Abstract
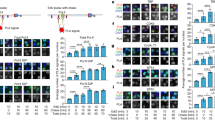
IN eukaryotic cells, active genes and their regulatory sequences are organized into open chromatin conformations in which nucleosomes can be modified, disrupted or totally absent1–3. It has been proposed that these characteristic chromatin structures and their associated factors might be directly inherited by the newly synthesized daughter strands during chromosome duplication4–6. Here we show that in the yeast Saccharomyces cerevisiae, replication machinery entering upstream of a transcriptionally active ribosomal RNA gene generates two newly replicated coding regions regularly packaged into nucleosomes, indicating that the active chromatin structure cannot be directly inherited at the replication fork. Whereas the establishment of an exposed chromatin conformation at some newly replicated rRNA gene promoters can occur shortly after the passage of the replication fork, regeneration of the active chromatin structure along the coding region is always a post-replicative process involving disruption of preformed nucleosomes.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Felsenfeld, G. Nature 355, 219–224 (1992).
Kornberg, R. D. & Lorch, Y. A. Rev. Cell Biol. 8, 563–587 (1992).
Sogo, J. M., Ness., P. J., Widmer, R. M., Parish, R. W. & Koller, T. J. molec. Biol. 178, 897–928 (1984).
Alberts, B., Worcel, A. & Weintruab, H. Proc. int. Symp. on the Eukaiyotic Genome (eds Bradbury, M. & Javaherian, K.) 165–191 (Academic, New York, 1977).
Brown, D. D. Cell 37, 359–365 (1984).
Weintraub, H. Cell 42, 705–711 (1985).
Dammann, R., Lucchini, R., Koller, T. & Sogo, J. M. Nucleic Acids Res. 10, 2331–2338 (1993).
Conconi, A., Losa, R., Koller, T. & Sogo, J. M. J. molec. Biol. 178, 920–928 (1984).
Conconi, A., Widmer, R. M., Koller, T. & Sogo, J. M. Cell 57, 753–761 (1989).
Lucchini, R. & Sogo, J. M. Molec. cell. Biol. 12, 4288–4296 (1992).
Lucchini, R. & Sogo, J. M. Molec. cell. Biol. 14, 318–326 (1994).
Sogo, J. M., Stahl, H., Koller, T. & Knippers, R. J. molec. Biol. 189, 189–204 (1986).
Brewer, B. J. & Fangman, W. L. Cell 55, 637–643 (1988).
Linskens, M. H. K. & Huberman, J. A. Molec. cell. Biol. 8, 4927–4935 (1988).
Jackson, V. Biochemistry 29, 719–731 (1990).
Svaren, J. & Chalkley, R. Trends Genet. 6, 52–56 (1990).
Wolffe, A. P. J. Cell Sci. 99, 201–206 (1991).
Kulkens, T., van Heerikhuizen, H., Klootwijk, J., Oliemans, J. & Planta, R. J. Curr. Genet. 16, 351–359 (1989).
Morrow, B. E., Johnson, S. P. & Warner, J. R. J. biol. Chem. 264, 9061–9068 (1989).
Chasman, D. I. et al. Genes Dev. 4, 403–514 (1990).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Lucchini, R., Sogo, J. Replication of transcriptionally active chromatin. Nature 374, 276–280 (1995). https://doi.org/10.1038/374276a0
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/374276a0
This article is cited by
-
Mutation of a Nopp140 gene dao-5 alters rDNA transcription and increases germ cell apoptosis in C. elegans
Cell Death & Disease (2014)
-
Ribosomen-Biogenese: Hierarchie oder koordiniertes Miteinander?
BIOspektrum (2011)
-
Transcript counting in single cells reveals dynamics of rDNA transcription
Molecular Systems Biology (2010)
-
Chromatin structure of ribosomal genes in Chironomus thummi (Diptera: Chironomidae): tissue specificity and behaviour under drug treatment
Chromosome Research (2007)
-
The chromatin remodeling complex NoRC controls replication timing of rRNA genes
The EMBO Journal (2005)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.