Abstract
Simple-sequence repeats (SSRs) have increasingly become the marker of choice for population genetic analyses. Unfortunately, the development of traditional ‘anonymous’ SSRs from genomic DNA is costly and time-consuming. These problems are further compounded by a paucity of resources in taxa that lack clear economic importance. However, the advent of the genomics age has resulted in the production of vast amounts of publicly available DNA sequence data, including large collections of expressed sequence tags (ESTs) from a variety of different taxa. Recent research has revealed that ESTs are a potentially rich source of SSRs that reveal polymorphisms not only within the source taxon, but in related taxa, as well. In this paper, we review what is known about the transferability of EST-SSRs from one taxon to another with particular reference to the potential of these markers to facilitate population genetic studies. As an example of the utility of these resources, we then cross-reference existing EST databases against lists of rare, endangered and invasive plant species and conclude that half of all suitable EST databases could be exploited for the population genetic analysis of species of conservation concern. We then discuss the advantages and disadvantages of EST-SSRs in the context of population genetic applications.
Similar content being viewed by others
Introduction
Population genetics analyses can provide data on a variety of important evolutionary parameters, including standing levels of genetic variation, the partitioning of this variability within/between populations, overall levels of inbreeding, selfing vs outcrossing rates, effective population sizes and the dynamics of recent population bottlenecks. Beyond providing basic evolutionary insights, such analyses are also an important tool for developing effective management strategies for endangered and/or invasive species (Hedrick, 2001; Sakai et al., 2001). Owing to their codominant and highly polymorphic nature, simple-sequence repeats (SSRs, a.k.a. microsatellites) have increasingly become the marker of choice for these sorts of analyses.
Unfortunately, the de novo development of SSRs is a costly and time-consuming endeavor (Zane et al., 2002; Squirrell et al., 2003), and these problems are often compounded by a paucity of resources in taxa that lack clear economic importance. Adding to this difficulty is the fact that the polymerase chain reaction primers used to amplify SSRs are frequently species-specific, meaning that markers developed in one taxon cannot be readily transferred to another. Beyond reducing the general utility of existing markers, this lack of transferability also means that interspecific comparisons are often based on disparate sets of markers, effectively confounding species differences with possible locus-specific effects.
One possible solution to these sorts of problems would be to exploit publicly available genomic resources for the development of gene-based SSR markers that are more likely to be transferable across taxonomic boundaries. In fact, the rapid and inexpensive development of SSRs from expressed sequence tag (EST) databases has been shown to be a feasible option for obtaining high-quality nuclear markers (Gupta et al., 2003; Bhat et al., 2005). Moreover, the National Center for Biotechnology Information (NCBI) EST database (dbEST; Boguski et al., 1993) contains an ever-increasing number of these ‘single-pass’ cDNA sequences, meaning that the resources necessary for the efficient development of large numbers of so-called EST-SSRs already exist for a wide variety of taxa.
In general, EST-SSRs have been found to be significantly more transferable across taxonomic boundaries than are traditional ‘anonymous’ SSRs (Chagne et al., 2004; Liewlaksaneeyanawin et al., 2004; Gutierrez et al., 2005; Pashley et al., 2006), and reports of EST-SSR transferability have become increasingly common. This is particularly true in plants, where transferability among economically important crop taxa has been demonstrated on a number of occasions (Decroocq et al., 2003; Thiel et al., 2003; Bandopadhyay et al., 2004; Saha et al., 2004; Varshney et al., 2005b). Until recently, however, little attention had been paid to the potential for transferring EST-SSRs from well-characterized taxa to lesser studied relatives as a means for facilitating evolutionary analyses (but see Arnold et al., 2002; Ellis et al., 2006).
In the present paper, we provide an overview of what is known about EST-SSR transferability with particular reference to the potential for these markers to facilitate population genetic analyses of previously understudied plant taxa. As an example of the utility of these resources, we further cross-reference existing EST databases against the US Fish and Wildlife Service threatened and endangered species database, the 2006 IUCN Red List of threatened species and the US State and Federal Composite List of Noxious Weeds to determine the extent of overlap between suitable EST databases and plant species of conservation concern. Finally, we provide a discussion of the advantages and disadvantages of EST-SSRs in the context of population genetic applications.
Transferability and polymorphism of EST-SSRs
Perhaps the most common applications of EST-SSRs to date have involved analyses of functional diversity, genetic mapping and/or marker-assisted selection in crop species (reviewed by Varshney et al., 2005a). To the extent that these markers are transferable across taxa, however, EST-SSRs also have clear potential for use in basic evolutionary applications, such as population genetic analyses (Ellis et al., 2006). In this section, we provide an overview of what is known about EST-SSR transferability in plants.
As noted in the Introduction, the ability to effectively transfer polymorphic EST-SSRs across taxa has now been demonstrated in a number of cases, most commonly in studies involving economically important crop species (Table 1). Taken together, the results of these studies indicate that EST-SSRs can often be transferred across relatively large taxonomic distances, spanning not just species within a genus, but in some instances multiple genera within a family. For example, Scott et al. (2000) tested the transferability of 10 Vitis EST-SSRs among grape cultivars, other grape species and related genera, and found high levels of transferability, with over 60% of markers tested working across taxa. Moreover, all of the transferable markers proved to be polymorphic at the level of cultivars, Vitis species and between related genera. Similarly, Decroocq et al. (2003) used EST-SSRs developed from grape and apricot sequences to investigate transferability across 46 related grape species and 29 members of the Rosaceae. Overall, the grape primers were transferable to, and revealed polymorphisms within, most Vitaceae accessions tested. In contrast, the apricot primers were found to be most useful within the subgenus Prunophora. In the cereals, Gupta et al. (2003) demonstrated extensive transferability of Triticum aestivum L. (bread wheat) EST-SSRs to 18 related wild species in the Triticum–Aegilops complex and to five cereal species of barley, oat, rye, rice and maize. Over 80% of primer pairs tested were transferable to the 18 related species, while nearly 60% showed transferability to one or more of the more distantly related cereal species. In other grass species, EST-SSRs from the turf grass Festuca arundinacea Schreb. (tall fescue) were tested for transferability in seven grasses from four genera varying in mating system and ploidy level (Saha et al., 2004). This work revealed greater than 90% transferability to one or more of the other species tested. Moreover, the surveyed loci revealed ample levels of polymorphism for elucidating relationships among these species. Finally, high levels transferability and substantial polymorphism were observed among 23 cotton (Gossypium) species (Guo et al., 2006).
In general terms, this sort of transferability is unique to EST-SSRs, with anonymous SSRs being significantly less portable (Chagne et al., 2004; Liewlaksaneeyanawin et al., 2004; Gutierrez et al., 2005; Pashley et al., 2006; but see Fitzsimmons et al., 1995; Dayanandan et al., 1997). EST-SSRs have also been shown to produce substantially ‘cleaner’ data (that is, easier to analyze/interpret amplification profiles) as compared to their anonymous counterparts (Pashley et al., 2006). There is also evidence that EST-SSRs located in coding regions are significantly more transferable than those found in untranslated regions (UTRs) (Pashley et al., 2006). Despite their potential to cause selectively deleterious frameshift mutations, however, EST-SSRs located in coding regions appear to reveal equivalent levels of polymorphism as compared to those located in UTRs, most likely due to a general trend toward trinucleotide repeats in coding regions. In fact, this trend toward trinucleotide repeats in exons has been observed in a variety of other taxa, including wheat (Gupta et al., 2003) barrel medic (Eujayl et al., 2004), tall fescue (Saha et al., 2004), and pine (Chagne et al., 2004). Regardless of the cause, if this observed tendency toward higher transferability and equivalent levels of polymorphism turns out to be a general feature of EST-SSRs located in protein-coding regions, the targeting of exonic trinucleotide repeat motifs might be the best strategy for developing portable sets of polymorphic EST-SSR markers.
EST resources and SSR frequencies
Although EST-SSR transferability has now been documented in a number of cases, the utility of these sorts of markers for facilitating evolutionary genetic research in non-target taxa (that is, taxa that lack genomic resources) has received relatively little attention. However, when the generally high transferability of EST-SSRs is combined with the fact that population genetic analyses often rely on a relatively small number of markers (Richards et al., 2004; Vornam et al., 2004; Szczys et al., 2005), it seems likely that even modest EST collections could prove to be of great value to evolutionary biologists. In fact, an estimated 2–5% of all plant-derived ESTs are thought to harbor SSRs (Kantety et al., 2002), although the actual frequency of SSR-bearing ESTs in any particular analysis is highly dependent on the search parameters (see below). Moreover, a large fraction of EST-SSRs (on the order of 80–90%) are typically found to be polymorphic (Bandopadhyay et al., 2004; Fraser et al., 2004; Pashley et al., 2006). Taking into account typical marker development attrition rates, it therefore seems likely that EST databases containing as few as 1000 sequences could provide enough markers to facilitate population genetic analyses.
To highlight the potential utility of such resources, we surveyed available EST collections and cross-referenced them against several databases that list either rare/endangered or invasive plant species. As of May 2006, the NCBI EST database (dbEST) contained over 36 million publicly available EST sequences from over 1100 taxa. Of these, 542 taxa accounted for greater than 1000 EST sequences apiece, including 211 different spermatophytes (that is, seed plants), representing 126 unique genera. The taxonomic databases that we cross-referenced these sequence collections against included the US Fish and Wildlife Service threatened and endangered species database (http://www.fws.gov/endangered/wildlife.html), the 2006 IUCN Red List of threatened species (http://www.redlist.org/), and the US State and Federal Composite List of Noxious Weeds (http://plants.usda.gov/).
At the time of our survey, the US Fish and Wildlife list of threatened and endangered species contained 51 plant species (representing 25 different genera) that were congeneric with at least one species for which there were ⩾1000 publicly available ESTs (Table 2). The IUCN Red List contained an additional 576 species from 45 genera that had congeners with similar EST resources. Turning to the US Composite List of Noxious Weeds, 80 species from 21 genera had at least one congener with ⩾1000 publicly available ESTs. In some cases, the source taxa for the ESTs were themselves either endangered or invasive; these species were excluded from the tallies, as noted in Table 2. In a handful of cases, the invasive species list cited only a genus name without specific epithet (for example, Vitis L.). Such instances were included in our tabulation, but only counted as a single taxon.
After accounting for overlap across lists, we found that half (68/136) of all plant-derived EST collections of sufficient size (that is, ⩾1000 sequences) could potentially serve as a source of EST-SSRs for the genetic analysis of rare, endangered or invasive plants species worldwide (Table 2). It is important to note here that this is most likely a somewhat conservative estimate, as: (1) this survey was primarily based on data from US agencies, although we did include the most critically endangered species from elsewhere, and (2) only those EST collections that were derived from a congener of the focal species were included in the tally. As noted above, EST-SSRs are also often transferable across greater taxonomic distances; for example, Rossetto (2001) found that the average rate of intergeneric transfer was ca. 35% in a variety of plant taxa. It should also be kept in mind that, while rare and invasive plants were chosen to illustrate the likely utility of existing EST resources for population genetic analyses, these resources have the potential to facilitate evolutionary research in a much wider variety of taxa.
In order to better gauge the utility of the smallest EST collections identified above, we surveyed all data sets consisting of 1000–10 000 ESTs for the presence of unique SSRs (Table 3). We did this by first downloading from dbEST all ESTs for each genus that showed overlap with one or more taxa of conservation interest. We then assembled them using CAP3 (Huang and Madan, 1999) and analyzed the resulting unigene set for each genus using SSRIT (Temnykh et al., 2001; http://www.gramene.org/db/searches/ssrtool), which is a perl script that identifies all SSRs within a set of sequences. We set the script to identify all possible di-, tri- and tetranucleotide repeats with a minimum of five, four and three subunits, respectively. While some researchers have employed higher cutoffs (Kantety et al., 2002), relaxing the thresholds maximizes SSR discovery while still producing a high percentage of polymorphic loci (Pashley et al., 2006).
Inspection of Table 3 reveals that nearly one in 10 unique ESTs (9.0±0.1%; mean±s.e.) contained at least one SSR (range=2.5–21.1%). Thus, it seems reasonable to assume that EST collections consisting of 1000 or more sequences have the potential to provide ample candidate SSRs for use in conservation genetic analyses. This is especially true in view of the potentially high percentage of EST-SSRs that turn out to be polymorphic (Bandopadhyay et al., 2004; Fraser et al., 2004; Pashley et al., 2006).
Prospects and pitfalls
As noted at the outset, the codominant and highly polymorphic nature of SSRs has increasingly made them the marker of choice for population genetics analyses. Unfortunately, the development of traditional ‘anonymous’ SSRs requires a substantial investment of both time and money, putting them out of reach for many researchers. Given that EST-based SSRs can be developed directly from existing sequence resources and can often be transferred from one species to another, EST databases are an attractive source of markers for the genetic analysis of understudied taxa.
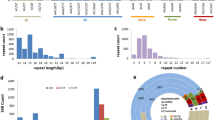
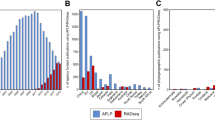
Looking beyond the relative ease with which EST-SSRs can be developed, one of their clearest advantages is that they allow one to make direct comparisons among taxa without running the risk that locus-specific differences might mask true species-level differences in things like overall levels of genetic diversity, the extent of population structure, so on. For example, Ellis et al. (2006) used EST-SSRs derived from the cultivated sunflower (Helianthus annuus L.) to investigate levels of genetic diversity in an extremely rare sunflower (H. verticillatus) and a more common congener (H. angustifolius). Based on a simple comparison of the mean level of genetic diversity present within each taxon, the two species are statistically indistinguishable. After controlling for inherent differences in variability from one locus to the next, however, it becomes clear that H. verticillatus actually harbors more genetic diversity than does H. angustifolius despite its rarity (Figure 1). Beyond providing more statistical power in paired comparisons, EST-SSRs also produce cleaner results for scoring as there are fewer null alleles (Leigh et al., 2003; Rungis et al., 2004) and fewer stutter bands (Leigh et al., 2003; Woodhead et al., 2003; Eujayl et al., 2004; Pashley et al., 2006). Despite these advantages, however, EST-SSRs are not without their drawbacks.
Comparison of genetic diversity in H. verticillatus and H. angustifolius. Each point represents one of the 19 loci that the two species have in common, and the lines connect data points derived from an individual locus (Ellis et al., 2006). Note that, although the diversity estimates overlap broadly between species, there is a clear tendency toward decreased genetic diversity in H. angustifolius when viewed on a per-locus basis, with 13 of 19 loci showing clear evidence of a decline.
One concern with SSRs in general is the possibility of null alleles, which fail to amplify due to primer site variation, and thus do not produce a visible amplicon. Individuals that are heterozygous for a null allele appear to be homozygous for the visible allele, whereas null homozygotes appear to be failed reactions. When present in a population, null alleles will bias allele frequencies, reduce the observed heterozygosity, and therefore increase apparent levels of inbreeding (DeWoody et al., 2006). While EST-SSRs are subject to these sorts of difficulties, the same can be said of anonymous SSRs. Moreover, the primers flanking EST-SSRs are derived from relatively conserved sequences; therefore, it is likely that null alleles will be less of a problem for EST-SSRs as compared to their anonymous counterparts. Indeed, Rungis et al. (2004) found that measures of inbreeding were significantly lower in EST-SSRs versus genomic SSRs in spruce, and they suggested that this resulted from a lower frequency of null alleles in the former.
Because the cDNAs from which ESTs are derived lack introns, one possible concern with EST-SSRs is that unrecognized intron splice sites could disrupt priming sites, resulting in failed amplification. Alternatively, large introns could fall between the primers, resulting in a product that is either too large or, in extreme cases, failed amplification. Fortunately, intron locations are relatively well-conserved across taxa (Strand et al., 1997; Ku et al., 2000; Wu et al., 2006). Thus, it is possible to minimize this sort of problem by aligning ESTs of interest against the genomic sequence of model species such as Arabidopsis or rice. Putative intron positions can then be noted, and primers can be designed accordingly. Of course, this is not a perfect solution, as intron gain and loss are still distinct possibilities. In some cases, however, it may be possible to redesign the primers to exclude troublesome introns.
Another obvious concern is that since EST-SSRs are located within genes, and thus more conserved across species, they may be less polymorphic than anonymous SSRs. This concern has been borne out in a number of taxa, including rice (Cho et al., 2000), bread wheat (Gupta et al., 2003), pines (Liewlaksaneeyanawin et al., 2004), barley (Chabane et al., 2005) and sunflower (Pashley et al., 2006). However, the levels of genetic diversity revealed by these markers are still considerably higher than those revealed by most alternative marker types, such as allozymes (Hamrick and Godt, 1996). Thus, even though EST-SSRs reveal less variability than do anonymous SSRs, these markers still reveal sufficient levels of variation for the vast majority of population genetic applications.
Perhaps the greatest concern with regard to the utility of EST-SSRs in the present context is that selection on these loci might influence the estimation of population genetic parameters. Indeed, divergent selection will increase differentiation among and reduce variability within populations, whereas balancing selection will have the opposite effect. While a recent study by Woodhead et al. (2005) revealed that estimates of population differentiation based on EST-SSRs are comparable to those based on both anonymous SSRs and AFLPs in ferns, and large-scale comparative analyses suggest that only a very small percentage of all genes are experiencing positive selection (Tiffin and Hahn, 2002; Clark et al., 2003), some small fraction of all EST-SSRs will inevitably be subject to selection. Indeed, there are examples from the literature wherein certain genic SSRs are known to be associated with various diseases in animals (Zoghbi and Orr, 2000; Mao et al., 2002; Yamada et al., 2002) or pathogenicity/virulence in microbes (Peak et al., 1996; Grimwood et al., 2001). While more studies are needed before we will have a better understanding of the possible effects of genic SSRs in plants (Li et al., 2004), it seems safe to assume that at least a small percentage of loci will be evolving in a non-neutral manner. It remains unclear, however, whether this problem will be more or less frequent than in other gene-based marker systems, such as allozymes.
There are, of course, a number of potential applications of EST-SSRs that will be less sensitive to the effects of selection. For example, single-generation applications such as paternity studies, mating system analyses and direct estimates of gene flow will be relatively robust to deviations from neutrality. In the case of analyses that rely on equilibrium assumptions, such as studies focusing on population structure and/or indirect estimates of gene flow, the effects of selection can best be minimized by increasing the number of markers utilized, so as to reduce the potential biases introduced by any one locus, and by endeavoring to employ a common set of markers across taxa when working in a comparative manner. Assuming that a sufficiently large number of markers are employed, it should also be possible to statistically identify and exclude loci with extreme FST values (Wright's (1951) measure of population genetic structure), as such outliers are likely the result of selection (Lewontin and Krakauer, 1973; Beaumont and Nichols, 1996).
Conclusions
The advent of the genomics age has resulted in the production of an ever-expanding body of DNA sequence data, including vast EST collections. These ESTs represent a potentially valuable source of gene-based SSR markers for population genetic analyses. While EST-SSRs are not without their drawbacks, they offer a number of clear benefits, including rapid and inexpensive development and high levels of cross-taxon portability. Thus, EST-SSRs have the potential to facilitate evolutionary analyses in a wide variety of taxa, and may well represent the best way forward for the analysis of species for which only limited resources are available.
References
Arnold C, Rossetto M, McNally J, Henry RJ (2002). The application of SSRs characterized for grape (Vitis vinifera) to conservation studies in Vitaceae. Am J Bot 89: 22–28.
Bandopadhyay R, Sharma S, Rustgi S, Singh R, Kumar A, Balyan HS et al. (2004). DNA polymorphism among 18 species of Triticum–Aegilops complex using wheat EST-SSRs. Plant Sci 166: 349–356.
Beaumont MA, Nichols RA (1996). Evaluating loci for use in the genetic analysis of population structure. Proc R Soc Lond B 263: 1619–1626.
Bhat PR, Krishnakumar V, Hendre PS, Rajendrakumar P, Varshney RK, Aggarwal RK (2005). Identification and characterization of expressed sequence tags-derived simple sequence repeats, markers from robusta coffee variety ‘C × R’ (an interspecific hybrid of Coffea canephora × Coffea congensis). Mol Ecol Notes 5: 80–83.
Boguski MS, Lowe TMJ, Tolstoshev CM (1993). dbEST – database for expressed sequence tags. Nat Genet 4: 332–333.
Chabane K, Ablett GA, Cordeiro GM, Valkoun J, Henry RJ (2005). EST versus genomic derived microsatellite markers for genotyping wild and cultivated barley. Genet Resour Crop Evol 52: 903–909.
Chagne D, Chaumeil P, Ramboer A, Collada C, Guevara A, Cervera MT et al. (2004). Cross-species transferability and mapping of genomic and cDNA SSRs in pines. Theor Appl Genet 109: 1204–1214.
Cho YG, Ishii T, Temnykh S, Chen X, Lipovich L, McCouch SR et al. (2000). Diversity of microsatellites derived from genomic libraries and GenBank sequences in rice (Oryza sativa L.). Theor Appl Genet 100: 713–722.
Clark AG, Glanowski S, Nielsen R, Thomas PD, Kejariwal A, Todd MA et al. (2003). Inferring non-neutral evolution from human-chimp-mouse orthologous gene trios. Science 302: 1960–1963.
Cordeiro GM, Casu R, McIntyre CL, Manners JM, Henry RJ (2001). Microsatellite markers from sugarcane (Saccharum spp.) ESTs cross transferable to erianthus and sorghum. Plant Sci 160: 1115–1123.
Dayanandan S, Bawa KS, Kesseli R (1997). Conservation of microsatellites among tropical trees (Leguminosae). Am J Bot 84: 1658–1663.
Decroocq V, Fave MG, Hagen L, Bordenave L, Decroocq S (2003). Development and transferability of apricot and grape EST microsatellite markers across taxa. Theor Appl Genet 106: 912–922.
DeWoody J, Nason JD, Hipkins VD (2006). Mitigating scoring errors in microsatellite data from wild populations. Mol Ecol Notes 6: 951–957.
Ellis JR, Pashley CH, Burke JM, McCauley DE (2006). High genetic diversity in a rare and endangered sunflower as compared to a common congener. Mol Ecol 15: 2345–2355.
Eujayl I, Sledge MK, Wang L, May GD, Chekhovskiy K, Zwonitzer JC et al. (2004). Medicago truncatula EST-SSRs reveal cross-species genetic markers for Medicago spp. Theor Appl Genet 108: 414–422.
Fitzsimmons NN, Moritz C, Moore SS (1995). Conservation and dynamics of microsatellite loci over 300-million years of marine turtle evolution. Mol Biol Evol 12: 432–440.
Fraser LG, Harvey CF, Crowhurst RN, De Silva HN (2004). EST-derived microsatellites from Actinidia species and their potential for mapping. Theor Appl Genet 108: 1010–1016.
Grimwood J, Olinger L, Stephens RS (2001). Expression of Chlamydia pneumoniae polymorphic membrane protein family genes. Infect Immun 69: 2383–2389.
Guo WZ, Wang W, Zhou BL, Zhang TZ (2006). Cross-species transferability of G. arboreum-derived EST-SSRs in the diploid species of Gossypium. Theor Appl Genet 112: 1573–1581.
Gupta PK, Rustgi S, Sharma S, Singh R, Kumar N, Balyan HS (2003). Transferable EST-SSR markers for the study of polymorphism and genetic diversity in bread wheat. Mol Genet Genom 270: 315–323.
Gutierrez MV, Patto MCV, Huguet T, Cubero JI, Moreno MT, Torres AM (2005). Cross-species amplification of Medicago truncatula microsatellites across three major pulse crops. Theor Appl Genet 110: 1210–1217.
Hamrick JL, Godt MJW (1996). Conservation genetics of endemic plant species. In: Avise JC and Hamrick JL (eds). Conservation Genetics. Case Histories From Nature. Chapman and Hall: New York, NY. pp 281–304.
Hedrick PW (2001). Conservation genetics: where are we now? Trends Ecol Evol 16: 629–636.
Huang XQ, Madan A (1999). CAP3: A DNA sequence assembly program. Genome Res 9: 868–877.
Kantety RV, La Rota M, Matthews DE, Sorrells ME (2002). Data mining for simple sequence repeats in expressed sequence tags from barley, maize, rice, sorghum and wheat. Plant Mol Biol 48: 501–510.
Ku HM, Vision T, Liu J, Tanksley SD (2000). Comparing sequenced segments of the tomato and Arabidopsis genomes: large-scale duplication followed by selective gene loss creates a network of synteny. Proc Natl Acad Sci USA 97: 9121–9126.
Leigh F, Lea V, Law J, Wolters P, Powell W, Donini P (2003). Assessment of EST- and genomic microsatellite markers for variety discrimination and genetic diversity studies in wheat. Euphytica 133: 359–366.
Lewontin RC, Krakauer J (1973). Distribution of gene frequency as a test of the theory of the selective neutrality of polymorphisms. Genetics 74: 175–195.
Li YC, Korol AB, Fahima T, Nevo E (2004). Microsatellites within genes: structure, function, and evolution. Mol Biol Evol 21: 991–1007.
Liewlaksaneeyanawin C, Ritland CE, El-Kassaby YA, Ritland K (2004). Single-copy, species-transferable microsatellite markers developed from loblolly pine ESTs. Theor Appl Genet 109: 361–369.
Mao R, Aylsworth AS, Potter N, Wilson WG, Breningstall G, Wick MJ et al. (2002). Childhood-onset ataxia: testing for large CAG-repeats in SCA2 and SCA7. Am J Med Genet 110: 338–345.
Pashley CH, Ellis JR, McCauley DE, Burke JM (2006). EST databases as a source for molecular markers: lessons from Helianthus. J Hered 97: 381–388.
Peak IRA, Jennings MP, Hood DW, Bisercic M, Moxon ER (1996). Tetrameric repeat units associated with virulence factor phase variation in Haemophilus also occur in Neisseria spp and Moraxella catarrhalis. FEMS Microbiol Lett 137: 109–114.
Richards CM, Reilley A, Touchell D, Antolin MF, Walters C (2004). Microsatellite primers for Texas wild rice (Zizania texana), and a preliminary test of the impact of cryogenic storage on allele frequency at these loci. Conser Genet 5: 853–859.
Rossetto M (2001). Sourcing of SSR markers from related plant species. In: Henry RJ (ed). Plant Genotyping: the DNA Fingerprinting of Plants. CABI Publishing: Wallingford. pp 211–224.
Rungis D, Berube Y, Zhang J, Ralph S, Ritland CE, Ellis BE et al. (2004). Robust simple sequence repeat markers for spruce (Picea spp.) from expressed sequence tags. Theor Appl Genet 109: 1283–1294.
Saha MC, Mian MAR, Eujayl I, Zwonitzer JC, Wang LJ, May GD (2004). Tall fescue EST-SSR markers with transferability across several grass species. Theor Appl Genet 109: 783–791.
Sakai AK, Allendorf FW, Holt JS, Lodge DM, Molofsky J, With KA et al. (2001). The population biology of invasive species. Annu Rev Ecol Syst 32: 305–332.
Scott KD, Eggler P, Seaton G, Rossetto M, Ablett EM, Lee LS et al. (2000). Analysis of SSRs derived from grape ESTs. Theor Appl Genet 100: 723–726.
Squirrell J, Hollingsworth PM, Woodhead M, Russell J, Lowe AJ, Gibby M et al. (2003). How much effort is required to isolate nuclear microsatellites from plants? Mol Ecol 12: 1339–1348.
Strand AE, Leebens-Mack J, Milligan BG (1997). Nuclear DNA-based markers for plant evolutionary biology. Mol Ecol 6: 113–118.
Szczys P, Hughes CR, Kessel RV (2005). Novel microsatellite markers used to determine the population genetic structure of the endangered Roseate Tern, Sterna dougallii, in Northwest Atlantic and Western Australia. Conser Genet 6: 461–466.
Temnykh S, DeClerck G, Lukashova A, Lipovich L, Cartinhour S, McCouch S (2001). Computational and experimental analysis of microsatellites in rice (Oryza sativa L.): frequency, length variation, transposon associations, and genetic marker potential. Genome Res 11: 1441–1452.
Thiel T, Michalek W, Varshney RK, Graner A (2003). Exploiting EST databases for the development and characterization of gene-derived SSR-markers in barley (Hordeum vulgare L.). Theor Appl Genet 106: 411–422.
Tiffin P, Hahn MW (2002). Coding sequence divergence between two closely related plant species: Arabidopsis thaliana and Brassica rapa ssp pekinensis. J Mol Evol 54: 746–753.
Varshney RK, Graner A, Sorrells ME (2005a). Genic microsatellite markers in plants: features and applications. Trends Biotechnol 23: 48–55.
Varshney RK, Sigmund R, Borner A, Korzun V, Stein N, Sorrells ME et al. (2005b). Interspecific transferability and comparative mapping of barley EST-SSR markers in wheat, rye and rice. Plant Sci 168: 195–202.
Vornam B, Decarli N, Gailing O (2004). Spatial distribution of genetic variation in a natural beech stand (Fagus sylvatica L.) based on microsatellite markers. Conser Genet 5: 561–570.
Woodhead M, Russell J, Squirrell J, Hollingsworth M, Cardle L, Ramsay L et al. (2003). Development of EST-SSRs from the alpine lady-fern, Athyrium distentifolium. Mol Ecol Notes 3: 287–290.
Woodhead M, Russell J, Squirrell J, Hollingsworth PM, Mackenzie K, Gibby M et al. (2005). Comparative analysis of population genetic structure in Athyrium distentifolium (Pteridophyta) using AFLPs and SSRs from anonymous and transcribed gene regions. Mol Ecol 14: 1681–1695.
Wright S (1951). The genetical structure of populations. Ann Eugen 15: 323–354.
Wu F, Mueller LA, Crouzillat D, Pétiard V, Tanksley SD (2006). Combining bioinformatics and phylogenetics to identify large sets of single-copy orthologous genes (COSII) for comparative, evolutionary and systematic studies: a test case in the Euasterid plant clade. Genetics 174: 1407–1420.
Yamada M, Tsuji S, Takahashi H (2002). Involvement of lysosomes in the pathogenesis of CAG repeat diseases. Ann Neurol 52: 498–503.
Zane L, Bargelloni L, Patarnello T (2002). Strategies for microsatellite isolation: a review. Mol Ecol 11: 1–16.
Zoghbi HY, Orr HT (2000). Glutamine repeats and neurodegeneration. Annu Rev Neurosci 23: 217–247.
Acknowledgements
We thank Catherine Pashley for her contributions to Table 1 and Mark Chapman, David McCauley, Aizhong Liu, Natasha Sherman and David Wills for comments on earlier versions of this paper. The writing of this paper was supported in part by grants from the National Science Foundation (DBI-0332411) and the National Research Initiative of the USDA Cooperative State Research, Education and Extension Service (03-35300-13104).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Ellis, J., Burke, J. EST-SSRs as a resource for population genetic analyses. Heredity 99, 125–132 (2007). https://doi.org/10.1038/sj.hdy.6801001
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.hdy.6801001
Keywords
This article is cited by
-
Genetic diversity of Thailand reserved mulberry germplasm based on morphological characteristics and newly developed EST-SSR and SRAP markers
Horticulture, Environment, and Biotechnology (2024)
-
Assessing genetic diversity of European white elm (Ulmus laevis Pallas) in Bavaria as an indicator for in-situ conservation and sustainable use of the species genetic resources
European Journal of Forest Research (2023)
-
Applying molecular and genetic methods to trees and their fungal communities
Applied Microbiology and Biotechnology (2023)
-
Genetic diversity and population structure in wild seabuckthorn (Hippophae salicifolia D. Don) growing in the Uttarakhand region of the Indian Himalayas
Genetic Resources and Crop Evolution (2023)
-
Genome survey sequencing-based SSR marker development and their validation in Dendrocalamus longispathus
Functional & Integrative Genomics (2023)