Abstract
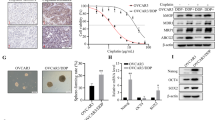
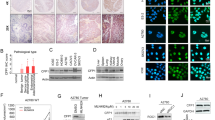
To investigate the mechanism by which HSulf-1 expression is downregulated in ovarian cancer, DNA methylation and histone acetylation of HSulf-1 was analysed in ovarian cancer cell lines and primary tumors. Treatment of OV207 and SKOV3 by 5-aza-2′-deoxycytidine resulted in increased transcription of HSulf-1. Sequence analysis of bisulfite-modified genomic DNA from ovarian cell lines and primary tumors without HSulf-1 expression revealed an increase in the frequency of methylation of 12 CpG sites in exon 1A. Chromatin immunoprecipitation assays showed an increase in histone H3 methylation in cell lines without HSulf-1 expression. To assess the significance of HSulf-1 downregulation in ovarian cancer, OV167 and OV202 cells were transfected with HSulf-1 siRNA. Downregulation of HSulf-1 expression in OV167 and OV202 cells lead to an attenuation of cisplatin-induced cytotoxicity. Moreover, patients with ovarian tumors expressing higher levels of HSulf-1 showed a 90% response rate (27/30) to chemotherapy compared to a response rate of 63% (19/30) in those with weak or moderate levels (P=0.0146, χ2 test). Collectively, these data indicate that HSulf-1 is epigenetically silenced in ovarian cancer and that epigenetic therapy targeting HSulf-1 might sensitize ovarian tumors to conventional first-line therapies.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Balch C, Huang TH, Brown R, Nephew KP . (2004). The epigenetics of ovarian cancer drug resistance and resensitization. Am J Obstet Gynecol 191: 1552–1572.
Blagosklonny MV, Robey R, Sackett DL, Du L, Traganos F, Darzynkiewicz Z et al. (2002). Histone deacetylase inhibitors all induce p21 but differentially cause tubulin acetylation, mitotic arrest, and cytotoxicity. Mol Cancer Ther 1: 937–941.
Chadee DN, Hendzel MJ, Tylipski CP, Allis CD, Bazett-Jones DP, Wright JA et al. (1999). Increased Ser-10 phosphorylation of histone H3 in mitogen-stimulated and oncogene-transformed mouse fibroblasts. J Biol Chem 274: 24914–24920.
Chien J, Aletti G, Baldi A, Catalano V, Muretto P, Keeney GL et al. (2006). Serine protease HtrA1 modulates chemotherapy-induced cytotoxicity. J Clin Invest 116: 1994–2004.
Conover CA, Hartmann LC, Bradley S, Stalboerger P, Klee GG, Kalli KR et al. (1998). Biological characterization of human epithelial ovarian carcinoma cells in primary culture: the insulin-like growth factor system. Exp Cell Res 238: 439–449.
Cross SH, Charlton JA, Nan X, Bird AP . (1994). Purification of CpG islands using a methylated DNA binding column. Nat Genet 6: 236–244.
Dai Y, Yang Y, MacLeod V, Yue X, Rapraeger AC, Shriver Z . (2005). HSulf-1 and HSulf-2 are potent inhibitors of myeloma tumor growth in vivo. J Biol Chem 280: 40066–40073.
Douglas DB, Akiyama Y, Carraway H, Belinsky SA, Esteller M, Gabrielson E et al. (2004). Hypermethylation of a small CpGuanine-rich region correlates with loss of activator protein-2alpha expression during progression of breast cancer. Cancer Res 64: 1611–1620.
Esteller M, Gaidano G, Goodman SN, Zagonel V, Capello D, Botto B, Gloghini A et al. (2002). Hypermethylation of the DNA repair gene O(6)-methylguanine DNA methyltransferase and survival of patients with diffuse large B-cell lymphoma. J Natl Cancer Inst 94: 26–32.
Fujii S, Luo RZ, Yuan J, Kadota M, Oshimura M, Dent SR et al. (2003). Reactivation of the silenced and imprinted alleles of ARHI is associated with increased histone H3 acetylation and decreased histone H3 lysine 9 methylation. Hum Mol Genet 12: 1791–1800.
Galm O, Rountree MR, Bachman KE, Jair KW, Baylin SB, Herman JG . (2002). Enzymatic regional methylation assay: a novel method to quantify regional CpG methylation density. Genome Res 12: 153–157.
Gifford G, Paul J, Vasey PA, Kaye SB, Brown R et al. (2004). The acquisition of hMLH1 methylation in plasma DNA after chemotherapy predicts poor survival for ovarian cancer patients. Clin Cancer Res 10: 4420–4426.
Gius D, Cui H, Bradbury CM, Cook J, Smart DK, Zhao SS et al. (2004). Distinct effects on gene expression of chemical and genetic manipulation of the cancer epigenome revealed by a multimodality approach. Cancer Cell 6: 361–371.
Greenlee RT, Hill-Harmon MB, Murray T, Thun M . (2001). Cancer statistics, 2001. CA Cancer J Clin 51: 15–36.
Han JW, Ahn SH, Park SH, Wang SY, Bae GU, Seo D et al. (2000). Apicidin, a histone deacetylase inhibitor, inhibits proliferation of tumor cells via induction of p21WAF1/Cip1 and gelsolin. Cancer Res 60: 6068–6074.
Herman JG, Graff JR, Myohanen S, Nelkin BD, Baylin SB . (1996). Methylation-specific PCR: a novel PCR assay for methylation status of CpG islands. Proc Natl Acad Sci USA 93: 9821–9826.
Johnstone RW . (2002). Histone-deacetylase inhibitors: novel drugs for the treatment of cancer. Nat Rev Drug Discov 1: 287–299.
Johnstone RW, Ruefli AA, Lowe SW . (2002). Apoptosis: a link between cancer genetics and chemotherapy. Cell 108: 153–164.
Jones PA, Baylin SB . (2002). The fundamental role of epigenetic events in cancer. Nat Rev Genet 3: 415–428.
Katsaros D, Cho W, Singal R, Fracchioli S, Rigault De La Longrais IA, Arisio R et al. (2004). Methylation of tumor suppressor gene p16 and prognosis of epithelial ovarian cancer. Gynecol Oncol 94: 685–692.
Kondo Y, Shen L, Yan PS, Huang TH, Issa JP . (2004). Chromatin immunoprecipitation microarrays for identification of genes silenced by histone H3 lysine 9 methylation. Proc Natl Acad Sci USA 101: 7398–7403.
Kopelovich L, Crowell JA, Fay JR . (2003). The epigenome as a target for cancer chemoprevention. J Natl Cancer Inst 95: 1747–1757.
Lai J, Chien J, Staub J, Avula R, Greene EL, Matthews TA et al. (2003). Loss of HSulf-1 up-regulates heparin-binding growth factor signaling in cancer. J Biol Chem 278: 23107–23117.
Lai JP, Chien J, Strome SE, Staub J, Montoya DP, Greene EL et al. (2004a). HSulf-1 modulates HGF-mediated tumor cell invasion and signaling in head and neck squamous carcinoma. Oncogene 23: 1439–1447.
Lai JP, Chien JR, Moser DR, Staub JK, Aderca I, Montoya DP et al. (2004b). hSulf1 sulfatase promotes apoptosis of hepatocellular cancer cells by decreasing heparin-binding growth factor signaling. Gastroenterology 126: 231–248.
Lai JP, Yu C, Moser CD, Aderca I, Han T, Garvey TD et al. (2006). SULF1 inhibits tumor growth and potentiates the effects of histone deacetylase inhibitors in hepatocellular carcinoma. Gastroenterology 130: 2130–2144.
LaVoie HA . (2005). Epigenetic control of ovarian function: the emerging role of histone modifications. Mol Cell Endocrinol 243: 12–18.
Makarla PB, Saboorian MH, Ashfaq R, Toyooka KO, Toyooka S, Minna JD et al. (2005). Promoter hypermethylation profile of ovarian epithelial neoplasms. Clin Cancer Res 11: 5365–5369.
Marks PA, Richon VM, Breslow R, Rifkind RA . (2001). Histone deacetylase inhibitors as new cancer drugs. Curr Opin Oncol 13: 477–483.
Narita K, Staub J, Chien J, Meyer K, Bauer M, Friedl A et al. (2006). HSulf-1 Inhibits Angiogenesis Tumorigenesis in vivo. Cancer Res 66: 6025–6032.
Nguyen CT, Gonzales FA, Jones PA . (2001). Altered chromatin structure associated with methylation-induced gene silencing in cancer cells: correlation of accessibility methylation MeCP2 binding acetylation. Nucleic Acids Res 29: 4598–4606.
Orlando V . (2000). Mapping chromosomal proteins in vivo by formaldehyde-crosslinked-chromatin immunoprecipitation. Trends Biochem Sci 25: 99–104.
Overbergh L, Valckx D, Waer M, Mathieu C . (1999). Quantification of murine cytokine mRNAs using real time quantitative reverse transcriptase PCR. Cytokine 11: 305–312.
Plumb JA, Strathdee G, Sludden J, Kaye SB, Brown R . (2000). Reversal of drug resistance in human tumor xenografts by 2′-deoxy-5-azacytidine-induced demethylation of the hMLH1 gene promoter. Cancer Res 60: 6039–6044.
Qian X, Guerrero RB, Plummer TB, Alves VF, Lloyd RV . (2004). Detection of hepatitis C virus RNA in formalin-fixed paraffin-embedded sections with digoxigenin-labeled cRNA probes. Diagn Mol Pathol 13: 9–14.
Seidl S, Ackermann J, Kaufmann H, Keck A, Nosslinger T, Zielinski et al. (2004). DNA-methylation analysis identifies the E-cadherin gene as a potential marker of disease progression in patients with monoclonal gammopathies. Cancer 100: 2598–2606.
Shridhar V, Bible KC, Staub J, Avula R, Lee YK, Kalli K et al. (2001). Loss of expression of a new member of the DNAJ protein family confers resistance to chemotherapeutic agents used in the treatment of ovarian cancer. Cancer Res 61: 4258–4265.
Strathdee G, Vass JK, Oien KA, Siddiqui N, Curto-Garcia J& Brown R . (2005). Demethylation of the MCJ gene in stage III/IV epithelial ovarian cancer response to chemotherapy. Gynecol Oncol 97: 898–903.
Terasawa K, Sagae S, Toyota M, Tsukada K, Ogi K, Satoh AT et al. (2004). Epigenetic inactivation of TMS1/ASC in ovarian cancer. Clin Cancer Res 10: 2000–2006.
Xing M, Cohen Y, Mambo E, Tallini G, Udelsman R, Ladenson PW et al. (2004). Early occurrence of RASSF1A hypermethylation its mutual exclusion with BRAF mutation in thyroid tumorigenesis. Cancer Res 64: 1664–1668.
Yamashita K, Upadhyay S, Osada M, Hoque MO, Xiao Y, Mori M et al. (2002). Pharmacologic unmasking of epigenetically silenced tumor suppressor genes in esophageal squamous cell carcinoma. Cancer Cell 2: 485–495.
Yang QW, Liu S, Tian Y, Salwen HR, Chlenski A, Weinstein J et al. (2003). Methylation-associated silencing of the thrombospondin-1 gene in human neuroblastoma. Cancer Res 63: 6299–6310.
Yoon JH, Dammann R, Pfeifer GP . (2001). Hypermethylation of the CpG island of the RASSF1A gene in ovarian renal cell carcinomas. Int J Cancer 94: 212–217.
Acknowledgements
This work was supported by grants from the US National Cancer Institute (CA106954-02), the John W Anderson Foundation, Minnesota Ovarian Cancer Alliance (MOCA), Edith and Bernard Waterman Foundation and the Mayo Foundation (all to VS). We would like to acknowledge Dr Lynn Hartmann and Dr Kimberly Kalli for providing us with the ovarian tissue microarray.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Staub, J., Chien, J., Pan, Y. et al. Epigenetic silencing of HSulf-1 in ovarian cancer:implications in chemoresistance. Oncogene 26, 4969–4978 (2007). https://doi.org/10.1038/sj.onc.1210300
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.onc.1210300
Keywords
This article is cited by
-
Abnormal methylation characteristics predict chemoresistance and poor prognosis in advanced high-grade serous ovarian cancer
Clinical Epigenetics (2021)
-
The Role of Epigenetic Changes in Ovarian Cancer: A Review
Indian Journal of Gynecologic Oncology (2021)
-
Methylome analysis of extreme chemoresponsive patients identifies novel markers of platinum sensitivity in high-grade serous ovarian cancer
BMC Medicine (2017)
-
Loss of HSulf-1: The Missing Link between Autophagy and Lipid Droplets in Ovarian Cancer
Scientific Reports (2017)
-
The heparan sulfate sulfotransferase 3-OST3A (HS3ST3A) is a novel tumor regulator and a prognostic marker in breast cancer
Oncogene (2016)