-
PDF
- Split View
-
Views
-
Cite
Cite
Armelle Logié, Claire Dunois-Lardé, Christophe Rosty, Olivier Levrel, Martine Blanche, Agnès Ribeiro, Jean-Marie Gasc, Jose Jorcano, Sabine Werner, Xavier Sastre-Garau, Jean Paul Thiery, François Radvanyi, Activating mutations of the tyrosine kinase receptor FGFR3 are associated with benign skin tumors in mice and humans, Human Molecular Genetics, Volume 14, Issue 9, 1 May 2005, Pages 1153–1160, https://doi.org/10.1093/hmg/ddi127
Close - Share Icon Share
Abstract
Specific germline activating point mutations in the gene encoding the tyrosine kinase receptor FGFR3 (fibroblast growth factor receptor 3) result in autosomal dominant human skeletal dysplasias. The identification in multiple myeloma and in two epithelial cancers—bladder and cervical carcinomas—of somatic FGFR3 mutations identical to the germinal activating mutations found in skeletal dysplasias, together with functional studies, have suggested an oncogenic role for this receptor. Although acanthosis nigricans, a benign skin tumor, has been found in some syndromes associated with germinal activating mutations of FGFR3, the role of activated FGFR3 in the epidermis has never been investigated. Here, we targeted an activated receptor mutant (S249C FGFR3) to the basal cells of the epidermis of transgenic mice. Mice expressing the transgene developed benign epidermal tumors with no sign of malignancy. These skin lesions had features in common with acanthosis nigricans and other benign human skin tumors, including seborrheic keratosis, one of the most common benign epidermal tumors in humans. We therefore screened a series of 62 cases of seborrheic keratosis for FGFR3 mutations. A large proportion of these tumors (39%) harbored somatic activating FGFR3 mutations, identical to those associated with skeletal dysplasia syndromes and bladder and cervical neoplasms. Our findings directly implicate FGFR3 activation as a major cause of benign epidermal tumors in humans.
INTRODUCTION
Fibroblast growth factor receptor 3 (FGFR3) belongs to a class of transmembrane tyrosine kinase receptors (FGFR1–4) involved in signal transduction regulating cell growth, differentiation, migration, wound healing and angiogenesis, depending on target cell type and developmental stage. These receptors are glycoproteins composed of two or three extracellular immunoglobulin (Ig)-like domains, a transmembrane domain and a split tyrosine kinase domain. Ligand binding induces receptor dimerization, leading to phosphorylation of the kinase domain (1,2). Specific activating point mutations affecting different domains of FGFR3 result in autosomal dominant human skeletal dysplasias: craniosynostoses (Crouzonodermoskeletal syndrome and Muenke craniosynostosis) and dwarfing chondrodysplasias of various degrees of severity (hypochondroplasia, achondroplasia, severe achondroplasia with developmental delay and acanthosis nigricans—also known as ‘SADDAN’—and thanatophoric dysplasia) (3,4). The degree of activation of FGFR3 is correlated with the severity of dwarfing chondrodysplasias (5). Functional studies have shown that some of these mutations activate the receptor either by generating a cysteine residue that can form an intermolecular disulfide bond leading to ligand-independent dimerization of the receptor (R248C) or by changing the conformation of the activation loop in the tyrosine kinase of the receptor (K650/652E) (3,4). Additional evidence consistent with FGFR3 playing a negative role in bone growth has been obtained in several mouse models including Fgfr3-deficient mice and mice harboring activating mutations in FGFR3 (4). In contrast to this inhibitory role, an oncogenic role has been proposed for FGFR3 in cancer. Somatic activating FGFR3 mutations identical to those found in skeletal dysplasias have been shown to be associated with multiple myeloma and more recently with epithelial malignancies (bladder and cervix carcinomas) (6–8). Functional studies have provided several lines of evidence supporting an oncogenic role for FGFR3: activated FGFR3 induces the malignant transformation of NIH3T3 fibroblasts (9,10) and generates lymphoid malignancies in mice (11). Although acanthosis nigricans, a benign skin epidermal tumor, has been found in some syndromes associated with germinal activating mutations of FGFR3 (Crouzonodermoskeletal syndrome, SADDAN syndrome and rare long-surviving thanatophoric dysplasia patients) (12–15), the role of activated FGFR3 in the epidermis has never been investigated.
We investigated the role of activated FGFR3 in the epidermis by targeting the expression of an activated receptor (S249C FGFR3) to the basal layer of the epidermis, using the bovine keratin 5 (K5) promoter (16,17). The S249C mutation, which causes ligand-independent activation of the receptor, is the most common FGFR3 mutation in carcinomas (8,18,19). This mutation also causes thanaphoric dysplasia, a syndrome associated with acanthosis nigricans in the rare long-term survivors (14,15).
RESULTS
Transgenic mice expressing mutated FGFR3 (FGFR3 S249C) in the epidermis develop benign skin tumors
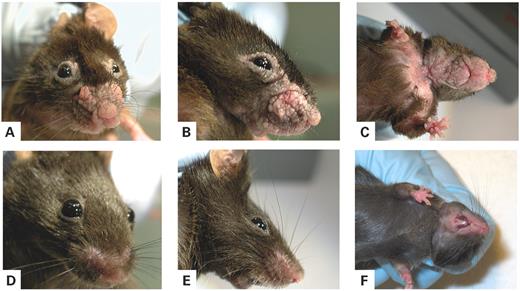
We targeted the expression of a mutated form of FGFR3 (FGFR3 S249C) to the epidermis using the keratin 5 promoter. The K5-(S249C)FGFR3 construct was made by subcloning the FGFR3 S249C cDNA (see Materials and Methods) into a vector containing a 5.2 kb fragment of the bovine keratin 5 promoter, the rabbit β-globin intron 2 and the SV40 polyadenylation signal. This construct was then microinjected into fertilized B6D2F1 mouse oocytes. Among the seven founders obtained, four expressed the K5-(S249C)FGFR3 transgene in the skin (data not shown). Three of these four founders developed a skin phenotype. These three founders were viable and fertile and transmitted the transgene to their offspring in a Mendelian fashion. The offspring developed a phenotype indistinguishable from that of the founders. At the age of 3–4 months, the transgenic mice presented skin verrucous tumors on the eyelids and the snout (Fig. 1A and B), which then gradually extended from the snout to the throat. In animals aged >5 months, another type of lesion appeared on the upper trunk, characterized by a marked thickening of the skin and hair loss (Fig. 1C). None of the skin lesions arising in the various transgenic lines showed any signs of regression. In some old mice, a thickening of the tongue and palate was observed. No lesions were observed in other tissues, although some expressed the transgene including the forestomach and the trachea (Supplementary Material Fig. S1).
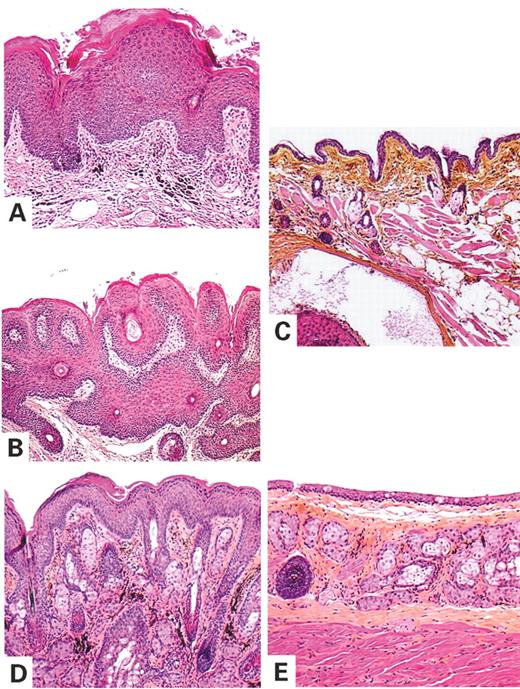
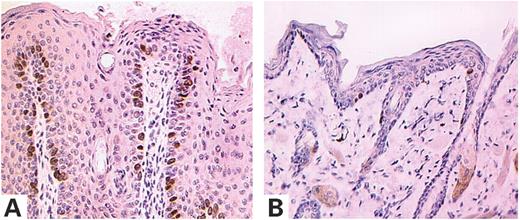
Histological examination of the skin lesions revealed common features, irrespective of body site and gross appearance: (i) pronounced acanthosis (thickening of the nucleated layers of the epithelium), (ii) hyperkeratosis (thickening of the stratum corneum) with occasional parakeratosis (hyperkeratosis, with retention of nuclei) and (iii) papillomatosis (numerous projections of the dermal papillae) (Fig. 2). The most exophytic lesions of the snout displayed a trabeculated pattern, with rare keratin-filled invaginations and small keratin cysts, which generally surrounded a hair shaft (Fig. 2B). No abnormalities of the hair follicles or sebaceous glands were observed. No histological features of malignancy were detected: mitotic figures were rare and always localized in the basal layer of the epidermis and cytological atypia was not observed. The absence of malignancy was evident, even in the lesions of transgenic mice >18 months old. Cell proliferation was studied by incorporation of the thymidine analog 5-bromo-2-deoxyuridine (BrdU). Consistent with the benign nature of the lesions, proliferating cells were restricted to the basal cell layer of the epidermis and to hair follicles in both transgenic and control mice (Fig. 3). However, the proportion of basal cells undergoing proliferation was significantly higher in transgenic mouse epidermis than in control skin (27 versus 1.8%, P<0.0001).
Frequent FGFR3 mutations are found in a common benign skin tumor in humans
The skin lesions observed in transgenic mice expressing mutated FGFR3 displayed several histological features (acanthosis, hyperkeratosis and papillomatosis) common to benign human epidermal tumors including seborrheic keratosis, one of the most common benign epidermal tumors in humans. This lesion, which has no malignant potential, is mainly located on the trunk and face and presents as two histological types—the hyperkeratotic and the acanthotic types. Seborrheic keratoses tend to appear in middle age and become more common and numerous with advancing age.
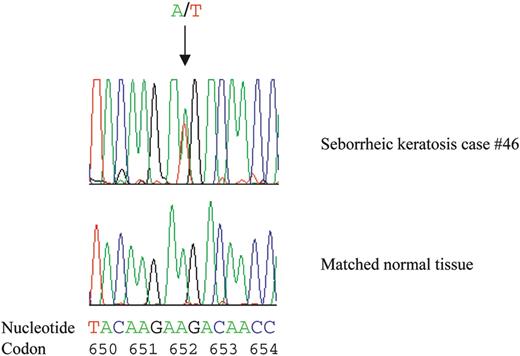
We screened a series of 62 seborrheic keratoses for FGFR3 mutations by using single strand conformation polymorphism (SSCP) analysis followed by DNA sequencing. The three regions of FGFR3 (located in exon 7, 10, 15) known to harbor most of the point mutations previously described in multiple myeloma, bladder and cervical carcinomas and skeletal dysplasias were analyzed. We identified point mutations in 24 samples (39%) (Fig. 4 and Table 1). Matched constitutional DNA, available in 10 skin tumor cases with FGFR3 mutations, contained wild-type sequences demonstrating the somatic nature of these mutations (Fig. 4) (data not shown).
We found no correlation among the presence of FGFR3 mutations, the sex or age of the patients, the location of the tumors (P=0.21) or the histological type (acanthotic or hyperkeratotic) (P=0.29). Among the four patients for whom two seborrheic keratoses were analyzed, we found a FGFR3 mutation in one of the two lesions for two patients (cases 47a and b and 53a and b), no FGFR3 mutation in either lesion for one patient (case 55a and b) and a different FGFR3 mutation in each seborrheic keratosis for one patient (case 11a and b) (Table 1).
The 24 mutations identified affected seven different codons: R248C, S249C, G372C, S373C, Y375C, K652E and K652M (Table 2). All these mutations were identical to the constitutional mutations present in patients with thanatophoric dysplasia and SADDAN (3,20,21). With the exception of the S373C mutation, all the mutations have already been described as somatic mutations in multiple myeloma and in bladder and cervical carcinomas (6–8,18,19). The mutations in exon 7 (R248C and S249C) were located at the junction between extracellular Ig-like domains II and III of FGFR3, the mutations in exon 10 (G372C, S373C and Y375C) were located in the transmembrane domain and the mutations in exon 15 (K652E and K652M) were located in the second part of the intracellular tyrosine kinase domain of FGFR3.
DISCUSSION
Since the identification of mutations of FGFR3 in human skeletal dysplasias, the role of FGFR3 in skeletal development has attracted considerable attention (4). The recent identification of somatic activating FGFR3 mutations in multiple myeloma and in two epithelial cancers—bladder and cervix carcinomas—has raised the question of an oncogenic role of mutated FGFR3 in tumorigenesis (6–8). In hematopoietic cells, functional studies have shown that mutated FGFR3 may contribute to the malignant phenotype (9–11). The oncogenic potential of mutated FGFR3 has yet to be investigated in bladder and cervix epithelia.
We show here that mutated FGFR3 can induce benign tumors with no malignant potential in mouse epidermis. This led us to identify frequent FGFR3 mutations in seborrheic keratosis, one of the most frequent benign epidermal tumors in older individuals. Here, we screened part of the coding sequence of FGFR3 (exons 7, 10 and 15) containing most of the mutations identified in severe forms of dwarfism and all the mutations already identified in bladder and cervix carcinomas. However, we cannot totally exclude the presence in seborrheic keratoses of FGFR3 mutations outside these exons.
Although the clonal nature of seborrheic keratoses has been suggested (22), no molecular alterations had been reported in these lesions. We report for the first time a molecular alteration associated with this benign skin tumor. The FGFR3 mutations that we found in seborrheic keratosis are the same as those found in thanatophoric dysplasia and SADDAN syndrome. Interestingly, the rare long-surviving thanatophoric dysplasia patients and all the patients with SADDAN syndrome develop acanthosis nigricans, a benign skin tumor sharing histological similarities with seborrheic keratosis and with the mouse skin lesions observed in our transgenic mice. Our results in mice strongly suggest that the occurrence of acanthosis nigricans observed in these human skeletal disorders results directly from the expression of mutated FGFR3 in the epidermis.
We expressed an activated form of FGFR3 in the mouse epidermis by targeting the FGFR3 S249C transgene to the epidermis with the keratin 5 promoter. A different approach—targeted homologous recombination—has been used in previous studies to generate several mouse models harboring a mutated endogenous FGFR3 gene (23–26). The various mutations inserted corresponded to mutations found in skeletal dysplasias, carcinomas and, as shown here, seborrheic keratoses. Surprisingly, although the endogenous mouse gene is expressed in the skin, no skin lesions have been reported in these models. The absence of a skin phenotype may be due to the genetic background of the mice used in these models, the level of expression of the mouse FGFR3 gene, differences in the regulation of the mouse and human promoters or slight differences between the mouse and the human FGFR3.
The skin lesions observed in our transgenic mouse model have several histological features in common with seborrheic keratoses (acanthosis, hyperkeratosis and papillomatosis). Human seborrheic keratoses characteristically display keratin-filled structures (horn cysts and pseudo-horn cysts). In the mouse model, horn cysts were present in some thick lesions, but were less numerous and smaller than those in human seborrheic keratoses. Furthermore, in mice, these cysts always surrounded a hair. The differences between human and mouse lesions may simply correspond to a difference between species. They may also be due to mutated FGFR3 being under the control of the keratin 5 promoter in the mouse model, whereas mutated FGFR3 is under the control of its own promoter in human seborrheic keratoses. The FGFR3 promoter gives suprabasal expression in resting human skin and basal expression after wounding (27). In human seborrheic keratosis, FGFR3 expression was observed in both the basal and the suprabasal layers. In transgenic mouse lesions, the mutated FGFR3 was observed in the basal layer and in the first suprabasal layers (Claire Dunois-Lardé and Jean-Marie Gasc, unpublished results).
The induction of benign skin tumors by FGFR3 mutations in mice, the occurrence of acanthosis nigricans in skeletal dysplasia syndromes caused by germinal FGFR3 mutations and the identification of somatic FGFR3 mutations in seborrheic keratoses support a role for mutated FGFR3 in benign epidermal tumorigenesis. The fact that we did not identify FGFR3 mutations in a series of epidermal carcinomas (23 squamous cell carcinomas and 22 basal cell carcinomas) (28) (data not shown) suggests that the role of activated FGFR3 in the epidermis is restricted to benign tumorigenesis. In the bladder, although FGFR3 mutations are found in a significant proportion of muscle-invasive carcinomas (15% of cases), it should be noted that such mutations occur at high frequency (>70% of cases) in low stage, low grade bladder carcinomas (TaG1–TaG2) (18,29). In addition, a high frequency of FGFR3 mutations was recently detected in urothelial papilloma, a rare entity with little or no risk of developing urothelial carcinomas (30,31).
The transgenic mouse model we have set up should be useful for studying the mechanisms of benign tumorigenesis and for developing new therapies targeting mutated FGFR3. Although not life threatening, seborrheic keratoses can be troublesome and annoying to the patient. They may become irritated and itch. Often unattractive, or even disfiguring, these lesions have a negative psychological impact—daily reminders of aging. Anti-tyrosine kinase drugs have recently been used successfully to treat several malignancies (32). The identification of activating mutations of a tyrosine kinase receptor in a significant proportion of seborrheic keratoses may provide new opportunities for the treatment of this condition.
MATERIALS AND METHODS
Generation of K5-(S249C)FGFR3 transgenic mice
We targeted the expression of an activated FGFR3 carrying the S249C mutation to the epidermis in mice, using the 5′-regulatory region of the bovine keratin 5 gene (16). The FGFR3b splice variant was used, as this is the main form of FGFR3 in the murine epidermis and human epithelia (33) (unpublished data). The S249C mutation, which causes ligand-independent activation of the receptor, is the most common mutation in carcinomas (8). The coding sequence of the human FGFR3b cDNA was amplified by RT–PCR from RNA of a normal urothelium, using the primers 5′-CGGGGCTGCCTGAGGAC-3′ and 5′-TGCTAGGGACCCCTCACATT-3′. The PCR product was inserted into a cytomegalovirus promoter-driven pcDNAI/Neo expression vector (Invitrogen, San Diego, CA, USA). The S249C mutation was then introduced into this cloned FGFR3b cDNA as follows: a 729 bp region of the FGFR3b cDNA carrying the S249C mutation was amplified by RT–PCR from RNA of a bladder carcinoma, using the primers 5′-CGTCGTGGAGAACAAGTTTGGCAG-3′ and 5′-CCGAGACAGCTCCCATTTG-3′. The Xho1–Aor51H1 fragment containing the mutation was excised from the PCR product and inserted in place of the corresponding sequence in the non-mutated FGFR3b construct. The pK5-(S249C)FGFR3b construct was then obtained by inserting the mutated FGFR3b cDNA excised with XbaI and HindIII into the SnaBI site of the pBK5 expression vector containing the 5.2 kb bovine keratin 5 (K5) regulatory sequences, β-globin intron 2 and the 3′ polyadenylation sequences of SV40. All PCR-generated segments were verified by sequencing both strands. The K5-(S249C)FGFR3 construct excised with SalI and NotI was purified and microinjected into fertilized B6D2F1 oocytes. Genomic DNA was extracted from mouse tails and screened by PCR for integration of the transgene. The care, housing and handling of the mice conformed to the recommendations of the French Ethics Committee and were overseen by authorized investigators.
Histology of mouse tissues
Tissue samples from transgenic mice and wild-type littermates matched with respect to sex and body site were collected and fixed by overnight incubation at room temperature in freshly prepared Methacarn (6 vol absolute methanol/3 vol chloroform/1 vol acetic acid). The samples were embedded in paraffin, and sections (5 µm) were cut and stained with hematoxylin–eosin–saffron for histological analysis.
BrdU labeling
Transgenic mice (n=5) from line 79 and control littermates (n=5) were injected (i.p.), with 0.25 mg g−1 body weight BrdU (Sigma). They were sacrificed 2 h later. BrdU immunohistochemistry was carried out with the BrdU in situ detection kit (BD PharMingen, San Diego, CA, USA), using the reagents and conditions suggested by the manufacturer. The number of BrdU-labeled cells among 300 cells from the basal epidermis layer was determined for each mouse.
Human skin samples
Alcohol/formalin/acetic acid-fixed paraffin-embedded specimens of seborrheic keratoses were obtained from the Pathology Department of the Institut Curie and cut into 8 µm sections. One section was used for hematoxylin–eosin–saffron staining, for distinguishing the tumor cell areas and their subsequent microdissection from the remaining unstained sections. Genomic DNA was isolated using the QIAmp tissue kit (Qiagen), according to the manufacturer's instructions.
Mutation analyses
We used SSCP and sequencing to analyze the three regions of FGFR3 in exons 7, 10 and 15 known to harbor most of the point mutations previously described in multiple myeloma, bladder and cervix carcinomas and skeletal dysplasias. SSCP and sequence analysis were performed as previously described (18), except that the following primer sequences were used: 5′-AGTGGCGGTGGTGGTGAGGGAG-3′ and 5′-GACGTTCCACATGTCACTGCGTGT-3′ for exon 7; 5′-CCTCAACGCCCATGTCTTTGCAGC-3′ and 5′-GTTCTAGAGGGCGAAGGGCGAGTTC-3′ for exon 10 and 5′-TGGTGACCGAGGACAACGTGATG-3′ and 5′-GTTGTGGCGGAAGGGTGTGGGA-3′ for exon 15.
SUPPLEMENTARY MATERIAL
Supplementary Material is available at HMG Online.
ACKNOWLEDGEMENTS
We thank Richard Ekué, Sabrina Etheve, David Cappellen, Marie-Thérèse Morin, David Ricol, Catherine de Oliveira and Monique Denoyelle for their participation in various aspects of the work. We also thank Dr Thierry Clerici, Dr Nathalie Franck, Dr Marie-Jeanne Himbert-Guillot and Dr Françoise Raynaud for fruitful discussions. This work was supported by the CNRS, the Institut Curie and Comité de Paris Ligue Nationale Contre le Cancer (Laboratoire associé).
The authors wish it to be known that, in their opinion, the first three authors should be regarded as joint First Authors.
Figure 1. Phenotype of K5-(S249C)FGFR3 transgenic mice. (A and B) Two K5-(S249C)FGFR3 transgenic mice (line 79) at 6 months of age. These mice exhibit verrucous cauliflower-like lesions on the snout and the eyelids. These lesions are well demarcated, of symmetrical localization and occasionally display partial pigmentation, as shown in (B). (C) Throat and upper chest skin lesions on a 10-month-old K5-(S249C)FGFR3 transgenic mouse (line 79) are shown. The throat lesions resemble those on the snout in (A and B), areas of flat skin thickening with hair loss are clearly apparent on the upper chest. (D–F) Control littermates are shown below each transgenic mouse.
Figure 2. Tumor histology of K5-(S249C)FGFR3 transgenic mouse skin lesions. (A, B and D) Hematoxylin, eosin and saffron sections of skin lesions from the snout (A and B) and the trunk (D) of a transgenic K5-(S249C)FGFR3 mouse. (C and E) Sections of normal snout (C) and trunk (E) from a control littermate are shown for comparison. These skin lesions showed various degrees of acanthosis, papillomatosis and hyperkeratosis. Lesion in (B) is thicker than in (A) and shows interlacing strands (trabeculated aspect) of basaloid and squamous cells enclosing hair follicles. The parakeratosis and spongiosis observed in (A) were probably caused by itching and repetitive scratching of the lesion. Magnification 100×.
Figure 3. K5-(S249C)FGFR3 transgenic mice, tumor cell proliferation, as assessed by BrdU labeling. Mice were injected with 5-bromodeoxyuridine (BrdU), and tissue sections were analyzed by immunohistochemical staining with an antibody against BrdU. Labeled cells were restricted to the basal layer in both transgenic K5-(S249C)FGFR3 mice (A) and wild-type littermates (B). Magnification 200×.
Figure 4. Example of a somatic FGFR3 mutation in a seborrheic keratosis sample. Sequencing revealed a missense mutation in codon 652 (AAG-->ATG) of FGFR3 in DNA from seborrheic keratosis 46 and a wild-type sequence in a normal matched DNA sample.
Clinical data and FGFR3 mutations found in the 62 seborrheic keratoses
| Case # . | Age (yr) . | Sex . | Location of the tumor . | Size (mm) . | Histological type . | FGFR3 codon . | FGFR3 Mutation . |
|---|---|---|---|---|---|---|---|
| 1 | 73 | F | Trunk | 10 | Acanthotic | wt | |
| 2 | 64 | F | Face | 10 | Acanthotic | 248 | CGC→TGC |
| 3 | 76 | F | Extremity | 9 | Hyperkeratotic | wt | |
| 4 | 51 | F | Extremity | 8 | Hyperkeratotic | wt | |
| 5 | 47 | M | Face | 12 | Acanthotic | wt | |
| 6 | 51 | M | Face | 8 | Acanthotic | 373 | AGT→TGT |
| 7 | 46 | F | Extremity | 8 | Acanthotic | wt | |
| 8 | 56 | M | Trunk | 10 | Acanthotic | wt | |
| 9 | 54 | M | Face | 9 | Hyperkeratotic | 248 | CGC→TGC |
| 10 | 76 | F | Trunk | 9 | Acanthotic | 249 | TCC→TGC |
| 11a | 91 | F | Trunk | 12 | Acanthotic | 249 | TCC→TGC |
| b | Trunk | 15 | Acanthotic | 375 | TAT→TGT | ||
| 12 | 59 | F | Trunk | 15 | Acanthotic | wt | |
| 13 | 86 | M | Face | 6 | Hyperkeratotic | wt | |
| 14 | 71 | M | Face | 8 | Acanthotic | wt | |
| 15 | 66 | F | Trunk | 8 | Hyperkeratotic | wt | |
| 16 | 54 | F | Face | 10 | Acanthotic | 372 | GGC→TGC |
| 17 | 53 | F | Trunk | 7 | Acanthotic | wt | |
| 18 | 84 | F | Trunk | 8 | Hyperkeratotic | 248 | CGC→TGC |
| 19 | 43 | F | Face | 12 | Acanthotic | 652 | AAG→ATG |
| 20 | 59 | F | Trunk | 15 | Acanthotic | wt | |
| 21 | 67 | F | Face | 6 | Acanthotic | 372 | GGC→TGC |
| 22 | 57 | F | Trunk | 6 | Hyperkeratotic | wt | |
| 23 | 63 | F | Trunk | 10 | Acanthotic | wt | |
| 24 | 44 | F | Trunk | 4 | Hyperkeratotic | wt | |
| 25 | 72 | F | Trunk | 13 | Acanthotic | 375 | TAT→TGT |
| 26 | 74 | M | Face | 6 | Acanthotic | wt | |
| 27 | 40 | F | Trunk | 6 | Acanthotic | wt | |
| 28 | 88 | F | Trunk | 7 | Hyperkeratotic | wt | |
| 29 | 67 | F | Face | 7 | Acanthotic | wt | |
| 30 | 66 | F | Trunk | 8 | Hyperkeratotic | wt | |
| 31 | 85 | M | Trunk | 6 | Acanthotic | 248 | CGC→TGC |
| 32 | 83 | F | Trunk | 5 | Hyperkeratotic | wt | |
| 33 | 67 | F | Face | 8 | Acanthotic | 249 | TCC→TGC |
| 34 | 31 | F | Trunk | 3 | Acanthotic | 248 | CGC→TGC |
| 35 | 76 | F | Face | 10 | Hyperkeratotic | wt | |
| 36 | 79 | F | Face | 4 | Acanthotic | wt | |
| 37 | 49 | F | Trunk | 6 | Hyperkeratotic | wt | |
| 38 | 63 | F | Face | 15 | Hyperkeratotic | wt | |
| 39 | 76 | F | Trunk | 6 | Acanthotic | wt | |
| 40 | 52 | F | Face | 14 | Hyperkeratotic | wt | |
| 41 | 56 | F | Trunk | 10 | Acanthotic | wt | |
| 42 | 59 | M | Face | 18 | Acanthotic | wt | |
| 43 | 83 | F | Trunk | 6 | Hyperkeratotic | 372 | GGC→TGC |
| 44 | 30 | F | Trunk | 10 | Hyperkeratotic | wt | |
| 45 | 77 | F | Trunk | 4 | Hyperkeratotic | wt | |
| 46 | 31 | F | Trunk | 5 | Acanthotic | 652 | AAG→ATG |
| 47a | 79 | M | Face | 11 | Hyperkeratotic | 373 | AGT→TGT |
| b | Face | 4 | Hyperkeratotic | wt | |||
| 48 | 67 | F | Trunk | 6 | Acanthotic | 375 | TAT→TGT |
| 49 | 65 | F | Trunk | 15 | Hyperkeratotic | 373 | AGT→TGT |
| 50 | 79 | F | Trunk | 7 | Hyperkeratotic | 652 | AAG→ATG |
| 51 | 75 | F | Face | 14 | Hyperkeratotic | wt | |
| 52 | 70 | F | Face | 15 | Acanthotic | 249 | TCC→TGC |
| 53a | 79 | F | Face | 6 | Hyperkeratotic | 652 | AAG→GAG |
| b | Face | 11 | Hyperkeratotic | wt | |||
| 54 | 45 | F | Trunk | 12 | Acanthotic | wt | |
| 55a | 46 | M | Extremity | 9 | Hyperkeratotic | wt | |
| b | Trunk | 6 | Acanthotic | wt | |||
| 56 | 76 | F | Face | 9 | Acanthotic | wt | |
| 57 | 31 | M | Face | 4 | Acanthotic | 375 | TAT→TGT |
| 58 | 75 | M | Face | 10 | Hyperkeratotic | 249 | TCC→TGC |
| Case # . | Age (yr) . | Sex . | Location of the tumor . | Size (mm) . | Histological type . | FGFR3 codon . | FGFR3 Mutation . |
|---|---|---|---|---|---|---|---|
| 1 | 73 | F | Trunk | 10 | Acanthotic | wt | |
| 2 | 64 | F | Face | 10 | Acanthotic | 248 | CGC→TGC |
| 3 | 76 | F | Extremity | 9 | Hyperkeratotic | wt | |
| 4 | 51 | F | Extremity | 8 | Hyperkeratotic | wt | |
| 5 | 47 | M | Face | 12 | Acanthotic | wt | |
| 6 | 51 | M | Face | 8 | Acanthotic | 373 | AGT→TGT |
| 7 | 46 | F | Extremity | 8 | Acanthotic | wt | |
| 8 | 56 | M | Trunk | 10 | Acanthotic | wt | |
| 9 | 54 | M | Face | 9 | Hyperkeratotic | 248 | CGC→TGC |
| 10 | 76 | F | Trunk | 9 | Acanthotic | 249 | TCC→TGC |
| 11a | 91 | F | Trunk | 12 | Acanthotic | 249 | TCC→TGC |
| b | Trunk | 15 | Acanthotic | 375 | TAT→TGT | ||
| 12 | 59 | F | Trunk | 15 | Acanthotic | wt | |
| 13 | 86 | M | Face | 6 | Hyperkeratotic | wt | |
| 14 | 71 | M | Face | 8 | Acanthotic | wt | |
| 15 | 66 | F | Trunk | 8 | Hyperkeratotic | wt | |
| 16 | 54 | F | Face | 10 | Acanthotic | 372 | GGC→TGC |
| 17 | 53 | F | Trunk | 7 | Acanthotic | wt | |
| 18 | 84 | F | Trunk | 8 | Hyperkeratotic | 248 | CGC→TGC |
| 19 | 43 | F | Face | 12 | Acanthotic | 652 | AAG→ATG |
| 20 | 59 | F | Trunk | 15 | Acanthotic | wt | |
| 21 | 67 | F | Face | 6 | Acanthotic | 372 | GGC→TGC |
| 22 | 57 | F | Trunk | 6 | Hyperkeratotic | wt | |
| 23 | 63 | F | Trunk | 10 | Acanthotic | wt | |
| 24 | 44 | F | Trunk | 4 | Hyperkeratotic | wt | |
| 25 | 72 | F | Trunk | 13 | Acanthotic | 375 | TAT→TGT |
| 26 | 74 | M | Face | 6 | Acanthotic | wt | |
| 27 | 40 | F | Trunk | 6 | Acanthotic | wt | |
| 28 | 88 | F | Trunk | 7 | Hyperkeratotic | wt | |
| 29 | 67 | F | Face | 7 | Acanthotic | wt | |
| 30 | 66 | F | Trunk | 8 | Hyperkeratotic | wt | |
| 31 | 85 | M | Trunk | 6 | Acanthotic | 248 | CGC→TGC |
| 32 | 83 | F | Trunk | 5 | Hyperkeratotic | wt | |
| 33 | 67 | F | Face | 8 | Acanthotic | 249 | TCC→TGC |
| 34 | 31 | F | Trunk | 3 | Acanthotic | 248 | CGC→TGC |
| 35 | 76 | F | Face | 10 | Hyperkeratotic | wt | |
| 36 | 79 | F | Face | 4 | Acanthotic | wt | |
| 37 | 49 | F | Trunk | 6 | Hyperkeratotic | wt | |
| 38 | 63 | F | Face | 15 | Hyperkeratotic | wt | |
| 39 | 76 | F | Trunk | 6 | Acanthotic | wt | |
| 40 | 52 | F | Face | 14 | Hyperkeratotic | wt | |
| 41 | 56 | F | Trunk | 10 | Acanthotic | wt | |
| 42 | 59 | M | Face | 18 | Acanthotic | wt | |
| 43 | 83 | F | Trunk | 6 | Hyperkeratotic | 372 | GGC→TGC |
| 44 | 30 | F | Trunk | 10 | Hyperkeratotic | wt | |
| 45 | 77 | F | Trunk | 4 | Hyperkeratotic | wt | |
| 46 | 31 | F | Trunk | 5 | Acanthotic | 652 | AAG→ATG |
| 47a | 79 | M | Face | 11 | Hyperkeratotic | 373 | AGT→TGT |
| b | Face | 4 | Hyperkeratotic | wt | |||
| 48 | 67 | F | Trunk | 6 | Acanthotic | 375 | TAT→TGT |
| 49 | 65 | F | Trunk | 15 | Hyperkeratotic | 373 | AGT→TGT |
| 50 | 79 | F | Trunk | 7 | Hyperkeratotic | 652 | AAG→ATG |
| 51 | 75 | F | Face | 14 | Hyperkeratotic | wt | |
| 52 | 70 | F | Face | 15 | Acanthotic | 249 | TCC→TGC |
| 53a | 79 | F | Face | 6 | Hyperkeratotic | 652 | AAG→GAG |
| b | Face | 11 | Hyperkeratotic | wt | |||
| 54 | 45 | F | Trunk | 12 | Acanthotic | wt | |
| 55a | 46 | M | Extremity | 9 | Hyperkeratotic | wt | |
| b | Trunk | 6 | Acanthotic | wt | |||
| 56 | 76 | F | Face | 9 | Acanthotic | wt | |
| 57 | 31 | M | Face | 4 | Acanthotic | 375 | TAT→TGT |
| 58 | 75 | M | Face | 10 | Hyperkeratotic | 249 | TCC→TGC |
F: female; M: male; wt: wild-type.
Two seborrheic keratoses were analyzed for cases 11, 47, 53 and 55.
Clinical data and FGFR3 mutations found in the 62 seborrheic keratoses
| Case # . | Age (yr) . | Sex . | Location of the tumor . | Size (mm) . | Histological type . | FGFR3 codon . | FGFR3 Mutation . |
|---|---|---|---|---|---|---|---|
| 1 | 73 | F | Trunk | 10 | Acanthotic | wt | |
| 2 | 64 | F | Face | 10 | Acanthotic | 248 | CGC→TGC |
| 3 | 76 | F | Extremity | 9 | Hyperkeratotic | wt | |
| 4 | 51 | F | Extremity | 8 | Hyperkeratotic | wt | |
| 5 | 47 | M | Face | 12 | Acanthotic | wt | |
| 6 | 51 | M | Face | 8 | Acanthotic | 373 | AGT→TGT |
| 7 | 46 | F | Extremity | 8 | Acanthotic | wt | |
| 8 | 56 | M | Trunk | 10 | Acanthotic | wt | |
| 9 | 54 | M | Face | 9 | Hyperkeratotic | 248 | CGC→TGC |
| 10 | 76 | F | Trunk | 9 | Acanthotic | 249 | TCC→TGC |
| 11a | 91 | F | Trunk | 12 | Acanthotic | 249 | TCC→TGC |
| b | Trunk | 15 | Acanthotic | 375 | TAT→TGT | ||
| 12 | 59 | F | Trunk | 15 | Acanthotic | wt | |
| 13 | 86 | M | Face | 6 | Hyperkeratotic | wt | |
| 14 | 71 | M | Face | 8 | Acanthotic | wt | |
| 15 | 66 | F | Trunk | 8 | Hyperkeratotic | wt | |
| 16 | 54 | F | Face | 10 | Acanthotic | 372 | GGC→TGC |
| 17 | 53 | F | Trunk | 7 | Acanthotic | wt | |
| 18 | 84 | F | Trunk | 8 | Hyperkeratotic | 248 | CGC→TGC |
| 19 | 43 | F | Face | 12 | Acanthotic | 652 | AAG→ATG |
| 20 | 59 | F | Trunk | 15 | Acanthotic | wt | |
| 21 | 67 | F | Face | 6 | Acanthotic | 372 | GGC→TGC |
| 22 | 57 | F | Trunk | 6 | Hyperkeratotic | wt | |
| 23 | 63 | F | Trunk | 10 | Acanthotic | wt | |
| 24 | 44 | F | Trunk | 4 | Hyperkeratotic | wt | |
| 25 | 72 | F | Trunk | 13 | Acanthotic | 375 | TAT→TGT |
| 26 | 74 | M | Face | 6 | Acanthotic | wt | |
| 27 | 40 | F | Trunk | 6 | Acanthotic | wt | |
| 28 | 88 | F | Trunk | 7 | Hyperkeratotic | wt | |
| 29 | 67 | F | Face | 7 | Acanthotic | wt | |
| 30 | 66 | F | Trunk | 8 | Hyperkeratotic | wt | |
| 31 | 85 | M | Trunk | 6 | Acanthotic | 248 | CGC→TGC |
| 32 | 83 | F | Trunk | 5 | Hyperkeratotic | wt | |
| 33 | 67 | F | Face | 8 | Acanthotic | 249 | TCC→TGC |
| 34 | 31 | F | Trunk | 3 | Acanthotic | 248 | CGC→TGC |
| 35 | 76 | F | Face | 10 | Hyperkeratotic | wt | |
| 36 | 79 | F | Face | 4 | Acanthotic | wt | |
| 37 | 49 | F | Trunk | 6 | Hyperkeratotic | wt | |
| 38 | 63 | F | Face | 15 | Hyperkeratotic | wt | |
| 39 | 76 | F | Trunk | 6 | Acanthotic | wt | |
| 40 | 52 | F | Face | 14 | Hyperkeratotic | wt | |
| 41 | 56 | F | Trunk | 10 | Acanthotic | wt | |
| 42 | 59 | M | Face | 18 | Acanthotic | wt | |
| 43 | 83 | F | Trunk | 6 | Hyperkeratotic | 372 | GGC→TGC |
| 44 | 30 | F | Trunk | 10 | Hyperkeratotic | wt | |
| 45 | 77 | F | Trunk | 4 | Hyperkeratotic | wt | |
| 46 | 31 | F | Trunk | 5 | Acanthotic | 652 | AAG→ATG |
| 47a | 79 | M | Face | 11 | Hyperkeratotic | 373 | AGT→TGT |
| b | Face | 4 | Hyperkeratotic | wt | |||
| 48 | 67 | F | Trunk | 6 | Acanthotic | 375 | TAT→TGT |
| 49 | 65 | F | Trunk | 15 | Hyperkeratotic | 373 | AGT→TGT |
| 50 | 79 | F | Trunk | 7 | Hyperkeratotic | 652 | AAG→ATG |
| 51 | 75 | F | Face | 14 | Hyperkeratotic | wt | |
| 52 | 70 | F | Face | 15 | Acanthotic | 249 | TCC→TGC |
| 53a | 79 | F | Face | 6 | Hyperkeratotic | 652 | AAG→GAG |
| b | Face | 11 | Hyperkeratotic | wt | |||
| 54 | 45 | F | Trunk | 12 | Acanthotic | wt | |
| 55a | 46 | M | Extremity | 9 | Hyperkeratotic | wt | |
| b | Trunk | 6 | Acanthotic | wt | |||
| 56 | 76 | F | Face | 9 | Acanthotic | wt | |
| 57 | 31 | M | Face | 4 | Acanthotic | 375 | TAT→TGT |
| 58 | 75 | M | Face | 10 | Hyperkeratotic | 249 | TCC→TGC |
| Case # . | Age (yr) . | Sex . | Location of the tumor . | Size (mm) . | Histological type . | FGFR3 codon . | FGFR3 Mutation . |
|---|---|---|---|---|---|---|---|
| 1 | 73 | F | Trunk | 10 | Acanthotic | wt | |
| 2 | 64 | F | Face | 10 | Acanthotic | 248 | CGC→TGC |
| 3 | 76 | F | Extremity | 9 | Hyperkeratotic | wt | |
| 4 | 51 | F | Extremity | 8 | Hyperkeratotic | wt | |
| 5 | 47 | M | Face | 12 | Acanthotic | wt | |
| 6 | 51 | M | Face | 8 | Acanthotic | 373 | AGT→TGT |
| 7 | 46 | F | Extremity | 8 | Acanthotic | wt | |
| 8 | 56 | M | Trunk | 10 | Acanthotic | wt | |
| 9 | 54 | M | Face | 9 | Hyperkeratotic | 248 | CGC→TGC |
| 10 | 76 | F | Trunk | 9 | Acanthotic | 249 | TCC→TGC |
| 11a | 91 | F | Trunk | 12 | Acanthotic | 249 | TCC→TGC |
| b | Trunk | 15 | Acanthotic | 375 | TAT→TGT | ||
| 12 | 59 | F | Trunk | 15 | Acanthotic | wt | |
| 13 | 86 | M | Face | 6 | Hyperkeratotic | wt | |
| 14 | 71 | M | Face | 8 | Acanthotic | wt | |
| 15 | 66 | F | Trunk | 8 | Hyperkeratotic | wt | |
| 16 | 54 | F | Face | 10 | Acanthotic | 372 | GGC→TGC |
| 17 | 53 | F | Trunk | 7 | Acanthotic | wt | |
| 18 | 84 | F | Trunk | 8 | Hyperkeratotic | 248 | CGC→TGC |
| 19 | 43 | F | Face | 12 | Acanthotic | 652 | AAG→ATG |
| 20 | 59 | F | Trunk | 15 | Acanthotic | wt | |
| 21 | 67 | F | Face | 6 | Acanthotic | 372 | GGC→TGC |
| 22 | 57 | F | Trunk | 6 | Hyperkeratotic | wt | |
| 23 | 63 | F | Trunk | 10 | Acanthotic | wt | |
| 24 | 44 | F | Trunk | 4 | Hyperkeratotic | wt | |
| 25 | 72 | F | Trunk | 13 | Acanthotic | 375 | TAT→TGT |
| 26 | 74 | M | Face | 6 | Acanthotic | wt | |
| 27 | 40 | F | Trunk | 6 | Acanthotic | wt | |
| 28 | 88 | F | Trunk | 7 | Hyperkeratotic | wt | |
| 29 | 67 | F | Face | 7 | Acanthotic | wt | |
| 30 | 66 | F | Trunk | 8 | Hyperkeratotic | wt | |
| 31 | 85 | M | Trunk | 6 | Acanthotic | 248 | CGC→TGC |
| 32 | 83 | F | Trunk | 5 | Hyperkeratotic | wt | |
| 33 | 67 | F | Face | 8 | Acanthotic | 249 | TCC→TGC |
| 34 | 31 | F | Trunk | 3 | Acanthotic | 248 | CGC→TGC |
| 35 | 76 | F | Face | 10 | Hyperkeratotic | wt | |
| 36 | 79 | F | Face | 4 | Acanthotic | wt | |
| 37 | 49 | F | Trunk | 6 | Hyperkeratotic | wt | |
| 38 | 63 | F | Face | 15 | Hyperkeratotic | wt | |
| 39 | 76 | F | Trunk | 6 | Acanthotic | wt | |
| 40 | 52 | F | Face | 14 | Hyperkeratotic | wt | |
| 41 | 56 | F | Trunk | 10 | Acanthotic | wt | |
| 42 | 59 | M | Face | 18 | Acanthotic | wt | |
| 43 | 83 | F | Trunk | 6 | Hyperkeratotic | 372 | GGC→TGC |
| 44 | 30 | F | Trunk | 10 | Hyperkeratotic | wt | |
| 45 | 77 | F | Trunk | 4 | Hyperkeratotic | wt | |
| 46 | 31 | F | Trunk | 5 | Acanthotic | 652 | AAG→ATG |
| 47a | 79 | M | Face | 11 | Hyperkeratotic | 373 | AGT→TGT |
| b | Face | 4 | Hyperkeratotic | wt | |||
| 48 | 67 | F | Trunk | 6 | Acanthotic | 375 | TAT→TGT |
| 49 | 65 | F | Trunk | 15 | Hyperkeratotic | 373 | AGT→TGT |
| 50 | 79 | F | Trunk | 7 | Hyperkeratotic | 652 | AAG→ATG |
| 51 | 75 | F | Face | 14 | Hyperkeratotic | wt | |
| 52 | 70 | F | Face | 15 | Acanthotic | 249 | TCC→TGC |
| 53a | 79 | F | Face | 6 | Hyperkeratotic | 652 | AAG→GAG |
| b | Face | 11 | Hyperkeratotic | wt | |||
| 54 | 45 | F | Trunk | 12 | Acanthotic | wt | |
| 55a | 46 | M | Extremity | 9 | Hyperkeratotic | wt | |
| b | Trunk | 6 | Acanthotic | wt | |||
| 56 | 76 | F | Face | 9 | Acanthotic | wt | |
| 57 | 31 | M | Face | 4 | Acanthotic | 375 | TAT→TGT |
| 58 | 75 | M | Face | 10 | Hyperkeratotic | 249 | TCC→TGC |
F: female; M: male; wt: wild-type.
Two seborrheic keratoses were analyzed for cases 11, 47, 53 and 55.
FGFR3 mutations in 62 seborrheic keratoses
| Codon . | nt Position . | Exon . | Mutation . | Predicted effect . | Number of tumors . |
|---|---|---|---|---|---|
| 248 | 742 | 7 | CGC→TGC | Arg→Cys | 5 |
| 249 | 746 | 7 | TCC→TGC | Ser→Cys | 5 |
| 372 (370) | 1114 | 10 | GGC→TGC | Gly→Cys | 3 |
| 373 (371) | 1117 | 10 | AGT→TGT | Ser→Cys | 3 |
| 375 (373) | 1124 | 10 | TAT→TGT | Tyr→Cys | 4 |
| 652 (650) | 1954 | 15 | AAG→GAG | Lys→Glu | 1 |
| 652 (650) | 1955 | 15 | AAG→ATG | Lys→Met | 3 |
| Codon . | nt Position . | Exon . | Mutation . | Predicted effect . | Number of tumors . |
|---|---|---|---|---|---|
| 248 | 742 | 7 | CGC→TGC | Arg→Cys | 5 |
| 249 | 746 | 7 | TCC→TGC | Ser→Cys | 5 |
| 372 (370) | 1114 | 10 | GGC→TGC | Gly→Cys | 3 |
| 373 (371) | 1117 | 10 | AGT→TGT | Ser→Cys | 3 |
| 375 (373) | 1124 | 10 | TAT→TGT | Tyr→Cys | 4 |
| 652 (650) | 1954 | 15 | AAG→GAG | Lys→Glu | 1 |
| 652 (650) | 1955 | 15 | AAG→ATG | Lys→Met | 3 |
Codons and mutated nucleotides (nt position) are numbered according to the open reading frame of the FGFR3b cDNA. This isoform, which is the predominant form in epithelial cells, contains two more amino acids than the FGFR3c isoform, which is found in bone. Codon numbers according to the open reading frame of the FGFR3c cDNA are indicated in brackets.
FGFR3 mutations in 62 seborrheic keratoses
| Codon . | nt Position . | Exon . | Mutation . | Predicted effect . | Number of tumors . |
|---|---|---|---|---|---|
| 248 | 742 | 7 | CGC→TGC | Arg→Cys | 5 |
| 249 | 746 | 7 | TCC→TGC | Ser→Cys | 5 |
| 372 (370) | 1114 | 10 | GGC→TGC | Gly→Cys | 3 |
| 373 (371) | 1117 | 10 | AGT→TGT | Ser→Cys | 3 |
| 375 (373) | 1124 | 10 | TAT→TGT | Tyr→Cys | 4 |
| 652 (650) | 1954 | 15 | AAG→GAG | Lys→Glu | 1 |
| 652 (650) | 1955 | 15 | AAG→ATG | Lys→Met | 3 |
| Codon . | nt Position . | Exon . | Mutation . | Predicted effect . | Number of tumors . |
|---|---|---|---|---|---|
| 248 | 742 | 7 | CGC→TGC | Arg→Cys | 5 |
| 249 | 746 | 7 | TCC→TGC | Ser→Cys | 5 |
| 372 (370) | 1114 | 10 | GGC→TGC | Gly→Cys | 3 |
| 373 (371) | 1117 | 10 | AGT→TGT | Ser→Cys | 3 |
| 375 (373) | 1124 | 10 | TAT→TGT | Tyr→Cys | 4 |
| 652 (650) | 1954 | 15 | AAG→GAG | Lys→Glu | 1 |
| 652 (650) | 1955 | 15 | AAG→ATG | Lys→Met | 3 |
Codons and mutated nucleotides (nt position) are numbered according to the open reading frame of the FGFR3b cDNA. This isoform, which is the predominant form in epithelial cells, contains two more amino acids than the FGFR3c isoform, which is found in bone. Codon numbers according to the open reading frame of the FGFR3c cDNA are indicated in brackets.
References
Johnson, D.E. and Williams, L.T. (
Robertson, S.C., Tynan, J.A. and Donoghue, D.J. (
Ornitz, D.M. and Marie, P.J. (
Naski, M.C., Wang, Q., Xu, J. and Ornitz, D.M. (
Chesi, M., Nardini, E., Brents, L.A., Schrock, E., Ried, T., Kuehl, W.M. and Bergsagel, P.L. (
Richelda, R., Ronchetti, D., Baldini, L., Cro, L., Viggiano, L., Marzella, R., Rocchi, M., Otsuki, T., Lombardi, L., Maiolo, A.T. et al. (
Cappellen, D., De Oliveira, C., Ricol, D., de Medina, S., Bourdin, J., Sastre-Garau, X., Chopin, D., Thiery, J.P. and Radvanyi, F. (
Chesi, M., Brents, L.A., Ely, S.A., Bais, C., Robbiani, D.F., Mesri, E.A., Kuehl, W.M. and Bergsagel, P.L. (
Ronchetti, D., Greco, A., Compasso, S., Colombo, G., Dell'Era, P., Otsuki, T., Lombardi, L. and Neri, A. (
Li, Z., Zhu, Y.X., Plowright, E.E., Bergsagel, P.L., Chesi, M., Patterson, B., Hawley, T.S., Hawley, R.G. and Stewart, A.K. (
Torley, D., Bellus, G.A. and Munro, C.S. (
Bellus, G.A., Bamshad, M.J., Przylepa, K.A., Dorst, J., Lee, R.R., Hurko, O., Jabs, E.W., Curry, C.J., Wilcox, W.R., Lachman, R.S. et al. (
Baker, K.M., Olson, D.S., Harding, C.O. and Pauli, R.M. (
Okajima, K., Asai, K., Niwa, T., Ohki, S., Sobajima, H., Tyson, J., Malcolm, S. and Wada, Y. (
Ramirez, A., Bravo, A., Jorcano, J.L. and Vidal, M. (
Fuchs, E. and Raghavan, S. (
Billerey, C., Chopin, D., Aubriot-Lorton, M.H., Ricol, D., Gil Diez de Medina, S., Van Rhijn, B., Bralet, M.P., Lefrere-Belda, M.A., Lahaye, J.B., Abbou, C.C. et al. (
van Rhijn, B.W., van Tilborg, A.A., Lurkin, I., Bonaventure, J., de Vries, A., Thiery, J.P., van der Kwast, T.H., Zwarthoff, E.C. and Radvanyi, F. (
Tavormina, P.L., Shiang, R., Thompson, L.M., Zhu, Y.Z., Wilkin, D.J., Lachman, R.S., Wilcox, W.R., Rimoin, D.L., Cohn, D.H. and Wasmuth, J.J. (
Tavormina, P.L., Bellus, G.A., Webster, M.K., Bamshad, M.J., Fraley, A.E., McIntosh, I., Szabo, J., Jiang, W., Jabs, E.W. and Wilcox, W.R. et al. (
Nakamura, H., Hirota, S., Adachi, S., Ozaki, K., Asada, H. and Kitamura, Y. (
Li, C., Chen, L., Iwata, T., Kitagawa, M., Fu, X.Y. and Deng, C.X. (
Iwata, T., Chen, L., Li, C., Ovchinnikov, D.A., Behringer, R.R., Francomano, C.A. and Deng, C.X. (
Chen, L., Li, C., Qiao, W., Xu, X. and Deng, C. (
Iwata, T., Li, C.L., Deng, C.X. and Francomano, C.A. (
Takenaka, H., Yasuno, H. and Kishimoto, S. (
Karoui, M., Hofmann-Radvanyi, H., Zimmermann, U., Couvelard, A., Degott, C., Faridoni-Laurens, L., Ahomadegbe, J.C., Gazzeri, S., Brambilla, E., Clerici, T. et al. (
Kimura, T., Suzuki, H., Ohashi, T., Asano, K., Kiyota, H. and Eto, Y. (
Cheng, L., Darson, M., Cheville, J.C., Neumann, R.M., Zincke, H., Nehra, A. and Bostwick, D.G. (
van Rhijn, B.W., Montironi, R., Zwarthoff, E.C., Jobsis, A.C. and van der Kwast, T.H. (
Mauro, M.J., O'Dwyer, M., Heinrich, M.C. and Druker, B.J. (
Werner, S., Weinberg, W., Liao, X., Peters, K.G., Blessing, M., Yuspa, S.H., Weiner, R.L. and Williams, L.T. (