Abstract
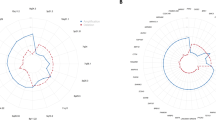
We analysed 148 primary breast cancers using BAC-arrays containing 287 clones representing cancer-related gene/loci to obtain genomic molecular portraits. Gains were detected in 136 tumors (91.9%) and losses in 123 tumors (83.1%). Eight tumors (5.4%) did not have any genomic aberrations in the 281 clones analysed. Common (more than 15% of the samples) gains were observed at 8q11–qtel, 1q21–qtel, 17q11–q12 and 11q13, whereas common losses were observed at 16q12–qtel, 11ptel–p15.5, 1p36–ptel, 17p11.2–p12 and 8ptel–p22. Patients with tumors registering either less than 5% (median value) or less than 11% (third quartile) total copy number changes had a better overall survival (log-rank test: P=0.0417 and P=0.0375, respectively). Unsupervised hierarchical clustering based on copy number changes identified four clusters. Women with tumors from the cluster with amplification of three regions containing known breast oncogenes (11q13, 17q12 and 20q13) had a worse prognosis. The good prognosis group (Nottingham Prognostic Index (NPI) ⩽3.4) tumors had frequent loss of 16q24–qtel. Genes significantly associated with estrogen receptor (ER), Grade and NPI were used to build k-nearest neighbor (KNN) classifiers that predicted ER, Grade and NPI status in the test set with an average misclassification rate of 24.7, 25.7 and 35.7%, respectively. These data raise the prospect of generating a molecular taxonomy of breast cancer based on copy number profiling using tumor DNA, which may be more generally applicable than expression microarray analysis.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Abbreviations
- NPI:
-
Nottingham Prognostic Index
- ER:
-
estrogen receptor
- HR:
-
hazard ratio
References
Al-Kuraya K, Schraml P, Torhorst J, Tapia C, Zaharieva B, Novotny H et al. (2004). Prognostic relevance of gene amplifications and coamplifications in breast cancer. Cancer Res 64: 8534–8540.
Bogliolo M, Cabre O, Callen E, Castillo V, Creus A, Marcos R et al. (2002). The Fanconi anaemia genome stability and tumour suppressor network. Mutagenesis 17: 529–538.
Callagy G, Pharoah P, Chin SF, Sangan T, Daigo Y, Jackson L et al. (2005). Identification and validation of prognostic markers in breast cancer with the complementary use of array-CGH and tissue microarrays. J Pathol 205: 388–396.
Chin SF, Wang Q, Puisieux A, Caldas C . (2001). Absence of rearrangements in the BRCA2 gene in human cancers. Br J Cancer 84: 193–195.
Cingoz S, Altungoz O, Canda T, Saydam S, Aksakoglu G, Sakizli M . (2003). DNA copy number changes detected by comparative genomic hybridization and their association with clinicopathologic parameters in breast tumors. Cancer Genet Cytogenet 145: 108–114.
Daigo Y, Chin SF, Gorringe KL, Bobrow LG, Ponder BA, Pharoah PD et al. (2001). Degenerate oligonucleotide primed-polymerase chain reaction-based array comparative genomic hybridization for extensive amplicon profiling of breast cancers: a new approach for the molecular analysis of paraffin-embedded cancer tissue. Am J Pathol 158: 1623–1631.
Elston CW, Ellis IO . (1991). Pathological prognostic factors in breast cancer. I. The value of histological grade in breast cancer: experience from a large study with long-term follow-up. Histopathology 19: 403–410.
Forozan F, Karhu R, Kononen J, Kallioniemi A, Kallioniemi OP . (1997). Genome screening by comparative genomic hybridization. Trends Genet 13: 405–409.
Fridlyand J, Snijders AM, Ylstra B, Li H, Olshen A, Segraves R et al. (2006). Breast tumor copy number aberration phenotypes and genomic instability. BMC Cancer. 6: 96.
Gilchrist KW, Kalish L, Gould VE, Hirschl S, Imbriglia JE, Levy WM et al. (1985). Interobserver reproducibility of histopathological features in stage II breast cancer. An ECOG study. Breast Cancer Res Treat 5: 3–10.
Glinsky GV, Higashiyama T, Glinskii AB . (2004). Classification of human breast cancer using gene expression profiling as a component of the survival predictor algorithm. Clin Cancer Res 10: 2272–2283.
Gunther K, Merkelbach-Bruse S, Amo-Takyi BK, Handt S, Schroder W, Tietze L . (2001). Differences in genetic alterations between primary lobular and ductal breast cancers detected by comparative genomic hybridization. J Pathol 193: 40–47.
Heidenblad M, Lindgren D, Veltman JA, Jonson T, Mahlamaki EH, Gorunova L et al. (2005). Microarray analyses reveal strong influence of DNA copy number alterations on the transcriptional patterns in pancreatic cancer: implications for the interpretation of genomic amplifications. Oncogene 24: 1794–1801.
Hodgson G, Hager JH, Volik S, Hariono S, Wernick M, Moore D et al. (2001). Genome scanning with array CGH delineates regional alterations in mouse islet carcinomas. Nat Genet 29: 459–464.
James LA, Mitchell EL, Menasce L, Varley JM . (1997). Comparative genomic hybridisation of ductal carcinoma in situ of the breast: identification of regions of DNA amplification and deletion in common with invasive breast carcinoma. Oncogene 14: 1059–1065.
Jones C, Ford E, Gillett C, Ryder K, Merrett S, Reis-Filho JS et al. (2004). Molecular cytogenetic identification of subgroups of grade III invasive ductal breast carcinomas with different clinical outcomes. Clin Cancer Res 10: 5988–5997.
Kallioniemi A, Kallioniemi OP, Piper J, Tanner M, Stokke T, Chen L et al. (1994). Detection and mapping of amplified DNA sequences in breast cancer by comparative genomic hybridization. Proc Natl Acad Sci USA 91: 2156–2160.
Kim YH, Girard L, Giacomini CP, Wang P, Hernandez-Boussard T, Tibshirani R et al. (2005). Combined microarray analysis of small cell lung cancer reveals altered apoptotic balance and distinct expression signatures of MYC family gene amplification. Oncogene 25: 130–138.
Korsching E, Packeisen J, Helms MW, Kersting C, Voss R, van Diest PJ et al. (2004). Deciphering a subgroup of breast carcinomas with putative progression of grade during carcinogenesis revealed by comparative genomic hybridisation (CGH) and immunohistochemistry. Br J Cancer 90: 1422–1428.
Landi MT, Kanetsky PA, Tsang S, Gold B, Munroe D, Rebbeck T et al. (2005). MC1R, ASIP, and DNA repair in sporadic and familial melanoma in a Mediterranean population. J Natl Cancer Inst 97: 998–1007.
Loo LW, Grove DI, Williams EM, Neal CL, Cousens LA, Schubert EL et al. (2004). Array comparative genomic hybridization analysis of genomic alterations in breast cancer subtypes. Cancer Res 64: 8541–8549.
Maass N, Rosel F, Schem C, Hitomi J, Jonat W, Nagasaki K . (2002). Amplification of the BCAS2 gene at chromosome 1p13.3–21 in human primary breast cancer. Cancer Lett 185: 219–223.
McShane LM, Radmacher MD, Freidlin B, Yu R, Li MC, Simon R . (2002). Methods for assessing reproducibility of clustering patterns observed in analyses of microarray data. Bioinformatics 18: 1462–1469.
Naylor TL, Greshock J, Wang Y, Colligon T, Yu QC, Clemmer V et al. (2005). High resolution genomic analysis of sporadic breast cancer using array-based comparative genomic hybridization. Breast Cancer Res 7: R1186–R1198.
Nessling M, Richter K, Schwaenen C, Roerig P, Wrobel G, Wessendorf S et al. (2005). Candidate genes in breast cancer revealed by microarray-based comparative genomic hybridization of archived tissue. Cancer Res 65: 439–447.
Nupponen NN, Isola J, Visakorpi T . (2000). Mapping the amplification of EIF3S3 in breast and prostate cancer. Genes Chromosomes Cancer 28: 203–210.
Perou CM, Sorlie T, Eisen MB, van de Rijn M, Jeffrey SS, Rees CA et al. (2000). Molecular portraits of human breast tumours. Nature 406: 747–752.
Pinkel D, Segraves R, Sudar D, Clark S, Poole I, Kowbel D et al. (1998). High resolution analysis of DNA copy number variation using comparative genomic hybridization to microarrays. Nat Genet 20: 207–211.
Piper J, Stegenga S, Pestova E, Marble H, Lucas M, Wilber K et al. (2002). An objective method for detecting copy number change in CGH microarray experiments. Proceedings of the Third Euroconference on Quantitative Molecular Cytogenetics. Rosenon, Stockholm, Sweden 4–6 July 2002; Rosenon, Stockholm, Sweden, pp 109–114.
Pollack JR, Sorlie T, Perou CM, Rees CA, Jeffrey SS, Lonning PE et al. (2002). Microarray analysis reveals a major direct role of DNA copy number alteration in the transcriptional program of human breast tumors. Proc Natl Acad Sci USA 99: 12963–12968.
Rennstam K, Ahlstedt-Soini M, Baldetorp B, Bendahl PO, Borg A, Karhu R et al. (2003). Patterns of chromosomal imbalances defines subgroups of breast cancer with distinct clinical features and prognosis. A study of 305 tumors by comparative genomic hybridization. Cancer Res 63: 8861–8868.
Reyal F, Stransky N, Bernard-Pierrot I, Vincent-Salomon A, de Rycke Y, Elvin P et al. (2005). Visualizing chromosomes as transcriptome correlation maps: evidence of chromosomal domains containing co-expressed genes – a study of 130 invasive ductal breast carcinomas. Cancer Res 65: 1376–1383.
Roylance R, Gorman P, Harris W, Liebmann R, Barnes D, Hanby A et al. (1999). Comparative genomic hybridization of breast tumors stratified by histological grade reveals new insights into the biological progression of breast cancer. Cancer Res 59: 1433–1436.
Smyth GK, Yang YH, Speed T . (2003). Statistical issues in cDNA microarray data analysis. Methods Mol Biol 224: 111–136.
Soares R, Balogh G, Guo S, Gartner F, Russo J, Schmitt F . (2004). Evidence for the notch signaling pathway on the role of estrogen in angiogenesis. Mol Endocrinol 18: 2333–2343.
Sorlie T, Perou CM, Tibshirani R, Aas T, Geisler S, Johnsen H et al. (2001). Gene expression patterns of breast carcinomas distinguish tumor subclasses with clinical implications. Proc Natl Acad Sci USA 98: 10869–10874.
Stange DE, Radlwimmer B, Schubert F, Traub F, Pich A, Toedt G et al. (2006). High-resolution genomic profiling reveals association of chromosomal aberrations on 1q and 16p with histologic and genetic subgroups of invasive breast cancer. Clin Cancer Res. 12: 345–352.
Storey JD, Tibshirani R . (2003). Statistical significance for genomewide studies. Proc Natl Acad Sci USA 100: 9440–9445.
Tirkkonen M, Tanner M, Karhu R, Kallioniemi A, Isola J, Kallioniemi OP . (1998). Molecular cytogenetics of primary breast cancer by CGH. Genes Chromosomes Cancer 21: 177–184.
Tsarouha H, Pandis N, Bardi G, Teixeira MR, Andersen JA, Heim S . (1999). Karyotypic evolution in breast carcinomas with i(1)(q10) and der(1;16)(q10;p10) as the primary chromosome abnormality. Cancer Genet Cytogenet 113: 156–161.
van‘t Veer LJ, Dai H, van de Vijver MJ, He YD, Hart AA, Mao M et al. (2002). Gene expression profiling predicts clinical outcome of breast cancer. Nature 415: 530–536.
Wang DY, Fulthorpe R, Liss SN, Edwards EA . (2004). Identification of estrogen-responsive genes by complementary deoxyribonucleic acid microarray and characterization of a novel early estrogen-induced gene: EEIG1. Mol Endocrinol 18: 402–411.
Wenger CR, Beardslee S, Owens MA, Pounds G, Oldaker T, Vendely P et al. (1993). DNA ploidy, S-phase, and steroid receptors in more than 127,000 breast cancer patients. Breast Cancer Res Treat 28: 9–20.
Zavadil J, Cermak L, Soto-Nieves N, Bottinger EP . (2004). Integration of TGF-beta/Smad and Jagged1/Notch signalling in epithelial-to-mesenchymal transition. EMBO J 23: 1155–1165.
Acknowledgements
We thank Vysis for providing all the reagents and Teresa Ruffalo for technical help. The work was funded by Cancer Research UK. NLB-M was supported by Fundação para a Ciência e a Tecnologia, Portugal (Fellowship SFRH/BD/2914/2000).
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on the Oncogene website (http://www.nature.com/onc).
Rights and permissions
About this article
Cite this article
Chin, SF., Wang, Y., Thorne, N. et al. Using array-comparative genomic hybridization to define molecular portraits of primary breast cancers. Oncogene 26, 1959–1970 (2007). https://doi.org/10.1038/sj.onc.1209985
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.onc.1209985
Keywords
This article is cited by
-
Leucine-rich repeat containing 4 act as an autophagy inhibitor that restores sensitivity of glioblastoma to temozolomide
Oncogene (2020)
-
Integrative analysis of copy number and gene expression in breast cancer using formalin-fixed paraffin-embedded core biopsy tissue: a feasibility study
BMC Genomics (2017)
-
Liprin-α1 is a regulator of vimentin intermediate filament network in the cancer cell adhesion machinery
Scientific Reports (2016)
-
Analysis of phosphatases in ER-negative breast cancers identifies DUSP4 as a critical regulator of growth and invasion
Breast Cancer Research and Treatment (2016)
-
Prognostic and therapeutic significance of ribonucleotide reductase small subunit M2 in estrogen-negative breast cancers
BMC Cancer (2014)