Acids Res 2000; 28: 446–453.
Lu PJ, Zhou XZ, Shen M, Lu KP . Function of WW domains as phosphoserine- or phosphothreonine-binding modules. Science 1999; 283: 1325–1328.
Gong JG, Costanzo A, Yang HQ, Melino G, Kaelin WG, Levrero M et al. The tyrosine kinase c-Abl regulates p73 in apoptotic response to cisplatin-induced DNA damage. Nature 1999; 399: 806–809.
Gregory MA, Qi Y, Hann SR . Phosphorylation by glycogen synthase kinase-3 controls c-myc proteolysis and subnuclear localization. J Biol Chem 2003; 278: 51606–51612.
Blackwood EM, Lugo TG, Kretzner L, King MW, Street AJ, Witte ON et al. Functional analysis of the AUG- and CUG-initiated forms of the c-Myc protein. Mol Biol Cell 1994; 5: 597–609.
Lugo TG, Witte ON . The BCR-ABL oncogene transforms Rat-1 cells and cooperates with v-myc. Mol Cell Biol 1989; 9: 1263–1270.
Srinivasan D, Sims JT, Plattner R . Aggressive breast cancer cells are dependent on activated Abl kinases for proliferation, anchorage-independent growth and survival. Oncogene 2008; 27: 1095–1105.
Srinivasan D, Plattner R . Activation of Abl tyrosine kinases promotes invasion of aggressive breast cancer cells. Cancer Res 2006; 66: 5648–5655.
Ganguly SS, Fiore LS, Sims JT, Friend JW, Srinivasan D, Thacker MA et al. c-Abl and Arg are activated in human primary melanomas, promote melanoma cell invasion via distinct pathways, and drive metastatic progression. Oncogene 2011; 31: 1804–1816.
Deininger MWN, Goldman JM, Melo JV . The molecular biology of chronic myeloid leukemia. Blood 2000; 96: 3343–3356.
Levy D, Adamovich Y, Reuven N, Shaul Y . Yap1 phosphorylation by c-Abl is a critical step in selective activation of proapoptotic genes in response to DNA damage. Mol Cell 2008; 29: 350–361.
Oh AS, Lahusen JT, Chien CD, Fereshteh MP, Zhang X, Dakshanamurthy S et al. Tyrosine phosphorylation of the nuclear receptor coactivator AIB1/SRC-3 is enhanced by Abl kinase and is required for its activity in cancer cells. Mol Cell Biol 2008; 28: 6580–6593.
Sánchez-Prieto R, Sanchez-Arevalo VJ, Servitja J-M, Gutkind JS . Regulation of p73 by c-Abl through the p38 MAP kinase pathway. Oncogene 2002; 21: 974–979.
Barilá D, Mangano R, Gonfloni S, Kretzschmar J, Moro M, Bohmann D et al. A nuclear tyrosine phosphorylation circuit: c-Jun as an activator and substrate of c-Abl and JNK. EMBO J 2000; 19: 273–281.
Kharbanda S, Pandey P, Yamauchi T, Kumar S, Kaneki M, Kumar V et al. Activation of MEK kinase 1 by the c-Abl protein tyrosine kinase in response to DNA damage. Mol Cell Biol 2000; 20: 4979–4989.
Pandey P, Raingeaud J, Kaneki M, Weichselbaum R, Davis RJ, Kufe D et al. Activation of p38 mitogen-activated protein kinase by c-Abl-dependent and -independent mechanisms. J Biol Chem 1996; 271: 23775–23779.
Lufei C, Koh TH, Uchida T, Cao X . Pin1 is required for the Ser727 phosphorylation-dependent Stat3 activity. Oncogene 2007; 26: 7656–7664.
Yi P, Wu R-C, Sandquist J, Wong J, Tsai SY, Tsai M-J et al. Peptidyl-prolyl isomerase 1 (Pin1) serves as a coactivator of steroid receptor by regulating the activity of phosphorylated steroid receptor coactivator 3 (SRC-3/AIB1). Mol Cell Biol 2005; 25: 9687–9699.
Zacchi P, Gostissa M, Uchida T, Salvagno C, Avolio F, Volinia S et al. The prolyl isomerase Pin1 reveals a mechanism to control p53 functions after genotoxic insults. Nature 2002; 419: 853–857.
Agami R, Blandino G, Oren M, Shaul Y . Interaction of c-Abl and p73alpha and their collaboration to induce apoptosis. Nature 1999; 399: 809–813.
Yuan ZM, Shioya H, Ishiko T, Sun X, Gu J, Huang YY et al. p73 is regulated by tyrosine kinase c-Abl in the apoptotic response to DNA damage. Nature 1999; 399: 814–817.
Sawyers CL, Callahan W, Witte ON . Dominant negative MYC blocks transformation by ABL oncogenes. Cell 1992; 70: 901–910.
Gomez-Casares MT, Garcia-Alegria E, Lopez-Jorge CE, Ferrandiz N, Blanco R, Alvarez S et al. MYC antagonizes the differentiation induced by imatinib in chronic myeloid leukemia cells through downregulation of p27KIP1. Oncogene 2013; 32: 2239–2246.
Adhikary S, Eilers M . Transcriptional regulation and transformation by Myc proteins. Nat Rev Mol Cell Biol 2005; 6: 635–645.
Conacci-Sorrell M, Ngouenet C, Eisenman RN . Myc-Nick: a cytoplasmic cleavage product of myc that promotes alpha-tubulin acetylation and cell differentiation. Cell 2010; 142: 480–493.
Huang CC, Liu CH, Chuang NN . An enhanced association of RACK1 with Abl in cells transfected with oncogenic ras. Int J Biochem Cell Biol 2008; 40: 423–431.
Craig RW, Buchan HL, Civin CI, Kastan MB . Altered cytoplasmic/nuclear distribution of the c-myc protein in differentiating ML-1 human myeloid leukemia cells. Cell Growth Differ 1993; 4: 349–357.
Sundaresan V, Forgacs IC, Wight DG, Wilson B, Evan GI, Watson JV . Abnormal distribution of c-myc oncogene product in familial adenomatous polyposis. J Clin Pathol 1987; 40: 1274–1281.
Williams AR, Piris J, Wyllie AH . Immunohistochemical demonstration of altered intracellular localization of the C-Myc oncogene product in human colorectal neoplasms. J Pathol 1990; 160: 287–293.
Haupt Y, Barak Y, Oren M . Cell type-specific inhibition of p53-mediated apoptosis by mdm2. EMBO J 1996; 15: 1596–1606.
Acknowledgements
We thank Theresia Kress, Stefano Campaner and Giorgio Scita for critical reading of the manuscript, Gianni Del Sal for scientific discussion and reagents, Warren Pear, Ricardo Sanchez-Prieto, Silvio Gutkind, Michael Cole and Pier-Giuseppe Pelicci for reagents, Celine Ngouenet, Arianna Vino, Lorenzo Spagnuolo and Teresa Moliné for technical assistance, and the Antibody facility at the IFOM-IEO Campus. STI571 was supplied by Novartis. VJSAL was supported by a post-doctoral fellowship from the Ministerio de Ciencia e Inovacion. Work in the Amati lab was supported by grants from the European Commission FP7 Program (EuroSyStem and MODHEP), the European Research Council, the Association for International Cancer Research (AICR), the Cariplo Foundation, the Italian Health Ministry and the Italian Association for Cancer Research (AIRC).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on the Oncogene website
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/3.0/
About this article
Cite this article
Sanchez-Arévalo Lobo, V., Doni, M., Verrecchia, A. et al. Dual regulation of Myc by Abl. Oncogene 32, 5261–5271 (2013). https://doi.org/10.1038/onc.2012.621
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/onc.2012.621
Keywords
This article is cited by
-
The ABL-MYC axis controls WIPI1-enhanced autophagy in lifespan extension
Communications Biology (2023)
-
MYC in liver cancer: mechanisms and targeted therapy opportunities
Oncogene (2023)
-
The acquisition order of leukemic drug resistance mutations is directed by the selective fitness associated with each resistance mechanism
Scientific Reports (2023)
-
Metabolic convergence on lipogenesis in RAS, BCR-ABL, and MYC-driven lymphoid malignancies
Cancer & Metabolism (2021)
-
The impact of phosphatases on proliferative and survival signaling in cancer
Cellular and Molecular Life Sciences (2018)