Key Points
-
Asymmetric cell division can be divided into four main steps: symmetry breaking, polarity establishment, determinant segregation and spindle positioning. As a result of these steps, a mother cell can give rise to two daughter cells with different fates.
-
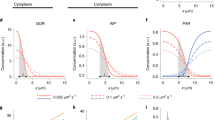
Symmetry breaking and polarity establishment are exemplified using the one-cell Caenorhabditis elegans embryo. This example illustrates the general principle whereby local modulation of the actin cytoskeleton is crucial for symmetry breaking and for polarity establishment.
-
The mechanisms by which cell polarity is translated into segregation of fate determinants are reviewed, with an emphasis on Drosophila melanogaster neuroblasts. In addition, the importance of regulated trafficking in ensuring proper cell-fate acquisition is illustrated using the example of D. melanogaster sensory organ precursors.
-
The mechanisms by which cell polarity is coupled to spindle positioning are covered, focusing again on the one-cell C. elegans embryo. Here, the available evidence indicates that spindle positioning is mediated by microtubule depolymerization and dynein function, as well as by heterotrimeric G proteins and associated components.
-
The consequences for proliferation control of defective asymmetric division of D. melanogaster neuroblasts are reviewed, and these underscore why findings in flies and worms are relevant for understanding self-renewing normal and cancer stem cells.
-
The review also includes a brief discussion of asymmetric cell division in vertebrates, and provides a taster of some promising directions for this exciting field.
Abstract
Asymmetric cell division is fundamental for generating diversity in multicellular organisms. The mechanisms that govern asymmetric cell division are increasingly well understood, owing notably to studies that were conducted in Drosophila melanogaster and Caenorhabditis elegans. Lessons learned from these two model organisms also apply to cells that divide asymmetrically in other metazoans, such as self-renewing stem cells in mammals.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Munro, E., Nance, J. & Priess, J. R. Cortical flows powered by asymmetrical contraction transport PAR proteins to establish and maintain anterior–posterior polarity in the early C. elegans embryo. Dev. Cell 7, 413–424 (2004).
Motegi, F. & Sugimoto, A. Sequential functioning of the ECT-2 RhoGEF, RHO-1 and CDC-42 establishes cell polarity in Caenorhabditis elegans embryos. Nature Cell Biol. 8, 978–985 (2006).
Schonegg, S. & Hyman, A. A. CDC-42 and RHO-1 coordinate acto-myosin contractility and PAR protein localization during polarity establishment in C. elegans embryos. Development 133, 3507–3516 (2006).
Goldstein, B. & Hird, S. N. Specification of the anteroposterior axis in Caenorhabditis elegans. Development 122, 1467–1474 (1996).
Jenkins, N., Saam, J. R. & Mango, S. E. CYK-4/GAP provides a localized cue to initiate anteroposterior polarity upon fertilization. Science 313, 1298–1301 (2006). This study reports that the GAP CYK-4 is donated to the zygote by the sperm and is required to initiate establishment of A–P polarity in C. elegans embryos, presumably through local downregulation of Rho activity.
O'Connell, K. F., Maxwell, K. N. & White, J. G. The spd-2 gene is required for polarization of the anteroposterior axis and formation of the sperm asters in the Caenorhabditis elegans zygote. Dev. Biol. 222, 55–70 (2000).
Hamill, D. R., Severson, A. F., Carter, J. C. & Bowerman, B. Centrosome maturation and mitotic spindle assembly in C. elegans require SPD-5, a protein with multiple coiled-coil domains. Dev. Cell 3, 673–684 (2002).
Cowan, C. R. & Hyman, A. A. Cyclin E–Cdk2 temporally regulates centrosome assembly and establishment of polarity in Caenorhabditis elegans embryos. Nature Cell Biol. 8, 1441–1447 (2006). In this study, a laser microbeam was used to ablate the centrosome, which demonstrated that the centrosome is normally needed to initiate A–P polarity establishment in C. elegans embryos.
Cowan, C. R. & Hyman, A. A. Centrosomes direct cell polarity independently of microtubule assembly in C. elegans embryos. Nature 431, 92–96 (2004).
Sonneville, R. & Gönczy, P. zyg-11 and cul-2 regulate progression through meiosis II and polarity establishment in C. elegans. Development 131, 3527–3543 (2004).
Tsai, M. C. & Ahringer, J. Microtubules are involved in anterior–posterior axis formation in C. elegans embryos. J. Cell Biol. 179, 397–402 (2007).
Etemad-Moghadam, B., Guo, S. & Kemphues, K. J. Asymmetrically distributed PAR-3 protein contributes to cell polarity and spindle alignement in early C. elegans embryos. Cell 83, 743–752 (1995).
Tabuse, Y. et al. Atypical protein kinase C cooperates with PAR-3 to establish embryonic polarity in Caenorhabditis elegans. Development 125, 3607–3614 (1998).
Hung, T. J. & Kemphues, K. J. PAR-6 is a conserved PDZ domain-containing protein that colocalizes with PAR-3 in Caenorhabditis elegans embryos. Development 126, 127–135 (1999).
Cuenca, A. A., Schetter, A., Aceto, D., Kemphues, K. & Seydoux, G. Polarization of the C. elegans zygote proceeds via distinct establishment and maintenance phases. Development 130, 1255–1265 (2003).
Boyd, L., Guo, S., Levitan, D., Stinchcomb, D. T. & Kemphues, K. J. PAR-2 is asymmetrically distributed and promotes association of P granules and PAR-1 with the cortex in C. elegans embryos. Development 122, 3075–3084 (1996).
Guo, S. & Kemphues, K. J. par-1, a gene required for establishing polarity in C. elegans embryos, encodes a putative Ser/Thr kinase that is asymmetrically distributed. Cell 81, 611–620 (1995).
Morton, D. G. et al. The Caenorhabditis elegans par-5 gene encodes a 14–3–3 protein required for cellular asymmetry in the early embryo. Dev. Biol. 241, 47–58 (2002).
Hao, Y., Boyd, L. & Seydoux, G. Stabilization of cell polarity by the C. elegans RING protein PAR-2. Dev. Cell 10, 199–208 (2006).
Hill, D. P. & Strome, S. An analysis of the role of microfilaments in the establishment and maintenance of asymmetry in Caenorhabditis elegans zygotes. Dev. Biol. 125, 75–84 (1988).
Guo, S. & Kemphues, K. J. A non-muscle myosin required for embryonic polarity in Caenorhabditis elegans. Nature 382, 455–458 (1996).
Shelton, C. A., Carter, J. C., Ellis, G. C. & Bowerman, B. The nonmuscle myosin regulatory light chain gene mlc-4 is required for cytokinesis, anterior–posterior polarity, and body morphology during Caenorhabditis elegans embryogenesis. J. Cell Biol. 146, 439–451 (1999).
Cheeks, R. J. et al. C. elegans PAR proteins function by mobilizing and stabilizing asymmetrically localized protein complexes. Curr. Biol. 14, 851–862 (2004).
Morton, D. G., Roos, J. M. & Kemphues, K. J. par-4, a gene required for cytoplasmic localization and determination of specific cell types in Caenorhabditis elegans embryogenesis. Genetics 130, 771–790 (1992).
Beers, M. & Kemphues, K. Depletion of the co-chaperone CDC-37 reveals two modes of PAR-6 cortical association in C. elegans embryos. Development 133, 3745–3754 (2006).
Gotta, M., Abraham, M. C. & Ahringer, J. CDC-42 controls early cell polarity and spindle orientation in C. elegans. Curr. Biol. 11, 482–488 (2001).
Aceto, D., Beers, M. & Kemphues, K. J. Interaction of PAR-6 with CDC-42 is required for maintenance but not establishment of PAR asymmetry in C. elegans. Dev. Biol. 299, 386–397 (2006).
Johnstone, O. & Lasko, P. Translational regulation and RNA localization in Drosophila oocytes and embryos. Annu. Rev. Genet. 35, 365–406 (2001).
DeRenzo, C., Reese, K. J. & Seydoux, G. Exclusion of germ plasm proteins from somatic lineages by cullin-dependent degradation. Nature 424, 685–689 (2003).
Doe, C. Q., Chu-LaGraff, Q., Wright, D. M. & Scott, M. P. The prospero gene specifies cell fates in the Drosophila central nervous system. Cell 65, 451–464 (1991).
Vaessin, H. et al. prospero is expressed in neuronal precursors and encodes a nuclear protein that is involved in the control of axonal outgrowth in Drosophila. Cell 67, 941–953 (1991).
Spana, E. P. & Doe, C. Q. The prospero transcription factor is asymmetrically localized to the cell cortex during neuroblast mitosis in Drosophila. Development 121, 3187–3195 (1995).
Hirata, J., Nakagoshi, H., Nabeshima, Y. & Matsuzaki, F. Asymmetric segregation of the homeodomain protein Prospero during Drosophila development. Nature 377, 627–630 (1995).
Knoblich, J. A., Jan, L. Y. & Jan, Y. N. Asymmetric segregation of Numb and Prospero during cell division. Nature 377, 624–627 (1995).
Schuldt, A. J. et al. Miranda mediates asymmetric protein and RNA localization in the developing nervous system. Genes Dev. 12, 1847–1857 (1998).
Broadus, J., Fuerstenberg, S. & Doe, C. Q. Staufen-dependent localization of prospero mRNA contributes to neuroblast daughter-cell fate. Nature 391, 792–795 (1998).
Bello, B., Reichert, H. & Hirth, F. The brain tumor gene negatively regulates neural progenitor cell proliferation in the larval central brain of Drosophila. Development 133, 2639–2648 (2006).
Betschinger, J., Mechtler, K. & Knoblich, J. A. Asymmetric segregation of the tumor suppressor Brat regulates self-renewal in Drosophila neural stem cells. Cell 124, 1241–1253 (2006). This article demonstrates that during asymmetric division of D. melanogaster neuroblasts, the protein Brat is inherited strictly by the GMC, where it prevents self-renewal, probably by interfering with MYC at the post-transcriptional level. See also references 37 and 39.
Lee, C. Y., Wilkinson, B. D., Siegrist, S. E., Wharton, R. P. & Doe, C. Q. Brat is a Miranda cargo protein that promotes neuronal differentiation and inhibits neuroblast self-renewal. Dev. Cell 10, 441–449 (2006).
Uemura, T., Shepherd, S., Ackerman, L., Jan, L. Y. & Jan, Y. N. numb, a gene required in determination of cell fate during sensory organ formation in Drosophila embryos. Cell 58, 349–360 (1989).
Lu, B., Rothenberg, M., Jan, L. Y. & Jan, Y. N. Partner of Numb colocalizes with Numb during mitosis and directs Numb asymmetric localization in Drosophila neural and muscle progenitors. Cell 95, 225–235 (1998).
Rhyu, M. S., Jan, L. Y. & Jan, Y. N. Asymmetric distribution of Numb protein during division of the sensory organ precursor cell confers distinct fates to daughter cells. Cell 76, 477–491 (1994).
Lee, C. Y. et al. Drosophila Aurora-A kinase inhibits neuroblast self-renewal by regulating aPKC/Numb cortical polarity and spindle orientation. Genes Dev. 20, 3464–3474 (2006).
Wang, H. et al. Aurora-A acts as a tumor suppressor and regulates self-renewal of Drosophila neuroblasts. Genes Dev. 20, 3453–3463 (2006).
Mayer, B., Emery, G., Berdnik, D., Wirtz-Peitz, F. & Knoblich, J. A. Quantitative analysis of protein dynamics during asymmetric cell division. Curr. Biol. 15, 1847–1854 (2005).
Barros, C. S., Phelps, C. B. & Brand, A. H. Drosophila nonmuscle myosin II promotes the asymmetric segregation of cell fate determinants by cortical exclusion rather than active transport. Dev. Cell 5, 829–840 (2003).
Ikeshima-Kataoka, H., Skeath, J. B., Nabeshima, Y., Doe, C. Q. & Matsuzaki, F. Miranda directs Prospero to a daughter cell during Drosophila asymmetric divisions. Nature 390, 625–629 (1997).
Shen, C. P., Jan, L. Y. & Jan, Y. N. Miranda is required for the asymmetric localization of Prospero during mitosis in Drosophila. Cell 90, 449–458 (1997).
Petritsch, C., Tavosanis, G., Turck, C. W., Jan, L. Y. & Jan, Y. N. The Drosophila myosin VI Jaguar is required for basal protein targeting and correct spindle orientation in mitotic neuroblasts. Dev. Cell 4, 273–281 (2003).
Shen, C. P. et al. Miranda as a multidomain adapter linking apically localized Inscuteable and basally localized Staufen and Prospero during asymmetric cell division in Drosophila. Genes Dev. 12, 1837–1846 (1998).
Matsuzaki, F., Ohshiro, T., Ikeshima-Kataoka, H. & Izumi, H. miranda localizes staufen and prospero asymmetrically in mitotic neuroblasts and epithelial cells in early Drosophila embryogenesis. Development 125, 4089–4098 (1998).
Wodarz, A., Ramrath, A., Kuchinke, U. & Knust, E. Bazooka provides an apical cue for Inscuteable localization in Drosophila neuroblasts. Nature 402, 544–547 (1999).
Wodarz, A., Ramrath, A., Grimm, A. & Knust, E. Drosophila atypical protein kinase C associates with Bazooka and controls polarity of epithelia and neuroblasts. J. Cell Biol. 150, 1361–1374 (2000).
Schober, M., Schaefer, M. & Knoblich, J. A. Bazooka recruits Inscuteable to orient asymmetric cell divisions in Drosophila neuroblasts. Nature 402, 548–551 (1999).
Petronczki, M. & Knoblich, J. A. DmPAR-6 directs epithelial polarity and asymmetric cell division of neuroblasts in Drosophila. Nature Cell Biol. 3, 43–49 (2001).
Parmentier, M. L. et al. Rapsynoid/Partner of Inscuteable controls asymmetric division of larval neuroblasts in Drosophila. J. Neurosci. 20, RC84 (2000).
Schaefer, M., Shevchenko, A. & Knoblich, J. A. A protein complex containing Inscuteable and the Gα -binding protein Pins orients asymmetric cell divisions in Drosophila. Curr. Biol. 10, 353–362 (2000).
Yu, F., Morin, X., Cai, Y., Yang, X. & Chia, W. Analysis of Partner of Inscuteable, a novel player of Drosophila asymmetric divisions, reveals two distinct steps in inscuteable apical localization. Cell 100, 399–409 (2000).
Schaefer, M., Petronczki, M., Dorner, D., Forte, M. & Knoblich, J. A. Heterotrimeric G proteins direct two modes of asymmetric cell division in the Drosophila nervous system. Cell 107, 183–194 (2001).
Yu, F. et al. Locomotion defects, together with Pins, regulates heterotrimeric G-protein signaling during Drosophila neuroblast asymmetric divisions. Genes Dev. 19, 1341–1353 (2005).
Siller, K. H., Cabernard, C. & Doe, C. Q. The NuMA-related Mud protein binds Pins and regulates spindle orientation in Drosophila neuroblasts. Nature Cell Biol. 8, 594–600 (2006).
Bowman, S. K., Neumuller, R. A., Novatchkova, M., Du, Q. & Knoblich, J. A. The Drosophila NuMA homolog Mud regulates spindle orientation in asymmetric cell division. Dev. Cell 10, 731–742 (2006).
Izumi, Y., Ohta, N., Hisata, K., Raabe, T. & Matsuzaki, F. Drosophila Pins-binding protein Mud regulates spindle-polarity coupling and centrosome organization. Nature Cell Biol. 8, 586–593 (2006).
Kraut, R., Chia, W., Jan, L. Y., Jan, Y. N. & Knoblich, J. A. Role of Inscuteable in orienting asymmetric cell divisions in Drosophila. Nature 383, 50–55 (1996).
Peng, C. Y., Manning, L., Albertson, R. & Doe, C. Q. The tumour-suppressor genes lgl and dlg regulate basal protein targeting in Drosophila neuroblasts. Nature 408, 596–600 (2000).
Fuse, N., Hisata, K., Katzen, A. L. & Matsuzaki, F. Heterotrimeric G proteins regulate daughter cell size asymmetry in Drosophila neuroblast divisions. Curr. Biol. 13, 947–954 (2003).
Yu, F., Cai, Y., Kaushik, R., Yang, X. & Chia, W. Distinct roles of Gαi and Gβ13F subunits of the heterotrimeric G protein complex in the mediation of Drosophila neuroblast asymmetric divisions. J. Cell Biol. 162, 623–633 (2003).
Ohshiro, T., Yagami, T., Zhang, C. & Matsuzaki, F. Role of cortical tumour-suppressor proteins in asymmetric division of Drosophila neuroblast. Nature 408, 593–596 (2000).
Betschinger, J., Eisenhaber, F. & Knoblich, J. A. Phosphorylation-induced autoinhibition regulates the cytoskeletal protein Lethal (2) giant larvae. Curr. Biol. 15, 276–282 (2005).
Betschinger, J., Mechtler, K. & Knoblich, J. A. The Par complex directs asymmetric cell division by phosphorylating the cytoskeletal protein Lgl. Nature 422, 326–330 (2003). The authors report that aPKC phosphorylates the tumour-suppressor protein Lgl, which can thus promote recruitment of Pon and Mira to the basal cell cortex and ensure their inheritance by the GMC.
Lehman, K., Rossi, G., Adamo, J. E. & Brennwald, P. Yeast homologues of tomosyn and lethal giant larvae function in exocytosis and are associated with the plasma membrane SNARE, Sec9. J. Cell Biol. 146, 125–140 (1999).
Strand, D. et al. The Drosophila lethal(2) giant larvae tumor suppressor protein forms homo-oligomers and is associated with nonmuscle myosin II heavy chain. J. Cell Biol. 127, 1361–1373 (1994).
Lu, B., Usui, T., Uemura, T., Jan, L. & Jan, Y. N. Flamingo controls the planar polarity of sensory bristles and asymmetric division of sensory organ precursors in Drosophila. Curr. Biol. 9, 1247–1250 (1999).
Roegiers, F., Younger-Shepherd, S., Jan, L. Y. & Jan, Y. N. Two types of asymmetric divisions in the Drosophila sensory organ precursor cell lineage. Nature Cell Biol. 3, 58–67 (2001).
Bellaiche, Y. et al. The Partner of Inscuteable/Discs-large complex is required to establish planar polarity during asymmetric cell division in Drosophila. Cell 106, 355–366 (2001).
Bellaiche, Y., Beaudoin-Massiani, O., Stuttem, I. & Schweisguth, F. The planar cell polarity protein Strabismus promotes Pins anterior localization during asymmetric division of sensory organ precursor cells in Drosophila. Development 131, 469–478 (2004).
Guo, M., Bier, E., Jan, L. Y. & Jan, Y. N. Tramtrack acts downstream of Numb to specify distinct daughter cell fates during asymmetric cell divisions in the Drosophila PNS. Neuron 14, 913–925 (1995).
Okabe, M., Imai, T., Kurusu, M., Hiromi, Y. & Okano, H. Translational repression determines a neuronal potential in Drosophila asymmetric cell division. Nature 411, 94–98 (2001).
Schweisguth, F. & Posakony, J. W. Antagonistic activities of Suppressor of Hairless and Hairless control alternative cell fates in the Drosophila adult epidermis. Development 120, 1433–1441 (1994).
Zeng, C., Younger-Shepherd, S., Jan, L. Y. & Jan, Y. N. Delta and Serrate are redundant Notch ligands required for asymmetric cell divisions within the Drosophila sensory organ lineage. Genes Dev. 12, 1086–1091 (1998).
Berdnik, D., Torok, T., Gonzalez-Gaitan, M. & Knoblich, J. A. The endocytic protein α-adaptin is required for numb-mediated asymmetric cell division in Drosophila. Dev. Cell 3, 221–231 (2002).
Skeath, J. B. & Doe, C. Q. Sanpodo and Notch act in opposition to Numb to distinguish sibling neuron fates in the Drosophila CNS. Development 125, 1857–1865 (1998).
Hutterer, A. & Knoblich, J. A. Numb and α-adaptin regulate Sanpodo endocytosis to specify cell fate in Drosophila external sensory organs. EMBO Rep. 6, 836–842 (2005).
Seugnet, L., Simpson, P. & Haenlin, M. Requirement for dynamin during Notch signaling in Drosophila neurogenesis. Dev. Biol. 192, 585–598 (1997).
Tang, H. et al. Numb proteins specify asymmetric cell fates via an endocytosis- and proteasome-independent pathway. Mol. Cell Biol. 25, 2899–2909 (2005).
Le Borgne, R. & Schweisguth, F. Unequal segregation of Neuralized biases Notch activation during asymmetric cell division. Dev. Cell 5, 139–148 (2003).
Lai, E. C., Deblandre, G. A., Kintner, C. & Rubin, G. M. Drosophila neuralized is a ubiquitin ligase that promotes the internalization and degradation of Delta. Dev. Cell 1, 783–794 (2001).
Pavlopoulos, E. et al. neuralized encodes a peripheral membrane protein involved in Delta signaling and endocytosis. Dev. Cell 1, 807–816 (2001).
Emery, G. et al. Asymmetric Rab 11 endosomes regulate Delta recycling and specify cell fate in the Drosophila nervous system. Cell 122, 763–773 (2005). This study revealed that the recycling endosome is clustered around the centrosome in one of the two daughters of SOP asymmetric cell division, in a Numb- and Neur-independent manner, and that this results in enhanced Delta activity.
Grill, S. W., Gönczy, P., Stelzer, E. H. & Hyman, A. A. Polarity controls forces governing asymmetric spindle positioning in the Caenorhabditis elegans embryo. Nature 409, 630–633 (2001).
Grill, S. W., Howard, J., Schaffer, E., Stelzer, E. H. & Hyman, A. A. The distribution of active force generators controls mitotic spindle position. Science 301, 518–521 (2003). The analysis of centrosome fragments induced by a laser microbeam revealed that the imbalance in pulling forces acting on the two spindle poles in C. elegans embryos results from a larger number of cortical force generators being active on the posterior side.
Coue, M., Lombillo, V. A. & McIntosh, J. R. Microtubule depolymerization promotes particle and chromosome movement in vitro. J. Cell Biol. 112, 1165–1175 (1991).
Lombillo, V. A., Stewart, R. J. & McIntosh, J. R. Minus-end-directed motion of kinesin-coated microspheres driven by microtubule depolymerization. Nature 373, 161–164 (1995).
Wang, H. W. et al. Architecture of the Dam1 kinetochore ring complex and implications for microtubule-driven assembly and force-coupling mechanisms. Nature Struct. Mol. Biol. 14, 721–726 (2007).
Labbé, J. C., Maddox, P. S., Salmon, E. D. & Goldstein, B. PAR proteins regulate microtubule dynamics at the cell cortex in C. elegans. Curr. Biol. 13, 707–714 (2003).
Kozlowski, C., Srayko, M. & Nédélec, F. Cortical microtubule contacts position the spindle in C. elegans embryos. Cell 129, 499–510 (2007).
Srayko, M., Kaya, A., Stamford, J. & Hyman, A. A. Identification and characterization of factors required for microtubule growth and nucleation in the early C. elegans embryo. Dev. Cell 9, 223–236 (2005).
Nguyen-Ngoc, T., Afshar, K. & Gönczy, P. Coupling of cortical dynein and Gα proteins mediates spindle positioning in Caenorhabditis elegans. Nature Cell Biol. 9, 1294–1302 (2007). This study shows that microtubule dynamics and dynein function are both required for the generation of pulling forces in C. elegans embryos, and that LIN-5–GPR-1/2–G α promote the presence of dynein at the cell cortex.
Couwenbergs, C. et al. Heterotrimeric G protein signaling functions with dynein to promote spindle positioning in C. elegans. J. Cell Biol. 179, 15–22 (2007).
Pecreaux, J. et al. Spindle oscillations during asymmetric cell division require a threshold number of active cortical force generators. Curr. Biol. 16, 2111–2122 (2006).
Carminati, J. L. & Stearns, T. Microtubules orient the mitotic spindle in yeast through dynein-dependent interactions with the cell cortex. J. Cell Biol. 138, 629–641 (1997).
Han, G. et al. The Aspergillus cytoplasmic dynein heavy chain and NUDF localize to microtubule ends and affect microtubule dynamics. Curr. Biol. 11, 719–724 (2001).
Gotta, M. & Ahringer, J. Distinct roles for Gα and Gβγ in regulating spindle position and orientation in Caenorhabditis elegans embryos. Nature Cell Biol. 3, 297–300 (2001).
Colombo, K. et al. Translation of polarity cues into asymmetric spindle positioning in Caenorhabditis elegans embryos. Science 300, 1957–1961 (2003).
Srinivasan, D. G., Fisk, R. M., Xu, H. & van den Heuvel, S. A complex of LIN-5 and GPR proteins regulates G protein signaling and spindle function in C. elegans. Genes Dev. 17, 1225–1239 (2003).
Gotta, M., Dong, Y., Peterson, Y. K., Lanier, S. M. & Ahringer, J. Asymmetrically distributed C. elegans homologs of AGS3/PINS control spindle position in the early embryo. Curr. Biol. 13, 1029–1037 (2003).
Lorson, M. A., Horvitz, H. R. & van den Heuvel, S. LIN-5 is a novel component of the spindle apparatus required for chromosome segregation and cleavage plane specification in Caenorhabditis elegans. J. Cell Biol. 148, 73–86 (2000).
Afshar, K., Willard, F. S., Colombo, K., Siderovski, D. P. & Gönczy, P. Cortical localization of the Gα protein GPA-16 requires RIC-8 function during C. elegans asymmetric cell division. Development 132, 4449–4459 (2005).
Du, Q. & Macara, I. G. Mammalian Pins is a conformational switch that links NuMA to heterotrimeric G proteins. Cell 119, 503–516 (2004). The authors discovered that the Pins- and GPR-1/2-related GoLoco-protein LGN acts as a conformational switch that bridges the Mud- and LIN-5-related coiled-coil protein NuMA with G α during mitosis in mammalian cells.
Basto, R. et al. Flies without centrioles. Cell 125, 1375–1386 (2006).
Afshar, K. et al. RIC-8 is required for GPR-1/2-dependent Gα function during asymmetric division of C. elegans embryos. Cell 119, 219–230 (2004).
Tsou, M. F., Hayashi, A. & Rose, L. S. LET-99 opposes Gα/GPR signaling to generate asymmetry for spindle positioning in response to PAR and MES-1/SRC-1 signaling. Development 130, 5717–5730 (2003).
Willard, F. S., Kimple, R. J. & Siderovski, D. P. Return of the GDI: the GoLoco Motif in cell division. Annu. Rev. Biochem. 73, 925–951 (2004).
Hess, H. A., Roper, J. C., Grill, S. W. & Koelle, M. R. RGS-7 completes a receptor-independent heterotrimeric G protein cycle to asymmetrically regulate mitotic spindle positioning in C. elegans. Cell 119, 209–218 (2004).
Couwembergs, C., Spilker, A. C. & Gotta, M. Control of embryonic spindle positioning and Gα activity by C. elegans RIC-8. Curr. Biol. 14, 1871–1876 (2004).
Hampoelz, B. & Knoblich, J. A. Heterotrimeric G proteins: new tricks for an old dog. Cell 119, 453–456 (2004).
Wang, H. et al. Ric-8 controls Drosophila neural progenitor asymmetric division by regulating heterotrimeric G proteins. Nature Cell Biol. 7, 1091–1098 (2005).
Gönczy, P., Pichler, S., Kirkham, M. & Hyman, A. A. Cytoplasmic dynein is required for distinct aspects of MTOC positioning, including centrosome separation, in the one cell stage Caenorhabditis elegans embryo. J. Cell Biol. 147, 135–150 (1999).
Lee, C. Y., Robinson, K. J. & Doe, C. Q. Lgl, Pins and aPKC regulate neuroblast self-renewal versus differentiation. Nature 439, 594–598 (2006).
Wang, H., Ouyang, Y., Somers, W. G., Chia, W. & Lu, B. Polo inhibits progenitor self-renewal and regulates Numb asymmetry by phosphorylating Pon. Nature 449, 96–100 (2007).
Berdnik, D. & Knoblich, J. A. Drosophila Aurora-A is required for centrosome maturation and actin-dependent asymmetric protein localization during mitosis. Curr. Biol. 12, 640–647 (2002).
Caussinus, E. & Gonzalez, C. Induction of tumor growth by altered stem-cell asymmetric division in Drosophila melanogaster. Nature Genet. 37, 1125–1129 (2005).
Gonzalez, C. Spindle orientation, asymmetric division and tumour suppression in Drosophila stem cells. Nature Rev. Genet. 8, 462–472 (2007).
Chenn, A. & McConnell, S. K. Cleavage orientation and the asymmetric inheritance of Notch1 immunoreactivity in mammalian neurogenesis. Cell 82, 631–641 (1995).
Cayouette, M. & Raff, M. The orientation of cell division influences cell-fate choice in the developing mammalian retina. Development 130, 2329–2339 (2003).
Haydar, T. F., Ang, E. Jr, & Rakic, P. Mitotic spindle rotation and mode of cell division in the developing telencephalon. Proc. Natl Acad. Sci. USA 100, 2890–2895 (2003).
Noctor, S. C., Martinez-Cerdeno, V., Ivic, L. & Kriegstein, A. R. Cortical neurons arise in symmetric and asymmetric division zones and migrate through specific phases. Nature Neurosci. 7, 136–144 (2004).
Konno, D. et al. Neuroepithelial progenitors undergo LGN-dependent planar divisions to maintain self-renewability during mammalian neurogenesis. Nature Cell Biol. 10, 93–101 (2008).
Zhong, W., Feder, J. N., Jiang, M. M., Jan, L. Y. & Jan, Y. N. Asymmetric localization of a mammalian Numb homolog during mouse cortical neurogenesis. Neuron 17, 43–53 (1996).
Rasin, M. R. et al. Numb and Numbl are required for maintenance of cadherin-based adhesion and polarity of neural progenitors. Nature Neurosci. 10, 819–827 (2007). The authors clarify the role of mouse Numb and Numblike by showing that the two proteins together are important for morphogenesis of the vertebrate cerebral cortex because they maintain adherens junctions of radial glial cells.
Zhong, W., Jiang, M. M., Weinmaster, G., Jan, L. Y. & Jan, Y. N. Differential expression of mammalian Numb, Numblike and Notch1 suggests distinct roles during mouse cortical neurogenesis. Development 124, 1887–1897 (1997).
Petersen, P. H., Zou, K., Krauss, S. & Zhong, W. Continuing role for mouse Numb and Numbl in maintaining progenitor cells during cortical neurogenesis. Nature Neurosci. 7, 803–811 (2004).
Zigman, M. et al. Mammalian inscuteable regulates spindle orientation and cell fate in the developing retina. Neuron 48, 539–545 (2005).
Sanada, K. & Tsai, L. H. G protein βγ subunits and AGS3 control spindle orientation and asymmetric cell fate of cerebral cortical progenitors. Cell 122, 119–131 (2005).
Morin, X., Jaouen, F. & Durbec, P. Control of planar divisions by the G-protein regulator LGN maintains progenitors in the chick neuroepithelium. Nature Neurosci. 10, 1440–1448 (2007).
Fischer, E. et al. Defective planar cell polarity in polycystic kidney disease. Nature Genet. 38, 21–23 (2006).
Lechler, T. & Fuchs, E. Asymmetric cell divisions promote stratification and differentiation of mammalian skin. Nature 437, 275–280 (2005).
Théry, M. et al. The extracellular matrix guides the orientation of the cell division axis. Nature Cell Biol. 7, 947–953 (2005).
Dalgaard, J. Z. & Klar, A. J. Does S. pombe exploit the intrinsic asymmetry of DNA synthesis to imprint daughter cells for mating-type switching? Trends Genet. 17, 153–157 (2001).
Yamada-Inagawa, T., Klar, A. J. & Dalgaard, J. Z. Schizosaccharomyces pombe switches mating type by the synthesis-dependent strand-annealing mechanism. Genetics 177, 255–265 (2007).
Shinin, V., Gayraud-Morel, B., Gomes, D. & Tajbakhsh, S. Asymmetric division and cosegregation of template DNA strands in adult muscle satellite cells. Nature Cell Biol. 8, 677–687 (2006).
Potten, C. S., Owen, G. & Booth, D. Intestinal stem cells protect their genome by selective segregation of template DNA strands. J. Cell Sci. 115, 2381–2388 (2002).
Smith, G. H. Label-retaining epithelial cells in mouse mammary gland divide asymmetrically and retain their template DNA strands. Development 132, 681–687 (2005).
Karpowicz, P. et al. Support for the immortal strand hypothesis: neural stem cells partition DNA asymmetrically in vitro. J. Cell Biol. 170, 721–732 (2005).
Rujano, M. A. et al. Polarised asymmetric inheritance of accumulated protein damage in higher eukaryotes. PLoS Biol. 4, e417 (2006).
Yamashita, Y. M., Mahowald, A. P., Perlin, J. R. & Fuller, M. T. Asymmetric inheritance of mother versus daughter centrosome in stem cell division. Science 315, 518–521 (2007).
Yamashita, Y. M., Jones, D. L. & Fuller, M. T. Orientation of asymmetric stem cell division by the APC tumor suppressor and centrosome. Science 301, 1547–1550 (2003).
Rebollo, E. et al. Functionally unequal centrosomes drive spindle orientation in asymmetrically dividing Drosophila neural stem cells. Dev. Cell 12, 467–474 (2007).
Rusan, N. M. & Peifer, M. A role for a novel centrosome cycle in asymmetric cell division. J. Cell Biol. 177, 13–20 (2007).
Park, D. H. & Rose, L. S. Dynamic localization of LIN-5 and GPR-1/2 to cortical force generation domains during spindle positioning. Dev. Biol. 315, 42–54 (2008).
Acknowledgements
I am grateful to K. Afshar, D. Constam, V. Hachet and J. Knoblich for critical reading of the manuscript, and to J. Knoblich also for providing the movie for Supplementary information S1. Apologies go to those authors whose work could not be covered owing to space constraints. My laboratory is supported by grants from the Swiss National Science Foundation (3100A0-102087) and Oncosuisse (OCS-01676-02-2005, OCS KLS 02024-02-2007).
Author information
Authors and Affiliations
Supplementary information
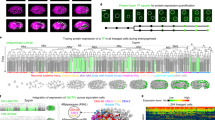
Supplementary information S1 (movie) | Asymmetric division of a Drosophila SOP
D. melanogaster sensory organ precursor (SOP) cell expressing GFP-Pon (green) and Histone2B-RFP (red). Anterior is to the left. The SOP is ∼ 10 µm in diameter. Note that GFPPon is segregated to the anterior of the SOP, and is thus inherited by pIIb. Modified with permission from ref. 1. (MOV 3318 kb)
Supplementary information S2 (movie) | Asymmetric division of a one-cell stage C. elegans embryo
C. elegans embryo expressing GFP-PIE-1 (green) and imaged using dual time-lapse DIC and fluorescence microscopy; the GFP and DIC images have been overlaid. Anterior is to the left. The embryo is ∼50 µm-long. Note that the bulk of GFP-PIE-1 is segregated to the posterior of the one-cell embryo. (MOV 8310 kb)
Related links
Glossary
- Mitotic spindle
-
A highly dynamic array of microtubules that forms during mitosis and serves to move the sister chromatids apart.
- Cell cortex
-
The region underlying the plasma membrane that is rich in actin filaments and associated proteins.
- Pronucleus
-
The haploid nucleus that is derived from either sperm or oocyte and that is present in the newly fertilized zygote.
- Centrosome
-
The principal microtubule-organizing centre of animal cells, an organelle that contains the centrioles and that anchors the minus end of microtubules.
- PAR proteins
-
A set of six proteins that were initially identified in C. elegans and whose inactivation results in a partitioning-defective phenotype in early embryos.
- PDZ domain
-
A protein-interaction domain that is often found in scaffolding proteins and that is named after the founding members of this protein family (PSD-95, discs-large and ZO-1).
- Ring finger
-
A protein domain that consists of two loops that are held together at their base by Cys and His residues that form a complex with two zinc ions. Many ring fingers function in protein degradation by facilitating protein ubiquitylation.
- 14–3–3 proteins
-
Adaptor/scaffold proteins that form homo- and heterodimers and bind, through specialized phosphorylated peptide motifs, to various proteins that are involved notably in signal transduction and in cell-cycle control.
- Neurectoderm
-
A portion of the ectoderm that is destined to become neural tissue.
- Coiled-coil domain
-
A structural domain that can mediate oligomerization. Coiled-coils contain two to five helices that twist around each other to form a supercoil.
- t-SNARE
-
An integral membrane protein that is present on a target membrane or organelle and that mediates fusion through interaction with a v-SNARE partner protein that is located on the fusion partner.
- Sheath cell
-
One of the four cells that comprises the bristle sensory organ that derives from the SOP in the peripheral nervous system of D. melanogaster.
- Socket cell
-
One of the four cells that comprises the bristle sensory organ that derives from the sensory organ precursors in the peripheral nervous system of D. melanogaster.
- Aggresome
-
Cytoplasmic inclusions bodies that contain misfolded proteins and that are located near centrosomes.
- Proteasome
-
A large protein complex that is responsible for degrading intracellular proteins that have been targeted for destruction, usually by the addition of ubiquitin polymers.
- Niche cell
-
A cell that is located in the vicinity of stem cells and that provides a suitable environment for maintaining the self-renewing capacity of stem cells.
- Gonial cell
-
A germ cell before it enters the meiotic cell cycle.
- Recycling endosome
-
An intracellular membrane compartment that mediates the recycling of endocytosed material back to the plasma membrane.
- Blastomere
-
A cell that is generated during embryonic cleavage divisions.
- Astral microtubules
-
Microtubules that radiate from the mitotic spindle poles to the cell cortex. They are involved in positioning the spindle poles during cell division.
- Kinetochore
-
A multiprotein complex that assembles on centromeric DNA and that mediates the attachment and movement of chromosomes along the microtubules of the mitotic spindle.
- Ventricular zone
-
A layer of cells that is located immediately adjacent to the cerebral ventricles in the vertebrate brain.
- Adherens junction
-
A cell–cell adhesion complex that contains cadherins and catenins that are attached to cytoplasmic actin filaments.
- Basement membrane
-
A thin layer of connective tissue that underlies the epithelium of many organs.
Rights and permissions
About this article
Cite this article
Gönczy, P. Mechanisms of asymmetric cell division: flies and worms pave the way. Nat Rev Mol Cell Biol 9, 355–366 (2008). https://doi.org/10.1038/nrm2388
Issue Date:
DOI: https://doi.org/10.1038/nrm2388
This article is cited by
-
Host-dependent impairment of parasite development and reproduction in the acanthocephalan model
Cell & Bioscience (2022)
-
Wnt signaling polarizes cortical actin polymerization to increase daughter cell asymmetry
Cell Discovery (2022)
-
Farnesoid X receptor via Notch1 directs asymmetric cell division of Sox9+ cells to prevent the development of liver cancer in a mouse model
Stem Cell Research & Therapy (2021)
-
Spindle positioning and its impact on vertebrate tissue architecture and cell fate
Nature Reviews Molecular Cell Biology (2021)
-
Force microscopy of the Caenorhabditis elegans embryonic eggshell
Microsystems & Nanoengineering (2020)