-
PDF
- Split View
-
Views
-
Cite
Cite
Akatsuki Kimura, Kazuko Matsubara, Masami Horikoshi, A Decade of Histone Acetylation: Marking Eukaryotic Chromosomes with Specific Codes, The Journal of Biochemistry, Volume 138, Issue 6, December 2005, Pages 647–662, https://doi.org/10.1093/jb/mvi184
Close - Share Icon Share
Abstract
Post-translational modification of histones, a major protein component of eukaryotic chromosomes, contributes to the epigenetic regulation of gene expression. Distinct patterns of histone modification are observed at specific chromosomal regions and affect various reactions on chromosomes (transcription, replication, repair, and recombination). Histone modification has long been proposed to have a profound effect on eukaryotic gene expression since its discovery in 1964. Verification of this idea, however, was difficult until the identification of enzymes responsible for histone modifications. Ten years ago (1995), histone acetyltransferases (HATs), which acetylate lysine residues in histone amino-terminal tail regions, were isolated. HATs are involved in the regulation of both promoter-specific transcription and long-range/chromosome-wide transcription. Analyses of HATs and other modification enzymes have revealed mechanisms of epigenetic regulation that are mediated by post-translational modifications of histones. Here we review some major advances in the field, with emphasis on the lysine specificity of the acetylation reaction and on the regulation of gene expression over broad regions.
1. Introduction: Before 1995
The nucleosome, a fundamental unit of eukaryotic chromosomes, is composed of DNA and histone proteins (Fig. 1) (1, 2). Histones have a mass roughly equal to that of the DNA which they are associated with. Each of the core histones (H2A, H2B, H3, and H4) exhibits a similar structural feature called the “histone fold,” which consists of a long central α-helix flanked by shorter helices and loops that interact with DNA (3). All the core histones have 15–30 unstructured residues at their amino termini, which are commonly referred to as “tails”; their carboxy termini consist of similarly unstructured tails.
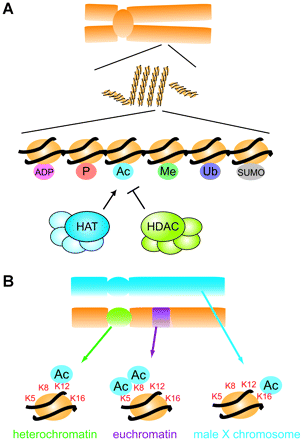
Chromosomes, chromatin, nucleosomes and histone modifications. (A) Schematic representation of a chromosome (upper), chromatin fiber (middle) and nucleosomes (lower). Histones are shown as yellow circles, and DNA is shown as a black line. The nucleosome, the fundamental unit of chromatin, consists of a histone octamer wrapped with 146 bp of DNA. Post-translational modifications of histones are shown in small circles: acetylation (Ac, blue), methylation (Me, green), phosphorylation (P, orange), ubiquitination (Ub, purple), sumoylation (SUMO, gray), and poly-ADP-ribosylation (ADP, pink). HATs (histone acetyltransferases) transfer acetyl groups to histones, and HDACs (histone deacetylases) remove them. (B) Specific acetylation of histone lysine residues in vivo. A schematic summary of the results of Turner et al. (8). K5, K8, K12, and K16 represent histone H4 lysine residues 5, 8, 12, and 16, respectively. Distinct patterns of acetylation are found in different chromosomal regions (see text for details).
The tail regions of core histones are subject to various post-translational modifications that are considered to be key reactions in the modulation of chromatin structure and function (Fig. 1A). Post-translational modifications of histones include the acetylation of lysine residues, lysine and arginine methylation, serine and threonine phosphorylation, lysine ubiquitination, lysine sumoylation, and the poly-ADP-ribosylation of glutamic acid (4, 5). Among these, acetylation has been the primary subject of research.
Histone acetylation was first discovered by Allfrey et al. in 1964 and proposed to regulate gene expression (6). This idea was supported by the observation that hyper-acetylation of histones correlates with transcriptional activation (7). Because acetylation of lysine residues neutralizes their positive charges, it was assumed that a decrease in the electrostatic interaction between DNA and histones is the major acetylation-dependent mechanism that regulates gene expression.
Turner et al. observed acetylation of distinct lysine residues in specific chromosomal regions in Drosophila melanogaster polytene nuclei (Fig. 1B) (8). Lysine 5 of histone H4 (H4-K5) or H4-K8 is frequently acetylated in euchromatic regions, where transcription is potentially active. In contrast, acetylation of H4-K12 is increased in heterochromatic regions, where transcription is potentially inactive. Acetylation of H4-K16 is found along the transcriptionally hyperactive male X chromosome (8). These observations suggested that residue-specific acetylation, rather than bulk neutralization of electrostatic charge, is important in regulation of gene expression through histone modification. They further suggested that histone modification mediates not only promoter-specific gene expression but also longer-range (and even chromosome-wide) gene expression.
Although assumed for more than 30 years, the cause-and-effect relationship between histone acetylation and transcriptional activity was not confirmed, mainly because the enzymes responsible for histone acetylation were unknown. Therefore the identification of such enzymes was a major objective in the field of eukaryotic gene expression.
2. Identification of HATs
In 1995, Brownell and Allis developed a histone acetyltransferase (HAT) assay and detected a single polypeptide of 55 kDa (p55) in macronuclear extracts of Tetrahymena thermophilia (9). Cloning of the cDNA encoding p55 revealed striking sequence similarity of the protein with a yeast transcriptional coactivator, Gcn5 (10), and this latter protein was then found to have HAT activity. The identification of a HAT in the transcriptional coactivator was a breakthrough in understanding the causal relationship between histone acetylation and gene expression.
Another gene encoding a HAT was identified in 1995. Kleff et al. screened for mutants defective in HAT activity among a collection of yeast temperature-sensitive mutants with an enzymatic assay that used fractionated cell extracts (11), which led to the discovery of the Hat1 protein. Because the hat1 mutant conferred no obvious phenotypes other than the enzymatic defect, the identification of Hat1 did not directly link histone acetylation and gene expression. However, Hat1 was found to share structural similarity with Gcn5, and both proteins were later assigned to the GNAT (GCN5-related N-acetyltransferases) superfamily (12). Hat1 was the first HAT whose tertiary structure was solved (13), thus contributing to an understanding of the relationship between HAT structure and function.
HAT activities of mammalian proteins were also reported. In 1996, HAT activity of PCAF (p300/CBP-associated factor), a protein that competes with the adenoviral oncoprotein E1A to bind to the coactivator p300/CBP, was reported (14). p300/CBP itself also turned out to be a HAT (15, 16), as did TAFII250 (CCG1/TAF1), the largest subunit of the general transcription initiation factor TFIID, which is conserved among eukaryotes (17). To date, various types of promoter-associated transcriptional coactivators and proteins in transcription machineries have been tested for HAT activity and some of these were shown to be HATs (Table 1) (18, 19). These findings strengthened the idea that localized acetylation of histones by transcription factors contributes to the activation of promoter-specific gene expression.
A list of identified histone acetyltransferases (HATs).
| Group . | HAT . | Organism . | Complex . | Possible function . | Ref. . |
|---|---|---|---|---|---|
| Gcn5 family | Gcn5 | yeast | SAGA, SLIK, SALSA, ADA, HAT-A2 | transcriptional activation | (10) |
| Gcn5L | mammal/fly | STAGA, TFTC | transcriptional activation | (95, 96) | |
| PCAF | mammal | PCAF complex | transcriptional activation | (14) | |
| MYST familya | Tip60 | mammal | TIP60 complex | transcriptional activation/DNA repair | (20) |
| HBO1 | mammal | HBO1 complex | gene expression?/DNA replication? | (97) | |
| MORF | mammal | transcriptional activation | (98) | ||
| MOZ | mammal | transcriptional activation | (99) | ||
| MOF | mammal/fly | MSL | dosage compensation | (25, 100–102) | |
| Esa1 | yeast | NuA4 | transcriptional activation | (36) | |
| Sas3 | yeast | NuA3 | transcriptional activation? | (103) | |
| Sas2 | yeast | SAS-I | anti-silencing | (41, 42, 45) | |
| Others | Hat1 | yeast | Hat1/2 complex | histone deposition | (11) |
| p300/CBP | mammal | transcriptional activation | (15, 16) | ||
| TAFII250 (TAF1) | mammal/fly/yeast | TFIID | RNA pol II transcription | (17) | |
| ACTR/SRC-1 | mammal | transcriptional activation | (104, 105) | ||
| Elp3 | yeast | elongator | transcriptional elongation | (106) | |
| hTFIIIC110 | mammal | TFIIIC | RNA pol III transcription | (107) | |
| hTFIIIC90 | mammal | TFIIIC | RNA pol III transcription | (108) | |
| Hpa2 | yeast | ? | (109) | ||
| Nut1 | yeast | mediator | RNA pol II transcription | (110) | |
| ATF-2 | mammal | transcriptional activation | (111) |
| Group . | HAT . | Organism . | Complex . | Possible function . | Ref. . |
|---|---|---|---|---|---|
| Gcn5 family | Gcn5 | yeast | SAGA, SLIK, SALSA, ADA, HAT-A2 | transcriptional activation | (10) |
| Gcn5L | mammal/fly | STAGA, TFTC | transcriptional activation | (95, 96) | |
| PCAF | mammal | PCAF complex | transcriptional activation | (14) | |
| MYST familya | Tip60 | mammal | TIP60 complex | transcriptional activation/DNA repair | (20) |
| HBO1 | mammal | HBO1 complex | gene expression?/DNA replication? | (97) | |
| MORF | mammal | transcriptional activation | (98) | ||
| MOZ | mammal | transcriptional activation | (99) | ||
| MOF | mammal/fly | MSL | dosage compensation | (25, 100–102) | |
| Esa1 | yeast | NuA4 | transcriptional activation | (36) | |
| Sas3 | yeast | NuA3 | transcriptional activation? | (103) | |
| Sas2 | yeast | SAS-I | anti-silencing | (41, 42, 45) | |
| Others | Hat1 | yeast | Hat1/2 complex | histone deposition | (11) |
| p300/CBP | mammal | transcriptional activation | (15, 16) | ||
| TAFII250 (TAF1) | mammal/fly/yeast | TFIID | RNA pol II transcription | (17) | |
| ACTR/SRC-1 | mammal | transcriptional activation | (104, 105) | ||
| Elp3 | yeast | elongator | transcriptional elongation | (106) | |
| hTFIIIC110 | mammal | TFIIIC | RNA pol III transcription | (107) | |
| hTFIIIC90 | mammal | TFIIIC | RNA pol III transcription | (108) | |
| Hpa2 | yeast | ? | (109) | ||
| Nut1 | yeast | mediator | RNA pol II transcription | (110) | |
| ATF-2 | mammal | transcriptional activation | (111) |
A list of identified histone acetyltransferases (HATs).
| Group . | HAT . | Organism . | Complex . | Possible function . | Ref. . |
|---|---|---|---|---|---|
| Gcn5 family | Gcn5 | yeast | SAGA, SLIK, SALSA, ADA, HAT-A2 | transcriptional activation | (10) |
| Gcn5L | mammal/fly | STAGA, TFTC | transcriptional activation | (95, 96) | |
| PCAF | mammal | PCAF complex | transcriptional activation | (14) | |
| MYST familya | Tip60 | mammal | TIP60 complex | transcriptional activation/DNA repair | (20) |
| HBO1 | mammal | HBO1 complex | gene expression?/DNA replication? | (97) | |
| MORF | mammal | transcriptional activation | (98) | ||
| MOZ | mammal | transcriptional activation | (99) | ||
| MOF | mammal/fly | MSL | dosage compensation | (25, 100–102) | |
| Esa1 | yeast | NuA4 | transcriptional activation | (36) | |
| Sas3 | yeast | NuA3 | transcriptional activation? | (103) | |
| Sas2 | yeast | SAS-I | anti-silencing | (41, 42, 45) | |
| Others | Hat1 | yeast | Hat1/2 complex | histone deposition | (11) |
| p300/CBP | mammal | transcriptional activation | (15, 16) | ||
| TAFII250 (TAF1) | mammal/fly/yeast | TFIID | RNA pol II transcription | (17) | |
| ACTR/SRC-1 | mammal | transcriptional activation | (104, 105) | ||
| Elp3 | yeast | elongator | transcriptional elongation | (106) | |
| hTFIIIC110 | mammal | TFIIIC | RNA pol III transcription | (107) | |
| hTFIIIC90 | mammal | TFIIIC | RNA pol III transcription | (108) | |
| Hpa2 | yeast | ? | (109) | ||
| Nut1 | yeast | mediator | RNA pol II transcription | (110) | |
| ATF-2 | mammal | transcriptional activation | (111) |
| Group . | HAT . | Organism . | Complex . | Possible function . | Ref. . |
|---|---|---|---|---|---|
| Gcn5 family | Gcn5 | yeast | SAGA, SLIK, SALSA, ADA, HAT-A2 | transcriptional activation | (10) |
| Gcn5L | mammal/fly | STAGA, TFTC | transcriptional activation | (95, 96) | |
| PCAF | mammal | PCAF complex | transcriptional activation | (14) | |
| MYST familya | Tip60 | mammal | TIP60 complex | transcriptional activation/DNA repair | (20) |
| HBO1 | mammal | HBO1 complex | gene expression?/DNA replication? | (97) | |
| MORF | mammal | transcriptional activation | (98) | ||
| MOZ | mammal | transcriptional activation | (99) | ||
| MOF | mammal/fly | MSL | dosage compensation | (25, 100–102) | |
| Esa1 | yeast | NuA4 | transcriptional activation | (36) | |
| Sas3 | yeast | NuA3 | transcriptional activation? | (103) | |
| Sas2 | yeast | SAS-I | anti-silencing | (41, 42, 45) | |
| Others | Hat1 | yeast | Hat1/2 complex | histone deposition | (11) |
| p300/CBP | mammal | transcriptional activation | (15, 16) | ||
| TAFII250 (TAF1) | mammal/fly/yeast | TFIID | RNA pol II transcription | (17) | |
| ACTR/SRC-1 | mammal | transcriptional activation | (104, 105) | ||
| Elp3 | yeast | elongator | transcriptional elongation | (106) | |
| hTFIIIC110 | mammal | TFIIIC | RNA pol III transcription | (107) | |
| hTFIIIC90 | mammal | TFIIIC | RNA pol III transcription | (108) | |
| Hpa2 | yeast | ? | (109) | ||
| Nut1 | yeast | mediator | RNA pol II transcription | (110) | |
| ATF-2 | mammal | transcriptional activation | (111) |
Meanwhile, our group identified a novel class of HATs that regulates long-range/chromosome-wide gene expression (20). A member of the MYST (MOZ, YBF2/SAS3, SAS2, Tip60) family of proteins, Tip60, was shown to possess HAT activity with novel substrate specificity. The HAT activities of other members of the MYST family, including Esa1, a yeast counterpart of Tip60, have been confirmed (Table 1). HAT activities for MYST family proteins were anticipated since they contain sequences similar to the acetyl-CoA binding motif (21) in an evolutionarily conserved region, the MYST domain (20). Interestingly, Esa1 has a sequence motif (the “ER motif ”) also found in a histone deacetylase (HDAC), yeast Rpd3 (22). Other MYST family members, yeast Sas2 and Sas3, are involved in long-range gene repression dependent on chromosomal location (23, 24), and Drosophila MOF is involved in hyperactivation of the male X chromosome (25). The identification of MYST proteins as HATs linked histone acetylation and long-range/chromosome-wide gene expression.
3. The residue-specificity of histone acetylation: A two-step classification and the allocation strategy
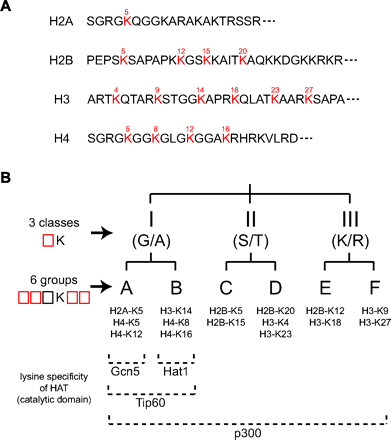
Many, but not all, lysines in the amino-terminal tail (N-tail) regions of core histones are acetylated in vivo (Fig. 2A) (26, 27). The site specificity of identified HATs is also divergent (18, 28). Analysis of primary sequences in the vicinity of lysines of core histone N-tails revealed that lysines acetylated in vivo can be classified into three classes and six groups (Fig. 2B) (28). This classification distinguishes lysines acetylated in vivo from others, and fits well with the in vitro site specificity of the catalytic domains of HATs. Therefore, it has been hypothesized that the catalytic domains of HATs recognize classes and groups according to this “two-step classification” hypothesis (28, 29).
Acetylation sites in histone amino-terminal tails and the “two-step classification” hypothesis. (A) Primary structures of histone tails in human cells. Acetylation sites, shown in red, were determined through primary structural analyses of histones in cellular extracts (26, 27). Acetylation of most of these lysines has also been detected using specific antibodies (e.g., Ref. 40). These sites coincide with those acetylated by identified HATs (e.g., Ref. 163). Recently, acetylation of four additional lysines in the amino-terminal tail regions of calf thymus histones has been detected (H2A-K9, H2A-K13, H2A-K15 and H4-K20) (164). Further analysis is required before concluding that these lysines are acetylated in vivo. (B) Two-step classification hypothesis. Lysines acetylated in vivo (shown in red in A) are subject to a “two-step classification.” Classes I to III are defined by residues located amino-terminal to acetylated lysines: glycine or alanine (G/A) for class I, serine or threonine (S/T) for class II, and lysine or arginine (K/R) for class III. Each class is subdivided into two groups according to additional flanking residues (groups A to F). For details, see (28). Of the four additional lysines that may be acetylated in vivo (164), the flanking sequences of three (H2A-K9, H2A-K13 and H2A-K15) are similar to those of classified lysines. H2A-K9 can be considered as a group B residue (28), and it is acetylated by a HAT in vitro (29); and H2A-K13 and H2A-K15 can be classified as group A lysines. Further analyses of the acetylation status of these lysine residues and comparison of obtained experimental results with the provisional classification will be of interest.
The hypothesis provides several insights into how HATs select specific lysines. First, the flanking sequences alone are unlikely to determine whether the lysine will be acetylated by a HAT. Some lysines that are not acetylated by HATs in the non-N-tail regions of histones or other proteins meet the classification criteria. HATs also acetylate non-histone substrates (Table 2). Some lysines in these substrates do not fall into any group in the classification. Because the higher order structure of the N-tail regions is considered flexible (3), this structure might be important for the N-tails of histones and non-histone substrates to be acetylated in addition to the flanking sequences of the target lysines. Second, the hypothesis provides a possible explanation for the broad but non-random specificity of Tip60 or p300 (Fig. 2B). For example, there is no apparent and strict consensus sequence for substrate recognition by a class I-specific HAT like Tip60. Because the ‘class I’ lysines consist solely of ‘group A’ and ‘group B’ lysines, the substrate recognition surfaces of the class I-specific HATs may consist of a surface that can recognize both sequences or two distinct surfaces of which one recognizes the consensus sequence of group A and the other that of group B. This speculation, based on the proposed classification, can explain specific lysine selection by HATs without an apparent target consensus sequence.
A list of nonhistone substrates for HATs.a
| Function . | Substrate . | Enzyme . | Effect of acetylation . | Ref. . |
|---|---|---|---|---|
| transcription factor | Sp1 | p300 | enhancement of DNA binding through interaction with p300, but not acetylation by p300 | (114) |
| KLF5 | p300 | enhancement of transcription activity | (115) | |
| FOXO1 | CBP | inhibition of transcription activity | (116) | |
| MEF2C | p300 | enhancement of DNA binding, transcription and myogenic differentiation | (117) | |
| SRY | p300 | increase in importin β binding and participate nuclear localization | (118) | |
| GATA-4 | p300 | enhancement of DNA binding, participates in transcription and involved in differentiation of embryonic stem cells into cardiac myocytes | (119, 120) | |
| HNF-6 | CBP | increase in stability and stimulate transcription | (121) | |
| signaling regulator | Stat3 | p300 | stimulation of DNA binding and transcription | (122) |
| Function . | Substrate . | Enzyme . | Effect of acetylation . | Ref. . |
|---|---|---|---|---|
| transcription factor | Sp1 | p300 | enhancement of DNA binding through interaction with p300, but not acetylation by p300 | (114) |
| KLF5 | p300 | enhancement of transcription activity | (115) | |
| FOXO1 | CBP | inhibition of transcription activity | (116) | |
| MEF2C | p300 | enhancement of DNA binding, transcription and myogenic differentiation | (117) | |
| SRY | p300 | increase in importin β binding and participate nuclear localization | (118) | |
| GATA-4 | p300 | enhancement of DNA binding, participates in transcription and involved in differentiation of embryonic stem cells into cardiac myocytes | (119, 120) | |
| HNF-6 | CBP | increase in stability and stimulate transcription | (121) | |
| signaling regulator | Stat3 | p300 | stimulation of DNA binding and transcription | (122) |
A comprehensive list can be obtained from Yang (2004) (123). Non-histone substrates that do not appear in the list of Yang (2004), are listed. The other non-histone substrates include chromosomal proteins, HAT autoacetylations, chromatin remodelers, transcriptional coregulators, general transcription factors, DNA metabolic enzymes, apoptosis regulator, nuclear import receptor, and viral proteins (123).
A list of nonhistone substrates for HATs.a
| Function . | Substrate . | Enzyme . | Effect of acetylation . | Ref. . |
|---|---|---|---|---|
| transcription factor | Sp1 | p300 | enhancement of DNA binding through interaction with p300, but not acetylation by p300 | (114) |
| KLF5 | p300 | enhancement of transcription activity | (115) | |
| FOXO1 | CBP | inhibition of transcription activity | (116) | |
| MEF2C | p300 | enhancement of DNA binding, transcription and myogenic differentiation | (117) | |
| SRY | p300 | increase in importin β binding and participate nuclear localization | (118) | |
| GATA-4 | p300 | enhancement of DNA binding, participates in transcription and involved in differentiation of embryonic stem cells into cardiac myocytes | (119, 120) | |
| HNF-6 | CBP | increase in stability and stimulate transcription | (121) | |
| signaling regulator | Stat3 | p300 | stimulation of DNA binding and transcription | (122) |
| Function . | Substrate . | Enzyme . | Effect of acetylation . | Ref. . |
|---|---|---|---|---|
| transcription factor | Sp1 | p300 | enhancement of DNA binding through interaction with p300, but not acetylation by p300 | (114) |
| KLF5 | p300 | enhancement of transcription activity | (115) | |
| FOXO1 | CBP | inhibition of transcription activity | (116) | |
| MEF2C | p300 | enhancement of DNA binding, transcription and myogenic differentiation | (117) | |
| SRY | p300 | increase in importin β binding and participate nuclear localization | (118) | |
| GATA-4 | p300 | enhancement of DNA binding, participates in transcription and involved in differentiation of embryonic stem cells into cardiac myocytes | (119, 120) | |
| HNF-6 | CBP | increase in stability and stimulate transcription | (121) | |
| signaling regulator | Stat3 | p300 | stimulation of DNA binding and transcription | (122) |
A comprehensive list can be obtained from Yang (2004) (123). Non-histone substrates that do not appear in the list of Yang (2004), are listed. The other non-histone substrates include chromosomal proteins, HAT autoacetylations, chromatin remodelers, transcriptional coregulators, general transcription factors, DNA metabolic enzymes, apoptosis regulator, nuclear import receptor, and viral proteins (123).
Most HATs form multisubunit complexes. In 1998, subunit compositions of human PCAF complex and yeast Gcn5 complexes (SAGA and ADA) were reported (30, 31). To date, subunits of various HAT complexes have been identified and revealed to play critical roles in regulating HAT activity, for example by targeting activity to specific chromosomal regions (Table 3) (32). The formation of multisubunit complexes also modulates the substrate specificity of HATs. The catalytic domains of most HATs alone are unable to acetylate the histones in the nucleosomal context. HAT complexes, however, can acetylate nucleosomal histones, and specific subunits of these complexes are required for targeting them to nucleosomes. For example, Gcn5 alone is unable to acetylate nucleosomal histones efficiently, whereas the Gcn5 complexes ADA and SAGA can (33). Ada2 and Ada3, subunits common to these two complexes, are required for their association with and acetylation of nucleosomal histones (34). Similarly, the Epl1 and Yng2 subunits of the Esa1 complex NuA4 are sufficient for Esa1 to acetylate nucleosomal histones (35).
A list of HAT complexes.
| complex . | Gcn5 family . | . | . | . | . | . | . | MYST family . | . | . | . | . | Others . | . | . | . | . | . | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| . | SAGA . | SLIK . | ADA . | HAT-A2 . | STAGA . | TFTC . | PCAF . | NuA4 . | NuA3 . | SAS-I . | TIP60 . | MSLd . | Hat1/2 . | Elongator . | TFIIDe . | TFIIICf . | ACTR/SRC-1 . | Mediator . | |||||||||||||||
| organism . | yeast . | yeast . | yeast . | yeast . | mammal . | mammal . | mammal . | yeast . | yeast . | yeast . | mammal . | fly . | yeast . | yeast . | mammal/fly/yeast . | mammal . | mammal . | yeast . | |||||||||||||||
| HAT vs. free histonea . | . | . | . | . | . | . | . | H2A . | H2A . | N.D.c . | H2A . | H2A . | H2A . | H2A . | . | . | . | N.A.c . | |||||||||||||||
| . | . | . | . | . | . | . | . | . | . | . | . | . | . | H2B . | . | . | . | . | |||||||||||||||
| . | H3 . | H3 . | H3 . | H3 . | H3 . | H3 . | H3 . | H3 . | H3 . | . | H3 . | H3 . | . | H3 . | H3 . | H3g . | H3 . | . | |||||||||||||||
. | H4 . | H4 . | H4 . | H4 . | H4 . | H4 . | H4 . | H4 . | H4 . | . | H4 . | H4 . | H4 . | H4 . | H4 . | . | H4 . | . | |||||||||||||||
| HAT complex vs. nucleosomal histonea . | H2B . | H2B . | H2B . | . | H2B . | . | . | H2A . | . | . | H2A . | . | N.D.c . | . | N.D.c . | H2A . | N.A.c . | . | |||||||||||||||
| . | H3 . | H3 . | H3 . | H3 . | H3 . | H3 . | H3 . | . | H3 . | . | . | . | . | H3 . | . | H3 . | . | H3 . | |||||||||||||||
| . | . | . | . | . | . | . | H4 . | H4 . | . | H4 . | H4 . | H4 . | . | H4 . | . | H4 . | . | H4 . | |||||||||||||||
| catalytic subunit(s) . | Gcn5 . | Gcn5 . | Gcn5 . | Gcn5 . | Gcn5L . | Gcn5L . | PCAF . | Esa1 . | Sas3 . | Sas2 . | Tip60 . | MOF . | Hat1 . | Elp3 . | TAFII250 (TAF1) . | TFIIIC220 TFIIIC110 TFIIIC90 . | ACTR/SRC-1 . | MED5 (Nut1) . | |||||||||||||||
| other . | Tral . | Tral . | . | . | TRRAP . | TRRAP . | PAF400 . | Yaf9 . | TAF14 . | Sas5 . | GAS41 . | . | Hat2 . | Elp1 . | TAF2 . | TFIIIC102 . | CBP . | MED1 . | |||||||||||||||
| subunitsb . | Ada1 . | Ada1 . | . | . | STAF42 . | . | . | Yng2 . | Yng1 . | . | ING3 . | . | Hif1 . | Elp2 . | TAF3 . | TFIIIC63 . | SRC-2 . | MED2 . | |||||||||||||||
| Ada2 | Ada2 | Ada2 | Ada2 | Ada2 | Arp4 | BAF53 | Elp4 | TAF4 | SS-A | MED3 | |||||||||||||||||||||||
| Ada3 | Ada3 | Ada3 | Ada3 | STAF54 | Ada3 | Ada3 | Epl1 | EPC | Elp5 | TAF5 | IKKβ | MED4 | |||||||||||||||||||||
| Ada5 | Ada5 | EPC-like | Elp6 | TAF6 | IKKγ | MED6 | |||||||||||||||||||||||||||
| Spt3 | Spt3 | Spt3 | Spt3 | Spt3 | Eaf3 | MRG15 | MSL3 | TAF7 | MED7 | ||||||||||||||||||||||||
| Spt7 | Spt7 | STAF65γ | MRGX | TAF8 | MED8 | ||||||||||||||||||||||||||||
| Spt8 | Eaf7 | MRGBP | TAF9 | MED9 | |||||||||||||||||||||||||||||
| TAF2 | Eaf6 | FLJ11730 | TAF10 | MED10 | |||||||||||||||||||||||||||||
| TAF4 | Eaf2 | DMAP1 | TAF11 | MED11 | |||||||||||||||||||||||||||||
| TAF5 | TAF5 | TAF5 | Tra1 | TRRAP | TAF12 | MED12 | |||||||||||||||||||||||||||
| TAF5L | TAF5L | TAF5L | Act1 | actin | TAF13 | MED13 | |||||||||||||||||||||||||||
| TAF6 | TAF6 | TAF6 | Eaf1 | P400 | TAF14 | MED14 | |||||||||||||||||||||||||||
| TAF6L | TAF6L | TAF6L | Eaf5 | TBP | MED15 | ||||||||||||||||||||||||||||
| TAF9 | TAF9 | TAF9 | TAF9 | TAF9 | Sas4 | MED16 | |||||||||||||||||||||||||||
| TAF10 | TAF10 | TAF10 | TAF10 | TAF10 | YL1 | MED17 | |||||||||||||||||||||||||||
| TAF12 | TAF12 | TAF12 | TAF12 | TAF12 | TIP49a | MED18 | |||||||||||||||||||||||||||
| Sgf29 | TIP49b | MED19 | |||||||||||||||||||||||||||||||
| Sgf73 | SCA7 | SCA7 | TRCp120 | MED20 | |||||||||||||||||||||||||||||
| Ubp8 | Ubp8 | MLE | MED21 | ||||||||||||||||||||||||||||||
| Sin4 | MSL1 | MED22 | |||||||||||||||||||||||||||||||
| Rtg2 | MSL2 | CDK8 | |||||||||||||||||||||||||||||||
| Ahc1 | Jil1 | CycC | |||||||||||||||||||||||||||||||
| STAF36 | roX RNA | ||||||||||||||||||||||||||||||||
| STAF46 | |||||||||||||||||||||||||||||||||
| STAF55 | |||||||||||||||||||||||||||||||||
| STAF60 | |||||||||||||||||||||||||||||||||
| SAP130 | SAP130 | ||||||||||||||||||||||||||||||||
| Ref.h | (31, 33) | (124, 125) | (31, 33) | (126) | (127, 128) | (129) | (30) | (37) | (43) | (45, 46, 130, 131) | (132–134) | (101) | (135, 136) | (137, 138) | (139) | (107) | (140) | (110) | |||||||||||||||
| complex . | Gcn5 family . | . | . | . | . | . | . | MYST family . | . | . | . | . | Others . | . | . | . | . | . | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| . | SAGA . | SLIK . | ADA . | HAT-A2 . | STAGA . | TFTC . | PCAF . | NuA4 . | NuA3 . | SAS-I . | TIP60 . | MSLd . | Hat1/2 . | Elongator . | TFIIDe . | TFIIICf . | ACTR/SRC-1 . | Mediator . | |||||||||||||||
| organism . | yeast . | yeast . | yeast . | yeast . | mammal . | mammal . | mammal . | yeast . | yeast . | yeast . | mammal . | fly . | yeast . | yeast . | mammal/fly/yeast . | mammal . | mammal . | yeast . | |||||||||||||||
| HAT vs. free histonea . | . | . | . | . | . | . | . | H2A . | H2A . | N.D.c . | H2A . | H2A . | H2A . | H2A . | . | . | . | N.A.c . | |||||||||||||||
| . | . | . | . | . | . | . | . | . | . | . | . | . | . | H2B . | . | . | . | . | |||||||||||||||
| . | H3 . | H3 . | H3 . | H3 . | H3 . | H3 . | H3 . | H3 . | H3 . | . | H3 . | H3 . | . | H3 . | H3 . | H3g . | H3 . | . | |||||||||||||||
. | H4 . | H4 . | H4 . | H4 . | H4 . | H4 . | H4 . | H4 . | H4 . | . | H4 . | H4 . | H4 . | H4 . | H4 . | . | H4 . | . | |||||||||||||||
| HAT complex vs. nucleosomal histonea . | H2B . | H2B . | H2B . | . | H2B . | . | . | H2A . | . | . | H2A . | . | N.D.c . | . | N.D.c . | H2A . | N.A.c . | . | |||||||||||||||
| . | H3 . | H3 . | H3 . | H3 . | H3 . | H3 . | H3 . | . | H3 . | . | . | . | . | H3 . | . | H3 . | . | H3 . | |||||||||||||||
| . | . | . | . | . | . | . | H4 . | H4 . | . | H4 . | H4 . | H4 . | . | H4 . | . | H4 . | . | H4 . | |||||||||||||||
| catalytic subunit(s) . | Gcn5 . | Gcn5 . | Gcn5 . | Gcn5 . | Gcn5L . | Gcn5L . | PCAF . | Esa1 . | Sas3 . | Sas2 . | Tip60 . | MOF . | Hat1 . | Elp3 . | TAFII250 (TAF1) . | TFIIIC220 TFIIIC110 TFIIIC90 . | ACTR/SRC-1 . | MED5 (Nut1) . | |||||||||||||||
| other . | Tral . | Tral . | . | . | TRRAP . | TRRAP . | PAF400 . | Yaf9 . | TAF14 . | Sas5 . | GAS41 . | . | Hat2 . | Elp1 . | TAF2 . | TFIIIC102 . | CBP . | MED1 . | |||||||||||||||
| subunitsb . | Ada1 . | Ada1 . | . | . | STAF42 . | . | . | Yng2 . | Yng1 . | . | ING3 . | . | Hif1 . | Elp2 . | TAF3 . | TFIIIC63 . | SRC-2 . | MED2 . | |||||||||||||||
| Ada2 | Ada2 | Ada2 | Ada2 | Ada2 | Arp4 | BAF53 | Elp4 | TAF4 | SS-A | MED3 | |||||||||||||||||||||||
| Ada3 | Ada3 | Ada3 | Ada3 | STAF54 | Ada3 | Ada3 | Epl1 | EPC | Elp5 | TAF5 | IKKβ | MED4 | |||||||||||||||||||||
| Ada5 | Ada5 | EPC-like | Elp6 | TAF6 | IKKγ | MED6 | |||||||||||||||||||||||||||
| Spt3 | Spt3 | Spt3 | Spt3 | Spt3 | Eaf3 | MRG15 | MSL3 | TAF7 | MED7 | ||||||||||||||||||||||||
| Spt7 | Spt7 | STAF65γ | MRGX | TAF8 | MED8 | ||||||||||||||||||||||||||||
| Spt8 | Eaf7 | MRGBP | TAF9 | MED9 | |||||||||||||||||||||||||||||
| TAF2 | Eaf6 | FLJ11730 | TAF10 | MED10 | |||||||||||||||||||||||||||||
| TAF4 | Eaf2 | DMAP1 | TAF11 | MED11 | |||||||||||||||||||||||||||||
| TAF5 | TAF5 | TAF5 | Tra1 | TRRAP | TAF12 | MED12 | |||||||||||||||||||||||||||
| TAF5L | TAF5L | TAF5L | Act1 | actin | TAF13 | MED13 | |||||||||||||||||||||||||||
| TAF6 | TAF6 | TAF6 | Eaf1 | P400 | TAF14 | MED14 | |||||||||||||||||||||||||||
| TAF6L | TAF6L | TAF6L | Eaf5 | TBP | MED15 | ||||||||||||||||||||||||||||
| TAF9 | TAF9 | TAF9 | TAF9 | TAF9 | Sas4 | MED16 | |||||||||||||||||||||||||||
| TAF10 | TAF10 | TAF10 | TAF10 | TAF10 | YL1 | MED17 | |||||||||||||||||||||||||||
| TAF12 | TAF12 | TAF12 | TAF12 | TAF12 | TIP49a | MED18 | |||||||||||||||||||||||||||
| Sgf29 | TIP49b | MED19 | |||||||||||||||||||||||||||||||
| Sgf73 | SCA7 | SCA7 | TRCp120 | MED20 | |||||||||||||||||||||||||||||
| Ubp8 | Ubp8 | MLE | MED21 | ||||||||||||||||||||||||||||||
| Sin4 | MSL1 | MED22 | |||||||||||||||||||||||||||||||
| Rtg2 | MSL2 | CDK8 | |||||||||||||||||||||||||||||||
| Ahc1 | Jil1 | CycC | |||||||||||||||||||||||||||||||
| STAF36 | roX RNA | ||||||||||||||||||||||||||||||||
| STAF46 | |||||||||||||||||||||||||||||||||
| STAF55 | |||||||||||||||||||||||||||||||||
| STAF60 | |||||||||||||||||||||||||||||||||
| SAP130 | SAP130 | ||||||||||||||||||||||||||||||||
| Ref.h | (31, 33) | (124, 125) | (31, 33) | (126) | (127, 128) | (129) | (30) | (37) | (43) | (45, 46, 130, 131) | (132–134) | (101) | (135, 136) | (137, 138) | (139) | (107) | (140) | (110) | |||||||||||||||
“HAT” indicates that a monomer of the catalytic subunit was examined as an enzyme, while “HAT complex” indicates that the complex forms were used. For the substrates, “free histones” and “nucleosomal histones” indicate that the corresponding forms of histones were used. The references for the specificity of the catalytic subunits alone against free histones are shown in Table 1.
Subunits conserved among complexes of the Gcn5 family and MYST family are represented on horizontal lines as in (32). Unified nomenclature for TFIID and mediator subunits is used (141, 142).
N.D. = not detected, N.A. = not analyzed.
A catalytic domain of MOF (aa. 518–827) acetylates H2A/H3/H4 (101), whereas the full-length MOF shows a strong preference for histone H4 (100). An N-terminal domain is involved in this gain of specificity (100). The full-length MOF is also capable of acetylating of nucleosomal histones with a strong preference for lysine 16 of histone H4 (100).
HAT activity toward nucleosomal histones of a TAFII250-containing complex (TFIIDβ) has not been detected (129). Subunit composition in yeast is shown.
Recombinant TFIIIC90 alone possesses a HAT activity toward nucleosomal histones with a strong preference for histone H3 (108).
Specificity of TFIIIC90. Specificity of TFIIIC220 or TFIIIC110 alone toward free histones has yet to be analyzed.
A list of HAT complexes.
| complex . | Gcn5 family . | . | . | . | . | . | . | MYST family . | . | . | . | . | Others . | . | . | . | . | . | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| . | SAGA . | SLIK . | ADA . | HAT-A2 . | STAGA . | TFTC . | PCAF . | NuA4 . | NuA3 . | SAS-I . | TIP60 . | MSLd . | Hat1/2 . | Elongator . | TFIIDe . | TFIIICf . | ACTR/SRC-1 . | Mediator . | |||||||||||||||
| organism . | yeast . | yeast . | yeast . | yeast . | mammal . | mammal . | mammal . | yeast . | yeast . | yeast . | mammal . | fly . | yeast . | yeast . | mammal/fly/yeast . | mammal . | mammal . | yeast . | |||||||||||||||
| HAT vs. free histonea . | . | . | . | . | . | . | . | H2A . | H2A . | N.D.c . | H2A . | H2A . | H2A . | H2A . | . | . | . | N.A.c . | |||||||||||||||
| . | . | . | . | . | . | . | . | . | . | . | . | . | . | H2B . | . | . | . | . | |||||||||||||||
| . | H3 . | H3 . | H3 . | H3 . | H3 . | H3 . | H3 . | H3 . | H3 . | . | H3 . | H3 . | . | H3 . | H3 . | H3g . | H3 . | . | |||||||||||||||
. | H4 . | H4 . | H4 . | H4 . | H4 . | H4 . | H4 . | H4 . | H4 . | . | H4 . | H4 . | H4 . | H4 . | H4 . | . | H4 . | . | |||||||||||||||
| HAT complex vs. nucleosomal histonea . | H2B . | H2B . | H2B . | . | H2B . | . | . | H2A . | . | . | H2A . | . | N.D.c . | . | N.D.c . | H2A . | N.A.c . | . | |||||||||||||||
| . | H3 . | H3 . | H3 . | H3 . | H3 . | H3 . | H3 . | . | H3 . | . | . | . | . | H3 . | . | H3 . | . | H3 . | |||||||||||||||
| . | . | . | . | . | . | . | H4 . | H4 . | . | H4 . | H4 . | H4 . | . | H4 . | . | H4 . | . | H4 . | |||||||||||||||
| catalytic subunit(s) . | Gcn5 . | Gcn5 . | Gcn5 . | Gcn5 . | Gcn5L . | Gcn5L . | PCAF . | Esa1 . | Sas3 . | Sas2 . | Tip60 . | MOF . | Hat1 . | Elp3 . | TAFII250 (TAF1) . | TFIIIC220 TFIIIC110 TFIIIC90 . | ACTR/SRC-1 . | MED5 (Nut1) . | |||||||||||||||
| other . | Tral . | Tral . | . | . | TRRAP . | TRRAP . | PAF400 . | Yaf9 . | TAF14 . | Sas5 . | GAS41 . | . | Hat2 . | Elp1 . | TAF2 . | TFIIIC102 . | CBP . | MED1 . | |||||||||||||||
| subunitsb . | Ada1 . | Ada1 . | . | . | STAF42 . | . | . | Yng2 . | Yng1 . | . | ING3 . | . | Hif1 . | Elp2 . | TAF3 . | TFIIIC63 . | SRC-2 . | MED2 . | |||||||||||||||
| Ada2 | Ada2 | Ada2 | Ada2 | Ada2 | Arp4 | BAF53 | Elp4 | TAF4 | SS-A | MED3 | |||||||||||||||||||||||
| Ada3 | Ada3 | Ada3 | Ada3 | STAF54 | Ada3 | Ada3 | Epl1 | EPC | Elp5 | TAF5 | IKKβ | MED4 | |||||||||||||||||||||
| Ada5 | Ada5 | EPC-like | Elp6 | TAF6 | IKKγ | MED6 | |||||||||||||||||||||||||||
| Spt3 | Spt3 | Spt3 | Spt3 | Spt3 | Eaf3 | MRG15 | MSL3 | TAF7 | MED7 | ||||||||||||||||||||||||
| Spt7 | Spt7 | STAF65γ | MRGX | TAF8 | MED8 | ||||||||||||||||||||||||||||
| Spt8 | Eaf7 | MRGBP | TAF9 | MED9 | |||||||||||||||||||||||||||||
| TAF2 | Eaf6 | FLJ11730 | TAF10 | MED10 | |||||||||||||||||||||||||||||
| TAF4 | Eaf2 | DMAP1 | TAF11 | MED11 | |||||||||||||||||||||||||||||
| TAF5 | TAF5 | TAF5 | Tra1 | TRRAP | TAF12 | MED12 | |||||||||||||||||||||||||||
| TAF5L | TAF5L | TAF5L | Act1 | actin | TAF13 | MED13 | |||||||||||||||||||||||||||
| TAF6 | TAF6 | TAF6 | Eaf1 | P400 | TAF14 | MED14 | |||||||||||||||||||||||||||
| TAF6L | TAF6L | TAF6L | Eaf5 | TBP | MED15 | ||||||||||||||||||||||||||||
| TAF9 | TAF9 | TAF9 | TAF9 | TAF9 | Sas4 | MED16 | |||||||||||||||||||||||||||
| TAF10 | TAF10 | TAF10 | TAF10 | TAF10 | YL1 | MED17 | |||||||||||||||||||||||||||
| TAF12 | TAF12 | TAF12 | TAF12 | TAF12 | TIP49a | MED18 | |||||||||||||||||||||||||||
| Sgf29 | TIP49b | MED19 | |||||||||||||||||||||||||||||||
| Sgf73 | SCA7 | SCA7 | TRCp120 | MED20 | |||||||||||||||||||||||||||||
| Ubp8 | Ubp8 | MLE | MED21 | ||||||||||||||||||||||||||||||
| Sin4 | MSL1 | MED22 | |||||||||||||||||||||||||||||||
| Rtg2 | MSL2 | CDK8 | |||||||||||||||||||||||||||||||
| Ahc1 | Jil1 | CycC | |||||||||||||||||||||||||||||||
| STAF36 | roX RNA | ||||||||||||||||||||||||||||||||
| STAF46 | |||||||||||||||||||||||||||||||||
| STAF55 | |||||||||||||||||||||||||||||||||
| STAF60 | |||||||||||||||||||||||||||||||||
| SAP130 | SAP130 | ||||||||||||||||||||||||||||||||
| Ref.h | (31, 33) | (124, 125) | (31, 33) | (126) | (127, 128) | (129) | (30) | (37) | (43) | (45, 46, 130, 131) | (132–134) | (101) | (135, 136) | (137, 138) | (139) | (107) | (140) | (110) | |||||||||||||||
| complex . | Gcn5 family . | . | . | . | . | . | . | MYST family . | . | . | . | . | Others . | . | . | . | . | . | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| . | SAGA . | SLIK . | ADA . | HAT-A2 . | STAGA . | TFTC . | PCAF . | NuA4 . | NuA3 . | SAS-I . | TIP60 . | MSLd . | Hat1/2 . | Elongator . | TFIIDe . | TFIIICf . | ACTR/SRC-1 . | Mediator . | |||||||||||||||
| organism . | yeast . | yeast . | yeast . | yeast . | mammal . | mammal . | mammal . | yeast . | yeast . | yeast . | mammal . | fly . | yeast . | yeast . | mammal/fly/yeast . | mammal . | mammal . | yeast . | |||||||||||||||
| HAT vs. free histonea . | . | . | . | . | . | . | . | H2A . | H2A . | N.D.c . | H2A . | H2A . | H2A . | H2A . | . | . | . | N.A.c . | |||||||||||||||
| . | . | . | . | . | . | . | . | . | . | . | . | . | . | H2B . | . | . | . | . | |||||||||||||||
| . | H3 . | H3 . | H3 . | H3 . | H3 . | H3 . | H3 . | H3 . | H3 . | . | H3 . | H3 . | . | H3 . | H3 . | H3g . | H3 . | . | |||||||||||||||
. | H4 . | H4 . | H4 . | H4 . | H4 . | H4 . | H4 . | H4 . | H4 . | . | H4 . | H4 . | H4 . | H4 . | H4 . | . | H4 . | . | |||||||||||||||
| HAT complex vs. nucleosomal histonea . | H2B . | H2B . | H2B . | . | H2B . | . | . | H2A . | . | . | H2A . | . | N.D.c . | . | N.D.c . | H2A . | N.A.c . | . | |||||||||||||||
| . | H3 . | H3 . | H3 . | H3 . | H3 . | H3 . | H3 . | . | H3 . | . | . | . | . | H3 . | . | H3 . | . | H3 . | |||||||||||||||
| . | . | . | . | . | . | . | H4 . | H4 . | . | H4 . | H4 . | H4 . | . | H4 . | . | H4 . | . | H4 . | |||||||||||||||
| catalytic subunit(s) . | Gcn5 . | Gcn5 . | Gcn5 . | Gcn5 . | Gcn5L . | Gcn5L . | PCAF . | Esa1 . | Sas3 . | Sas2 . | Tip60 . | MOF . | Hat1 . | Elp3 . | TAFII250 (TAF1) . | TFIIIC220 TFIIIC110 TFIIIC90 . | ACTR/SRC-1 . | MED5 (Nut1) . | |||||||||||||||
| other . | Tral . | Tral . | . | . | TRRAP . | TRRAP . | PAF400 . | Yaf9 . | TAF14 . | Sas5 . | GAS41 . | . | Hat2 . | Elp1 . | TAF2 . | TFIIIC102 . | CBP . | MED1 . | |||||||||||||||
| subunitsb . | Ada1 . | Ada1 . | . | . | STAF42 . | . | . | Yng2 . | Yng1 . | . | ING3 . | . | Hif1 . | Elp2 . | TAF3 . | TFIIIC63 . | SRC-2 . | MED2 . | |||||||||||||||
| Ada2 | Ada2 | Ada2 | Ada2 | Ada2 | Arp4 | BAF53 | Elp4 | TAF4 | SS-A | MED3 | |||||||||||||||||||||||
| Ada3 | Ada3 | Ada3 | Ada3 | STAF54 | Ada3 | Ada3 | Epl1 | EPC | Elp5 | TAF5 | IKKβ | MED4 | |||||||||||||||||||||
| Ada5 | Ada5 | EPC-like | Elp6 | TAF6 | IKKγ | MED6 | |||||||||||||||||||||||||||
| Spt3 | Spt3 | Spt3 | Spt3 | Spt3 | Eaf3 | MRG15 | MSL3 | TAF7 | MED7 | ||||||||||||||||||||||||
| Spt7 | Spt7 | STAF65γ | MRGX | TAF8 | MED8 | ||||||||||||||||||||||||||||
| Spt8 | Eaf7 | MRGBP | TAF9 | MED9 | |||||||||||||||||||||||||||||
| TAF2 | Eaf6 | FLJ11730 | TAF10 | MED10 | |||||||||||||||||||||||||||||
| TAF4 | Eaf2 | DMAP1 | TAF11 | MED11 | |||||||||||||||||||||||||||||
| TAF5 | TAF5 | TAF5 | Tra1 | TRRAP | TAF12 | MED12 | |||||||||||||||||||||||||||
| TAF5L | TAF5L | TAF5L | Act1 | actin | TAF13 | MED13 | |||||||||||||||||||||||||||
| TAF6 | TAF6 | TAF6 | Eaf1 | P400 | TAF14 | MED14 | |||||||||||||||||||||||||||
| TAF6L | TAF6L | TAF6L | Eaf5 | TBP | MED15 | ||||||||||||||||||||||||||||
| TAF9 | TAF9 | TAF9 | TAF9 | TAF9 | Sas4 | MED16 | |||||||||||||||||||||||||||
| TAF10 | TAF10 | TAF10 | TAF10 | TAF10 | YL1 | MED17 | |||||||||||||||||||||||||||
| TAF12 | TAF12 | TAF12 | TAF12 | TAF12 | TIP49a | MED18 | |||||||||||||||||||||||||||
| Sgf29 | TIP49b | MED19 | |||||||||||||||||||||||||||||||
| Sgf73 | SCA7 | SCA7 | TRCp120 | MED20 | |||||||||||||||||||||||||||||
| Ubp8 | Ubp8 | MLE | MED21 | ||||||||||||||||||||||||||||||
| Sin4 | MSL1 | MED22 | |||||||||||||||||||||||||||||||
| Rtg2 | MSL2 | CDK8 | |||||||||||||||||||||||||||||||
| Ahc1 | Jil1 | CycC | |||||||||||||||||||||||||||||||
| STAF36 | roX RNA | ||||||||||||||||||||||||||||||||
| STAF46 | |||||||||||||||||||||||||||||||||
| STAF55 | |||||||||||||||||||||||||||||||||
| STAF60 | |||||||||||||||||||||||||||||||||
| SAP130 | SAP130 | ||||||||||||||||||||||||||||||||
| Ref.h | (31, 33) | (124, 125) | (31, 33) | (126) | (127, 128) | (129) | (30) | (37) | (43) | (45, 46, 130, 131) | (132–134) | (101) | (135, 136) | (137, 138) | (139) | (107) | (140) | (110) | |||||||||||||||
“HAT” indicates that a monomer of the catalytic subunit was examined as an enzyme, while “HAT complex” indicates that the complex forms were used. For the substrates, “free histones” and “nucleosomal histones” indicate that the corresponding forms of histones were used. The references for the specificity of the catalytic subunits alone against free histones are shown in Table 1.
Subunits conserved among complexes of the Gcn5 family and MYST family are represented on horizontal lines as in (32). Unified nomenclature for TFIID and mediator subunits is used (141, 142).
N.D. = not detected, N.A. = not analyzed.
A catalytic domain of MOF (aa. 518–827) acetylates H2A/H3/H4 (101), whereas the full-length MOF shows a strong preference for histone H4 (100). An N-terminal domain is involved in this gain of specificity (100). The full-length MOF is also capable of acetylating of nucleosomal histones with a strong preference for lysine 16 of histone H4 (100).
HAT activity toward nucleosomal histones of a TAFII250-containing complex (TFIIDβ) has not been detected (129). Subunit composition in yeast is shown.
Recombinant TFIIIC90 alone possesses a HAT activity toward nucleosomal histones with a strong preference for histone H3 (108).
Specificity of TFIIIC90. Specificity of TFIIIC220 or TFIIIC110 alone toward free histones has yet to be analyzed.
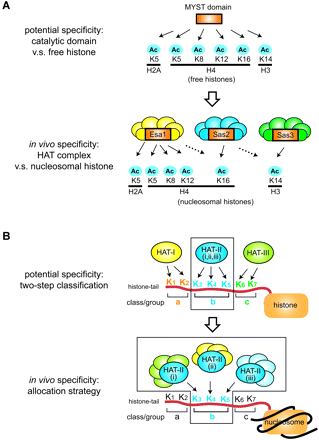
The lysine specificity of HATs is also modulated through multisubunit complex formation. In some cases, the lysine specificity of HAT complexes toward nucleosomal histones is more restricted than that of the catalytic subunit alone toward free histones. This is the case for HATs in the MYST family. Interestingly, there appears to be a systematic pattern underlying the in vivo lysine specificity of MYST family HATs, which we now call “allocation” (Fig. 3A lower). Six lysine residues in histone N-tails (K5 of H2A, K14 of H3, and K5/8/12/16 of H4) are potential targets for acetylation by MYST family HATs (Table 4, Fig. 3A upper) (29, 36). The MYST family has three members (Esa1, Sas2, and Sas3) in Saccharomyces cerevisiae. Esa1 has major effects on H2A-K5 and H4-K5/8/12 acetylation, both in vivo and in the form of multisubunit complexes (37–42), whereas Sas3 affects H3-K14 (43, 44) and Sas2 affects H4-K16 (Table 4 and Fig. 3A, lower) (41, 42, 45, 46). Based on these observations, we propose that there is an “allocation” strategy, such that HATs cover all potential acetylation sites while narrowing down the specificity of each member with little overlap (Fig. 3B, lower). Potential acetylation sites are defined by local structures of catalytic domains of HATs and histone N-tails, which may follow the “two-step classification” hypothesis (Fig. 3B, upper). This strategy does not decrease the number of lysines that are acetylated by all family members, while narrowing the specificity of an individual family member. This may explain why HATs form a family. Since the number of the MYST family members is increased in mammals (Table 1), it is intriguing to examine whether further allocation is observed and how it relates to complexity of gene regulation in mammals.
Lysine specificity of HAT complexes and the “allocation” hypothesis. (A) Allocation of lysine specificities among members of the MYST-HAT family. (Upper) Potential lysine specificity of the MYST-HAT domain toward free histones as substrates, based on experimental results found for Esa1, Sas3, and human Tip60 (Table 4). Positions of lysines are numbered according to the primary structures of vertebrate histones as in Fig. 2. (Lower) Lysine-specific acetylation of MYST-HAT family members in vivo (Table 4). The six lysines potentially acetylated by the MYST domain are allocated by the MYST-HAT family members for acetylation in vivo. H4-K16 may not be a preferred Esa1 acetylation site compared to other histone H4 lysines (dotted arrow; Table 4). (B) Two-step classification hypothesis and allocation strategy to select specific lysines by HAT. (Upper) The potential specificity of each enzyme is defined by catalytic domain structures, which correspond to the two-step classification of lysines (Fig. 2B). (Lower) Lysine specificity is allocated by family members, and each member regulates specific lysine(s) in vivo.
Lysine specificity of MYST-HATs in Saccharomyces cerevisiae.a
| . | In vitrob . | . | . | . | In vivoc . | . | Ref. . | ||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| . | MYST vs. free histone . | MYST vs. nuc. histone . | MYSTc vs. free histone . | MYSTc vs. nuc. histone . | Western blot . | Chromatin IP . | . | ||||
| Sas2 | not detected | not detected | (41, 42, 45, 46) | ||||||||
| H3-K14 | |||||||||||
| H4-K16 | H4 | H4-K16 | H4-K16 | ||||||||
| Sas3 | H2A (c) | not detected | (43, 44, 103) | ||||||||
| H3 | H3 | H3 | H3-K9/14 | ||||||||
| H4 | H4 | ||||||||||
| Esa1 | H2Ad-K5(c) | not detected | H2A (h/y) | H2Ad-K5(c) | H2A-K7(y) | (36–42, 143) | |||||
| H3-K14 | H3 | H2B-K16(y) | |||||||||
| H4-K5, K8, K12, K16 | H4 | H4-K5, K8, K12e | H4-K5, K8, K12 | H4-K5, K8, K12 | |||||||
| . | In vitrob . | . | . | . | In vivoc . | . | Ref. . | ||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| . | MYST vs. free histone . | MYST vs. nuc. histone . | MYSTc vs. free histone . | MYSTc vs. nuc. histone . | Western blot . | Chromatin IP . | . | ||||
| Sas2 | not detected | not detected | (41, 42, 45, 46) | ||||||||
| H3-K14 | |||||||||||
| H4-K16 | H4 | H4-K16 | H4-K16 | ||||||||
| Sas3 | H2A (c) | not detected | (43, 44, 103) | ||||||||
| H3 | H3 | H3 | H3-K9/14 | ||||||||
| H4 | H4 | ||||||||||
| Esa1 | H2Ad-K5(c) | not detected | H2A (h/y) | H2Ad-K5(c) | H2A-K7(y) | (36–42, 143) | |||||
| H3-K14 | H3 | H2B-K16(y) | |||||||||
| H4-K5, K8, K12, K16 | H4 | H4-K5, K8, K12e | H4-K5, K8, K12 | H4-K5, K8, K12 | |||||||
Experimental data shown are from the references indicated. Because the primary structures of amino-terminal tails of histones H2A and H2B vary from species to species (19), the species from which the histones were derived are shown in parentheses for histones H2A and H2B (i.e., c: chicken, h: human, y: yeast).
“MYST” indicates that the monomer of the corresponding member (Sas2, Sas3, or Esa1) was examined as an enzyme in the in vitro assays, while “MYSTc” indicates that the complex form of the member was used. For the substrates, “free histones” and “nuc. histones” indicate that free histones and nucleosomal histones were used in the assay, respectively.
The sites whose acetylation is reduced in the strains with mutations in the corresponding member (Sas2, Sas3, or Esa1) are detected either by “Western blot” or “Chromatin IP” assay.
Lysine 9 of chicken H2A is weakly acetylated by Esa1 compared to lysine 5 of chicken H2A. Because the site is not reported to be acetylated in vivo, the lysine 9 is omitted from further discussion (see Refs. 28 and 29).
The level of acetylation of H4-K16 by the Esa1 complex is slightly above the background level in vitro (38).
Lysine specificity of MYST-HATs in Saccharomyces cerevisiae.a
| . | In vitrob . | . | . | . | In vivoc . | . | Ref. . | ||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| . | MYST vs. free histone . | MYST vs. nuc. histone . | MYSTc vs. free histone . | MYSTc vs. nuc. histone . | Western blot . | Chromatin IP . | . | ||||
| Sas2 | not detected | not detected | (41, 42, 45, 46) | ||||||||
| H3-K14 | |||||||||||
| H4-K16 | H4 | H4-K16 | H4-K16 | ||||||||
| Sas3 | H2A (c) | not detected | (43, 44, 103) | ||||||||
| H3 | H3 | H3 | H3-K9/14 | ||||||||
| H4 | H4 | ||||||||||
| Esa1 | H2Ad-K5(c) | not detected | H2A (h/y) | H2Ad-K5(c) | H2A-K7(y) | (36–42, 143) | |||||
| H3-K14 | H3 | H2B-K16(y) | |||||||||
| H4-K5, K8, K12, K16 | H4 | H4-K5, K8, K12e | H4-K5, K8, K12 | H4-K5, K8, K12 | |||||||
| . | In vitrob . | . | . | . | In vivoc . | . | Ref. . | ||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| . | MYST vs. free histone . | MYST vs. nuc. histone . | MYSTc vs. free histone . | MYSTc vs. nuc. histone . | Western blot . | Chromatin IP . | . | ||||
| Sas2 | not detected | not detected | (41, 42, 45, 46) | ||||||||
| H3-K14 | |||||||||||
| H4-K16 | H4 | H4-K16 | H4-K16 | ||||||||
| Sas3 | H2A (c) | not detected | (43, 44, 103) | ||||||||
| H3 | H3 | H3 | H3-K9/14 | ||||||||
| H4 | H4 | ||||||||||
| Esa1 | H2Ad-K5(c) | not detected | H2A (h/y) | H2Ad-K5(c) | H2A-K7(y) | (36–42, 143) | |||||
| H3-K14 | H3 | H2B-K16(y) | |||||||||
| H4-K5, K8, K12, K16 | H4 | H4-K5, K8, K12e | H4-K5, K8, K12 | H4-K5, K8, K12 | |||||||
Experimental data shown are from the references indicated. Because the primary structures of amino-terminal tails of histones H2A and H2B vary from species to species (19), the species from which the histones were derived are shown in parentheses for histones H2A and H2B (i.e., c: chicken, h: human, y: yeast).
“MYST” indicates that the monomer of the corresponding member (Sas2, Sas3, or Esa1) was examined as an enzyme in the in vitro assays, while “MYSTc” indicates that the complex form of the member was used. For the substrates, “free histones” and “nuc. histones” indicate that free histones and nucleosomal histones were used in the assay, respectively.
The sites whose acetylation is reduced in the strains with mutations in the corresponding member (Sas2, Sas3, or Esa1) are detected either by “Western blot” or “Chromatin IP” assay.
Lysine 9 of chicken H2A is weakly acetylated by Esa1 compared to lysine 5 of chicken H2A. Because the site is not reported to be acetylated in vivo, the lysine 9 is omitted from further discussion (see Refs. 28 and 29).
The level of acetylation of H4-K16 by the Esa1 complex is slightly above the background level in vitro (38).
The formation of multisubunit complexes is also reported to broaden the lysine specificity of HATs. For example, the Gcn5 complexes ADA and SAGA can acetylate histone H2B in vitro, whereas Gcn5 alone cannot (33). Mechanisms underlying this gain of lysine specificity are unknown, although the Ada2 and Ada3 subunits of the complex are necessary and sufficient for broadening the lysine specificity of Gcn5 (34). Tertiary structure analyses of these complexes may provide insight into the underlying mechanisms.
4. The histone code hypothesis
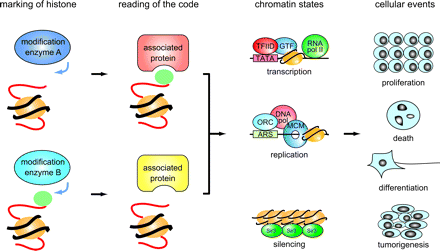
The residue specificity of identified HATs and HAT complexes was a strong evidence against the original proposal that the neutralization of the positive charge of lysine residues and a resultant decrease in electrostatic interactions between DNA and histones is the major consequence of histone acetylation. Instead, the idea that residue-specific modification of histones has unique and specific effects on chromatin function has become widely accepted (8). Post-translational modifications of histones, including acetylation, constitute a code that allows specific interactions or reactions with chromatin-associated components to take place in a chromosomal context. This idea has gained currency as the “histone code hypothesis”, which holds that the code is generated by histone-modifying enzymes of defined specificity and read by nonhistone proteins that bind in a modification-sensitive manner (Table 5 and Fig. 4) (5, 47). The mechanism is likely to be functioning in reactions other than transcription which are regulated by post-translational modification of histones, such as DNA replication, repair and recombination (Fig. 4) (48–50).
The histone code hypothesis. Schematic of the histone code hypothesis. Histones are labeled with “codes” by histone modification enzymes (“marking of histone” in the figure). These post-translational modifications are recognized by proteins that interact with histones in modification-dependent manners (“reading of the code”). Recruitment of these histone-interacting proteins triggers subsequent reactions on chromatin (“chromatin states”), which cause various changes (“cellular events”).
Examples of residue-specific histone modifications and induced downstream events.
| Histone . | Residue . | Modification . | Organism . | Modification enzyme . | Recognition . | Chromatin states (next reaction) . | Cellular events . | Ref. . |
|---|---|---|---|---|---|---|---|---|
| H4 | N.D.a | acetylation | mammal | CDY | BRDT | chromatin reorganization | spermatogenesis | (144, 145) |
| H3 | K4 | methylation | yeast | Set1 | Chd1 | acetylation of histone H3 by SLIK complex | response to transcriptional stress | (146–148) |
| K9 | methylation | yeast | Clr4 | Swi6 | silencing | maintenance of heterochromatin | (69, 70) | |
| K9 | methylation | mammal | Suv39h1 | HP1 | interfere with phosphorylation of H3-S10 | maintenance of heterochromatin | (68, 69, 149) | |
| K9 K27 | methylation | plant | Kryptonite (for K9) | CMT3 | DNA methylation | flowering | (150) | |
| Unknown (for K27) | ||||||||
| K27 | methylation | fly | E(Z) | Polycomb (PC) | silencing | homeotic gene repression | (151–153) | |
| K79 | methylation | mammal | DOT1L | 53BP1 | change higher-order chromatin structure | cell cycle | (154, 155) | |
| S10b | phosphorylation | tetrahymenayeast | Snf1 | GCN5 | acetylation of K14 of H3 by Gcn5 | transcription | (72, 156) | |
| H2A | S129 | phosphorylation | yeast | Mec1 | Arp4 | acetylation by NuA4 | DNA repair | (157, 158) |
| Histone . | Residue . | Modification . | Organism . | Modification enzyme . | Recognition . | Chromatin states (next reaction) . | Cellular events . | Ref. . |
|---|---|---|---|---|---|---|---|---|
| H4 | N.D.a | acetylation | mammal | CDY | BRDT | chromatin reorganization | spermatogenesis | (144, 145) |
| H3 | K4 | methylation | yeast | Set1 | Chd1 | acetylation of histone H3 by SLIK complex | response to transcriptional stress | (146–148) |
| K9 | methylation | yeast | Clr4 | Swi6 | silencing | maintenance of heterochromatin | (69, 70) | |
| K9 | methylation | mammal | Suv39h1 | HP1 | interfere with phosphorylation of H3-S10 | maintenance of heterochromatin | (68, 69, 149) | |
| K9 K27 | methylation | plant | Kryptonite (for K9) | CMT3 | DNA methylation | flowering | (150) | |
| Unknown (for K27) | ||||||||
| K27 | methylation | fly | E(Z) | Polycomb (PC) | silencing | homeotic gene repression | (151–153) | |
| K79 | methylation | mammal | DOT1L | 53BP1 | change higher-order chromatin structure | cell cycle | (154, 155) | |
| S10b | phosphorylation | tetrahymenayeast | Snf1 | GCN5 | acetylation of K14 of H3 by Gcn5 | transcription | (72, 156) | |
| H2A | S129 | phosphorylation | yeast | Mec1 | Arp4 | acetylation by NuA4 | DNA repair | (157, 158) |
Lysines acetylated by CDY are yet to be identified. Interaction between acetylated histones and BRDT was examined using a H4 amino-terminal peptide with acetylation at K5, K8, K12 and K16.
Snf1 and GCN5 derive from S. cerevisiae and Tetrahymena, respectively.
Examples of residue-specific histone modifications and induced downstream events.
| Histone . | Residue . | Modification . | Organism . | Modification enzyme . | Recognition . | Chromatin states (next reaction) . | Cellular events . | Ref. . |
|---|---|---|---|---|---|---|---|---|
| H4 | N.D.a | acetylation | mammal | CDY | BRDT | chromatin reorganization | spermatogenesis | (144, 145) |
| H3 | K4 | methylation | yeast | Set1 | Chd1 | acetylation of histone H3 by SLIK complex | response to transcriptional stress | (146–148) |
| K9 | methylation | yeast | Clr4 | Swi6 | silencing | maintenance of heterochromatin | (69, 70) | |
| K9 | methylation | mammal | Suv39h1 | HP1 | interfere with phosphorylation of H3-S10 | maintenance of heterochromatin | (68, 69, 149) | |
| K9 K27 | methylation | plant | Kryptonite (for K9) | CMT3 | DNA methylation | flowering | (150) | |
| Unknown (for K27) | ||||||||
| K27 | methylation | fly | E(Z) | Polycomb (PC) | silencing | homeotic gene repression | (151–153) | |
| K79 | methylation | mammal | DOT1L | 53BP1 | change higher-order chromatin structure | cell cycle | (154, 155) | |
| S10b | phosphorylation | tetrahymenayeast | Snf1 | GCN5 | acetylation of K14 of H3 by Gcn5 | transcription | (72, 156) | |
| H2A | S129 | phosphorylation | yeast | Mec1 | Arp4 | acetylation by NuA4 | DNA repair | (157, 158) |
| Histone . | Residue . | Modification . | Organism . | Modification enzyme . | Recognition . | Chromatin states (next reaction) . | Cellular events . | Ref. . |
|---|---|---|---|---|---|---|---|---|
| H4 | N.D.a | acetylation | mammal | CDY | BRDT | chromatin reorganization | spermatogenesis | (144, 145) |
| H3 | K4 | methylation | yeast | Set1 | Chd1 | acetylation of histone H3 by SLIK complex | response to transcriptional stress | (146–148) |
| K9 | methylation | yeast | Clr4 | Swi6 | silencing | maintenance of heterochromatin | (69, 70) | |
| K9 | methylation | mammal | Suv39h1 | HP1 | interfere with phosphorylation of H3-S10 | maintenance of heterochromatin | (68, 69, 149) | |
| K9 K27 | methylation | plant | Kryptonite (for K9) | CMT3 | DNA methylation | flowering | (150) | |
| Unknown (for K27) | ||||||||
| K27 | methylation | fly | E(Z) | Polycomb (PC) | silencing | homeotic gene repression | (151–153) | |
| K79 | methylation | mammal | DOT1L | 53BP1 | change higher-order chromatin structure | cell cycle | (154, 155) | |
| S10b | phosphorylation | tetrahymenayeast | Snf1 | GCN5 | acetylation of K14 of H3 by Gcn5 | transcription | (72, 156) | |
| H2A | S129 | phosphorylation | yeast | Mec1 | Arp4 | acetylation by NuA4 | DNA repair | (157, 158) |
Lysines acetylated by CDY are yet to be identified. Interaction between acetylated histones and BRDT was examined using a H4 amino-terminal peptide with acetylation at K5, K8, K12 and K16.
Snf1 and GCN5 derive from S. cerevisiae and Tetrahymena, respectively.
The histone codes are read by proteins that interact with histones in modification-dependent manners (Fig. 5A). One group, bromodomain proteins, is considered to bear acetylated histone interaction domains (51). To date, the bromodomains of various proteins have been reported to interact with acetylated histones in a lysine-specific manner in vitro (Table 6). Bromodomain-dependent binding of proteins to acetylated chromatin supports the binding of bromodomains to acetylated histone in vivo (52–56). These bromodomain-containing proteins are components of nucleosome-modulating complexes that also include ATPases and HATs. Acetylation at specific lysines is thought to stabilize these complexes through bromodomain interaction and to stimulate nucleosome remodeling, further acetylation, or the recruitment of TFIID (52, 57). Initial recruitment of a HAT to chromatin may require nucleosome remodeling. In vivo observations indicate that a HAT is recruited to a promoter after a nucleosome remodeling ATPase complex (58, 59). HAT is proposed to be required for the subsequent stable binding of the ATPase complex in these cases (60). HAT may be recruited to chromatin before the ATPase complex and recruit ATPases in some instances (52, 57, 60).
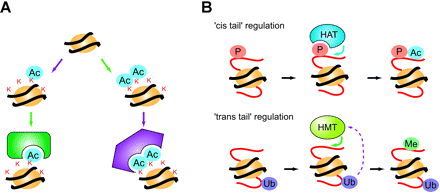
Reading histone codes. (A) Depending on specific patterns established by various histone-modification enzymes, distinct proteins are recruited to chromatin, with specific results (e.g. transcriptional activation/inactivation). (B) Schematic of “chromatin crosstalk.” The efficiency of modification at particular residues depends on pre-existing histone modification patterns. Such interdependency might involve residues in the same histone-tail (‘cis tail’ regulation) or those in different histone tails (‘trans tail’ regulation). Abbreviations: HAT, histone acetyltransferase; HMT, histone methyltransferase.
Summary of bromodomains with known acetyl-histone-binding ability.
| Protein . | Organism . | Acetyl-histone-binding . | Detection . | Ref. . |
|---|---|---|---|---|
| Gcn5 | human | K5 acetylated-H2A | NMR | (159) |
| K8 acetylated-H4 | ||||
| K16 acetylated-H4 | ||||
| yeast | K16 acetylated-H4 | NMR | (160) | |
| PCAF | human | acetylated-H3 | NMR | (51) |
| acetylated-H4 | ||||
| TAFII250 | human | K16 acetylated-H4 | NMR | (161) |
| K8/K16 acetylated-H4 | ||||
| K5/K12 acetylated-H4 | ||||
| K5/K8/K12/K16 acetylated-H4 | ||||
| K14 acetylated-H3 | Western blot | (56) | ||
| K8 acetylated-H4 | ||||
| K12 acetylated-H4 | ||||
| K16 acetylated-H4 | ||||
| Brd2 | mouse | K5 acetylated-H2B | Western blot | (56) |
| K12 acetylated-H2B | ||||
| K8 acetylated-H4 | ||||
| K12 acetylated-H4 | ||||
| K16 acetylated-H4 | ||||
| Brd4 | mouse | K14 acetylated-H3 | Western blot | (55) |
| K9/K14 acetylated-H3 | ||||
| K5/K12 acetylated-H4 | ||||
| K5/K8/K12/K16 acetylated-H4 | ||||
| BRDT | mouse | acetylated-H4 | Western blot | (145) |
| Bdf1 | yeast | acetylated-H3 acetylated-H4 | Coomassie brilliant blue staining | (53, 54) |
| Rsc4 | yeast | K14 acetylated-H3 | Western blot | (162) |
| Protein . | Organism . | Acetyl-histone-binding . | Detection . | Ref. . |
|---|---|---|---|---|
| Gcn5 | human | K5 acetylated-H2A | NMR | (159) |
| K8 acetylated-H4 | ||||
| K16 acetylated-H4 | ||||
| yeast | K16 acetylated-H4 | NMR | (160) | |
| PCAF | human | acetylated-H3 | NMR | (51) |
| acetylated-H4 | ||||
| TAFII250 | human | K16 acetylated-H4 | NMR | (161) |
| K8/K16 acetylated-H4 | ||||
| K5/K12 acetylated-H4 | ||||
| K5/K8/K12/K16 acetylated-H4 | ||||
| K14 acetylated-H3 | Western blot | (56) | ||
| K8 acetylated-H4 | ||||
| K12 acetylated-H4 | ||||
| K16 acetylated-H4 | ||||
| Brd2 | mouse | K5 acetylated-H2B | Western blot | (56) |
| K12 acetylated-H2B | ||||
| K8 acetylated-H4 | ||||
| K12 acetylated-H4 | ||||
| K16 acetylated-H4 | ||||
| Brd4 | mouse | K14 acetylated-H3 | Western blot | (55) |
| K9/K14 acetylated-H3 | ||||
| K5/K12 acetylated-H4 | ||||
| K5/K8/K12/K16 acetylated-H4 | ||||
| BRDT | mouse | acetylated-H4 | Western blot | (145) |
| Bdf1 | yeast | acetylated-H3 acetylated-H4 | Coomassie brilliant blue staining | (53, 54) |
| Rsc4 | yeast | K14 acetylated-H3 | Western blot | (162) |
Summary of bromodomains with known acetyl-histone-binding ability.
| Protein . | Organism . | Acetyl-histone-binding . | Detection . | Ref. . |
|---|---|---|---|---|
| Gcn5 | human | K5 acetylated-H2A | NMR | (159) |
| K8 acetylated-H4 | ||||
| K16 acetylated-H4 | ||||
| yeast | K16 acetylated-H4 | NMR | (160) | |
| PCAF | human | acetylated-H3 | NMR | (51) |
| acetylated-H4 | ||||
| TAFII250 | human | K16 acetylated-H4 | NMR | (161) |
| K8/K16 acetylated-H4 | ||||
| K5/K12 acetylated-H4 | ||||
| K5/K8/K12/K16 acetylated-H4 | ||||
| K14 acetylated-H3 | Western blot | (56) | ||
| K8 acetylated-H4 | ||||
| K12 acetylated-H4 | ||||
| K16 acetylated-H4 | ||||
| Brd2 | mouse | K5 acetylated-H2B | Western blot | (56) |
| K12 acetylated-H2B | ||||
| K8 acetylated-H4 | ||||
| K12 acetylated-H4 | ||||
| K16 acetylated-H4 | ||||
| Brd4 | mouse | K14 acetylated-H3 | Western blot | (55) |
| K9/K14 acetylated-H3 | ||||
| K5/K12 acetylated-H4 | ||||
| K5/K8/K12/K16 acetylated-H4 | ||||
| BRDT | mouse | acetylated-H4 | Western blot | (145) |
| Bdf1 | yeast | acetylated-H3 acetylated-H4 | Coomassie brilliant blue staining | (53, 54) |
| Rsc4 | yeast | K14 acetylated-H3 | Western blot | (162) |
| Protein . | Organism . | Acetyl-histone-binding . | Detection . | Ref. . |
|---|---|---|---|---|
| Gcn5 | human | K5 acetylated-H2A | NMR | (159) |
| K8 acetylated-H4 | ||||
| K16 acetylated-H4 | ||||
| yeast | K16 acetylated-H4 | NMR | (160) | |
| PCAF | human | acetylated-H3 | NMR | (51) |
| acetylated-H4 | ||||
| TAFII250 | human | K16 acetylated-H4 | NMR | (161) |
| K8/K16 acetylated-H4 | ||||
| K5/K12 acetylated-H4 | ||||
| K5/K8/K12/K16 acetylated-H4 | ||||
| K14 acetylated-H3 | Western blot | (56) | ||
| K8 acetylated-H4 | ||||
| K12 acetylated-H4 | ||||
| K16 acetylated-H4 | ||||
| Brd2 | mouse | K5 acetylated-H2B | Western blot | (56) |
| K12 acetylated-H2B | ||||
| K8 acetylated-H4 | ||||
| K12 acetylated-H4 | ||||
| K16 acetylated-H4 | ||||
| Brd4 | mouse | K14 acetylated-H3 | Western blot | (55) |
| K9/K14 acetylated-H3 | ||||
| K5/K12 acetylated-H4 | ||||
| K5/K8/K12/K16 acetylated-H4 | ||||
| BRDT | mouse | acetylated-H4 | Western blot | (145) |
| Bdf1 | yeast | acetylated-H3 acetylated-H4 | Coomassie brilliant blue staining | (53, 54) |
| Rsc4 | yeast | K14 acetylated-H3 | Western blot | (162) |
Sir3 (61, 62) and Tup1 (63) are proposed to interact with hypoacetylated histones, and both repress gene expression in S. cerevisiae. Sir3 spreads along chromatin and contributes to gene repression over a range of several kilobases (64). Deacetylation of H4-K16 by an HDAC, Sir2 (65), stimulates binding of Sir3 to chromatin and thus gene silencing, whereas acetylation of this lysine by a MYST-HAT, Sas2, prevents Sir3 from spreading on chromatin and contributes to anti-silencing (41, 42) (see next section). In contrast, Tup1 represses gene expression in a promoter-specific manner. The local recruitment of Tup1 is accomplished by sequence-specific DNA binding proteins such as α2/Mcm1, Mig1 and Sko1 (66). Tup1 acts in concert with a histone H2B/H3-specific HDAC, Hda1, to repress gene expression, possibly by binding to hypoacetylated histones (67).
Other modifications of histones also regulate their interaction with proteins, and these alterations also function as codes (Table 5). For example, methylation of H3-K9 is known to stimulate the binding of chromodomain-containing proteins such as HP1 and Swi6 to chromatin, leading to gene repression (68–70).
Modification of histones also influences other histone modifications, a phenomenon called cross talk (71) (Fig. 5B). In cis-tail crosstalk, a given modification affects modification of neighboring residues by physically stabilizing or inhibiting interaction between enzymes and substrates. For example, phosphorylation of serine 10 of histone H3 (H3-S10) enhances acetylation of H3-K14 by Gcn5 (72). In trans-tail crosstalk, a given modification affects modifications on other histone tails. For example, ubiquitination of H2B-K123 by Rad6/Ubc2 is required for methylation of H3-K4 and H3-K79 (73, 74). The interdependency of histone modifications led Fischle et al. to propose that histone modifications function as binary switches (75).
5. Long-range/chromosome-wide gene expression regulated by histone acetylation
Most HATs identified to date are involved mainly in promoter-specific gene expression. Analyses of these HATs have advanced our understanding of the molecular mechanisms by which histone acetylation controls gene expression at specific promoters. In contrast, the regulation of long-range/chromosome-wide gene expression by histone acetylation is poorly understood (Fig. 6). The MYST-type HAT family is a group of HATs involved mainly in long-range/chromosome-wide gene expression.
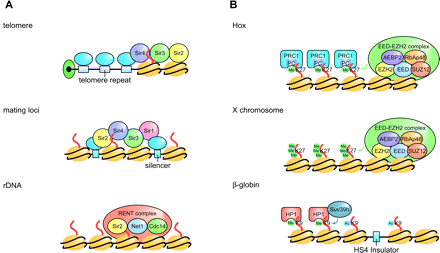
Long-range/chromosome-wide gene expression. (A) Examples of long-range gene regulation in yeast. At telomere-proximal regions (upper) and cryptic mating type loci (middle), Sir2 deacetylates H4-K16 and enhances the assembly of Sir2/Sir3/Sir4 proteins on chromatin in these regions. Sir1 is required for silencing at the mating type loci. The Sir proteins are recruited to the chromosomal regions through DNA-binding proteins that bind to telomere repeat or silencer elements. The RENT (Sir2, Net1, Cdc14) complex regulates silencing at the rDNA locus (lower). (B) Examples of long-range/chromosome-wide gene regulation in higher eukaryotes. At the HOX gene cluster (upper) and on the mammalian X chromosome (middle), H3-K27 is methylated by the EED-EZH2 complex (151, 165). At the HOX gene cluster, methylation enhances the binding of Polycomb (PC) in the PRC1 complex, which leads to silencing of the region. (Lower) Insulators at the β-globin locus are hyperacetylated, and H3-K9 in condensed chromatin region is methylated (80). A silencing protein, HP1, binds to histones methylated at H3-K9 by a histone methyltransferase, Suv39h.
Transcriptional silencing/anti-silencing of genes located near telomeres and the cryptic mating loci of yeast S. cerevisiae is a model system of long-range gene regulation in eukaryotes (64, 76–78). Deacetylation of H4-K16 by the Sir2 deacetylase is thought to enhance the binding of Sir3 to chromatin and to cause gene silencing at these regions. However, it was not clear whether deacetylation of this lysine residue was the primary determinant for Sir3 localization and gene silencing in vivo because the HAT acetylating this lysine in vivo was unknown. A MYST protein, Sas2, has been identified as the HAT that acetylates H4-K16 at telomere-distal regions (41, 42). The role of H4-K16 acetylation was evaluated by manipulating the acetylation level in vivo. Mutation of Sas2 and Sir2 caused the boundary between the hyper- and hypo-acetylated regions to advance and recede from its original position. Importantly, the position of this boundary coincided with that of Sir3 localization and of overall transcriptional activity. These results indicate that H4-K16 plays a decisive role in establishing transcriptionally active and inactive regions (Fig. 7A).
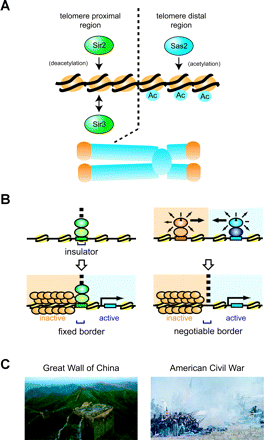
Negotiable and fixed borders. (A) Chromosomal gradient of histone acetylation near telomeres. Regions near the telomere end are hypoacetylated at H4-K16 through the function of Sir2, whereas telomere-distal regions are hyperacetylated through the function of Sas2. The hypoacetylated regions are enriched in Sir3 and gene expression is silenced. For details, see Ref. 41. (B) (Left) Establishment of fixed borders. An “insulator” DNA element is shown as a green box, and components recruited to this element are indicated as green circles. These factors function as a barrier, an enhancer blocker or both. (Right) Establishment of negotiable borders. The localization of each of a pair of modification enzymes can be defined by specific DNA elements on chromosomes. These enzymes are recruited by proteins bound to the DNA elements and can modify surrounding chromosomal regions. The border of the modified state is established somewhere between the two DNA elements. Distinct chromosome states are established according to specific modification states. Fixed borders (left) are tethered to an “insulator”, whereas negotiable borders (right) can shift according to the position of a “chromosomal gradient” based on the balance of two chromatin modification activities. For details, see (79). (C) Non-biological examples of a fixed border (left: Great Wall of China) and a negotiable border (right: American Civil War). The Great Wall of China was built by the first emperor of China as a first line of defense against the invading tribes north of China. The position of this border has been fixed for over 2,000 years. The American Civil War was fought in the United States in the 1860s between the northern states of the Union and the southern states of the Confederacy. The territories of each army changed during the war, but finally the Union prevailed and unified the states.
More importantly, these results inspired a novel view of chromosomal border establishment (79). A prevailing view of chromosome borders, which we term the “fixed border” model, is that they are ‘walls’ that actively inhibit the function of transcriptional enhancers or silencers between distinct regions on chromosomes (Fig. 7B; reviewed in Ref. 80). Positions of the borders will be defined by specific DNA elements and fixed in chromosomes (Fig. 7C). Analysis of Sas2, however, implied another mechanism that does not require ‘walls’ at chromosomal borders. Instead, it provided evidence for a mechanism (the “negotiable border” model) that defines a border through the balance of opposing enzymatic activities (Fig. 7B) (79). Positions of the borders depend on the strength of each activity and are thus movable (Fig. 7C).
Deacetylation of H4-K16 by Sir2 at the telomere-proximal regions stabilizes binding of Sir3 and Sir4 on the chromosome and inactivates gene expression (76). In contrast, acetylation of H4-K16 by Sas2 at the telomere-distal regions stabilizes binding of a bromodomain containing protein Bdf1 on the chromosome (53). Bdf1 is a component of a chromatin remodeling SWR1 complex, which exchanges conventional histone H2A with a histone variant H2A.Z (Htz1) in nucleosomes (81–83). This recruitment of H2A.Z on chromatin antagonizes telomeric silencing (84). Methylation of H3-K4 by Set1 and H3-K79 by Dot1 at telomere-distal regions may also play a role in anti-silencing by preventing Sir proteins to associate with chromatin of these regions (85–87). Ubiquitination of H2B-K123 by Rad6/Ubc2 should regulate methylation of these lysines (73, 74).
To establish a negotiable border, histone (or DNA) modification enzymes should be recruited to specific regions on chromosomes. A straightforward scenario for enzyme recruitment is to assume specific DNA elements that interact directly or indirectly with these enzymes. We previously predicted that boundary DNA elements (or insulators) originally assumed to define fixed borders may establish negotiable borders instead (79). Consistent with this prediction, a boundary DNA element in vertebrates recruits HATs to prevent the spread of silenced chromatin (88), and DNA elements that recruit histone modification enzymes function as boundary DNA elements in yeast (89). A temporal analysis of silent chromatin assembly in mammalian cells detected the bidirectional spread of histone and DNA modifications nucleated at a specific chromosome region (90). Other explanations of the recruitment of histone/DNA modification enzymes involve repetitive DNA elements and noncoding RNAs (91). Such mechanisms are important in dosage compensation in mammals and Drosophila and in gene silencing mediated by RNA interference (RNAi) (92). Establishment of eukaryotic chromosomal boundaries to regulate long-range/chromosome-wide gene expression involves many mechanisms (93, 94). How these mechanisms act in concert to regulate long-range/chromosome-wide transcriptional activity is one of the major challenges in the field of eukaryotic gene expression.
6. Perspectives
Here, we have reviewed major advances in histone acetylation research with emphases on lysine specificity in the acetylation reaction and on the regulation of long-range/chromosome-wide gene expression. Post-translational modification of histones also plays critical roles in other reactions involving chromatin, such as DNA replication, repair and recombination (48–50). Research in the past ten years has greatly advanced our knowledge not only of histone acetylation but also of acetylation of non-histone proteins, histone modification in general and epigenetic regulation in eukaryotes. Considering its highly packed structure and dynamic modulation throughout the cell cycle, and the tight control over efficient and specific gene expression, the eukaryotic chromosome is still full of mystery. Understanding the post-translational modifications of histones is a key for unlocking its secrets.
Present address: Graduate School of Science and Technology, Keio University, 3-14-1 Hiyoshi, Kohoku-ku, Yokohama 223-8522.
We are grateful to H. Fukuda and N. Sano for helpful discussions. We thank Drs. Wendy Zhow (http://www.meet-greatwall.org) and Tim Harrison (http://www.swcivilwar.com) for permission to use the images presented in Figure 7C. During this study, A. K. was a Research Fellow of the Japan Society for the Promotion of Science. This work was supported in part by Grants-in-Aid for Scientific Research from the Ministry of Education, Culture, Sports, Science and Technology of Japan and by Exploratory Research for Advanced Technology (ERATO) of the Japan Science and Technology Agency (JST).
REFERENCES
Kornberg, R.D. (
Kornberg, R.D. and Lorch, Y. (
Luger, K., Mader, A.W., Richmond, R.K., Sargent, D.F., and Richmond, T.J. (
Strahl, B.D. and Allis, C.D. (
Allfrey, V.G., Faulkner, R., and Mirsky, A.E. (
Hebbes, T.R., Thorne, A.W., and Crane-Robinson, C. (
Turner, B.M., Birley, A.J., and Lavender, J. (
Brownell, J. and Allis, C.D. (
Brownell, J., Zhou, J., Ranalli, T., Kobayashi, R., Edmondson, D., Roth, S., and Allis, C.D. (
Kleff, S., Andrulis, E.D., Anderson, C.W., and Sternglanz, R. (
Dyda, F., Klein, D.C., and Hickman, A.B. (
Dutnall, R.N., Tafrov, S.T., Sternglanz, R., and Ramakrishnan, V. (
Yang, X.-J., Ogryzko, V.V., Nishikawa, J., Howard, B.H., and Nakatani, Y. (
Ogryzko, V.V., Schiltz, R.L., Russanova, V., Howard, B.H., and Nakatani, Y. (
Bannister, A.J. and Kouzarides, T. (
Mizzen, C.A., Yang, X.J., Kokubo, T., Brownell, J.E., Bannister, A.J., Owen-Hughes, T., Workman, J., Wang, L., Berger, S.L., Kouzarides, T., Nakatani, Y., and Allis, C.D. (
Sterner, D.E. and Berger S.L. (
Roth, S.Y., Denu, J.M., and Allis, C.D. (
Yamamoto, T. and Horikoshi, M. (
Lu, L., Berkey, K.A., and Casero, Jr., R.A. (
Adachi, N., Kimura, A., and Horikoshi, M. (
Reifsnyder, C., Lowell, J., Clarke, A., and Pillus, L. (
Ehrenhofer-Murray, A.E., Rivier, D.H., and Rine, J. (
Hilfiker, A., Hilfiker-Kleiner, D., Pannuti, A., and Lucchesi, J.C. (
Marvin, K.W., Yau, P., and Bradbury, E.M. (
Thorne, A.W., Kmiciek, D., Mitchelsom, K., Sautiere, P., and Crane-Robinson, C. (
Kimura, A. and Horikoshi, M. (
Kimura, A. and Horikoshi, M. (
Ogryzko, V.V., Kotani, T., Zhang, X., Schiltz, R.L., Howard, T., Yang, X.J., Howard, B.H., Qin, J., and Nakatani, Y. (
Grant, P.A., Schieltz, D., Pray-Grant, M.G., Steger, D.J., Reese, J.C., Yates, J.R. 3rd, and Workman, J.L. (
Carrozza, M.J., Utley, R.T., Workman, J.L., and Côté, J. (
Grant, P.A., Duggan, L., Côté, J., Roberts, S.M., Brownell, J.E., Candau, R., Ohba, R., Owen-Hughes, T., Allis, C.D., Winston, F., Berger, S.L., and Workman, J.L. (
Balasubramanian, R., Pray-Grant, M.G., Selleck, W., Grant, P.A., and Tan, S. (
Boudreault, A.A., Cronier, D., Selleck, W., Lacoste, N., Utley, R.T., Allard, S., Savard, J., Lane, W.S., Tan, S., and Côté, J. (
Smith, E.R., Eisen, A., Gu, W., Sattah, M., Pannuti, A., Zhou, J., Cook, R.G., Lucchesi, J.C., and Allis, C.D. (
Allard, S., Utley, R.T., Savard, J., Clarke, A., Grant, P., Brandl, C.J., Pillus, L., Workman, J.L., and Côté, J. (
Ohba, R., Steger, D.J., Brownell, J.E., Mizzen, C.A., Cook, R.G., Côté, J., Workman, J.L., and Allis, C.D. (
Vogelauer, M., Wu, J., Suka, N., and Grunstein, M. (
Suka, N., Suka, Y., Carmen, A.A., Wu, J., and Grunstein, M. (
Kimura, A., Umehara, T., and Horikoshi, M. (
Suka, N., Luo, K., and Grunstein, M. (
John, S., Howe, L., Tafrov, S.T., Grant, P.A., Sternglanz, R., and Workman, J.L. (
Howe, L., Auston, D., Grant, P., John, S., Cook, R.G., Workman, J.L., and Pillus, L. (
Sutton, A., Shia, W.J., Band, D., Kaufman, P.D., Osada, S., Workman, J.L., and Sternglanz, R. (
Shia, W.J., Osada, S., Florens, L., Swanson, S.K., Washburn, M.P., and Workman, J.L. (
Donaldson, A.D. (
Peterson, C.L. and Côté, J. (
Thiriet, C. and Hayes, J.J. (
Dhalluin, C., Carlson, J.E., Zeng, L., He, C., Aggarwal, A.K., and Zhou, M.M. (
Hassan, A.H., Prochasson, P., Neely, K.E., Galasinski, S.C., Chandy, M., Carrozza, M.J., and Workman, J.L. (
Ladurner, A.G., Inouye, C., Jain, R., and Tjian, R. (
Matangkasombut, O. and Buratowski, S. (
Dey, A., Chitsaz, F., Abbasi, A., Misteli, T., and Ozato, K. (
Kanno, T., Kanno, Y., Siegel, R.M., Jang, M.K., Lenardo, M.J., and Ozato, K. (
Agalioti, T., Chen, G., and Thanos, D. (
Cosma, M.P., Tanaka, T., and Nasmyth, K. (
Krebs, J.E., Kuo, M.H., Allis, C.D., and Peterson, C.L. (
Hassan, A.H., Neely, K.E., and Workman, J.L. (
Hecht, A., Laroche, T., Strahl-Bolsinger, S., Gasser, S.M., and Grunstein, M. (
Carmen, A.A., Milne, L., and Grunstein, M. (
Edmondson, D.G., Smith, M.M., and Roth, S.Y. (
Grunstein, M. (
Imai, S., Armstrong, C.M., Kaeberlein, M., and Guarente, L. (
Smith, R.L. and Johnson, A.D. (
Wu, J., Suka, N., Carlson, M., and Grunstein, M. (
Lachner, M., O'Carroll, D., Rea, S., Mechtler, K., and Jenuwein, T. (
Bannister, A.J., Zegerman, P., Partridge, J.F., Miska, E.A., Thomas, J.O., Allshire, R.C., and Kouzarides, T. (
Nakayama, J., Rice, J.C., Strahl, B.D., Allis, C.D., and Grewal, S.I. (
Fischle, W., Wang, Y., and Allis, C.D. (
Clements, A., Poux, A.N., Lo, W.S., Pillus, L., Berger, S.L., and Marmorstein, R. (
Sun, Z.W. and Allis, C.D. (
Briggs, S.D., Xiao, T., Sun, Z.W., Caldwell, J.A., Shabanowitz, J., Hunt, D.F., Allis, C.D., and Strahl, B.D. (
Fischle, W., Wang, Y., and Allis, C.D. (
Kurdistani, S.K. and Grunstein, M. (
Rusche, L.N., Kirchmaier, A.L., and Rine, J. (
Pirrotta, V. and Gross, D.S. (
Kimura, A. and Horikoshi, M. (
West, A.G., Gaszner, M., and Felsenfeld, G. (
Mizuguchi, G., Shen, X., Landry, J., Wu, W.H., Sen, S., and Wu, C. (
Krogan, N.J., Keogh, M.C., Datta, N., Sawa, C., Ryan, O.W., Ding, H., Haw, R.A., Pootoolal, J., Tong, A., Canadien, V., Richards, D.P., Wu, X., Emili, A., Hughes, T.R., Buratowski, S., and Greenblatt, J.F. (
Kobor, M.S., Venkatasubrahmanyam, S., Meneghini, M.D., Gin, J.W., Jennings, J.L., Link, A.J., Madhani, H.D., and Rine, J. (
Meneghini, M.D., Wu, M., and Madhani, H.D. (
van Leeuwen, F., Gafken, P.R., and Gottschling, D.E. (
Ng, H.H., Feng, Q., Wang, H., Erdjument-Bromage, H., Tempst, P., Zhang, Y., and Struhl, K. (
Santos-Rosa, H., Bannister, A.J., Dehe, P.M., Geli, V., and Kouzarides, T. (
West, A.G., Huang, S., Gaszner, M., Litt, M.D., and Felsenfeld, G. (
Oki, M., Valenzuela, L., Chiba, T., Ito, T., and Kamakaka, R.T. (
Su, R.C., Brown, K.E., Saaber, S., Fisher, A.G., Merkenschlager, M., and Smale, S.T. (
Grewal, S.I. and Moazed, D. (
Verdel, A., Jia, S., Gerber, S., Sugiyama, T., Gygi, S., Grewal, S.I., and Moazed, D. (
Henikoff, S., Furuyama, T., and Ahmad, K. (
Swaminathan, J., Baxter, E.M., and Corces, V.G. (
Smith, E.R., Belote, J.M., Schiltz, R.L., Yang, X.J., Moore, P.A., Berger, S.L., Nakatani, Y., and Allis, C.D. (
Xu, W., Edmondson, D.G., and Roth, S.Y. (
Iizuka, M. and Stillman, B. (
Champagne, N., Bertos, N.R., Pelletier, N., Wang, A.H., Vezmar, M., Yang, Y., Heng, H.H., and Yang, X.J. (
Champagne, N., Pelletier, N., and Yang, X.J. (
Akhtar, A. and Becker, P.B. (
Smith, E.R., Pannuti, A., Gu, W., Steurnagel, A., Cook, R.G., Allis, C.D., and Lucchesi, J.C. (
Neal, K.C., Pannuti, A., Smith, E.R., and Lucchesi, J.C. (
Takechi, S. and Nakayama, T. (
Chen, H., Lin, R.J., Schiltz, R.L., Chakravarti, D., Nash, A., Nagy, L., Privalsky, M.L., Nakatani, Y., and Evans, R.M. (
Spencer, T.E., Jenster, G., Burcin, M.M., Allis, C.D., Zhou, J., Mizzen, C.A., McKenna, N.J., Onate, S.A., Tsai, S.Y., Tsai, M.J., and O'Malley, B.W. (
Wittschieben, B.O., Otero, G., de Bizemont, T., Fellows, J., Erdjument-Bromage, H., Ohba, R., Li, Y., Allis, C.D., Tempst, P., and Svejstrup, J.Q. (
Kundu, T.K., Wang, Z., and Roeder, R.G. (
Hsieh, Y.J., Kundu, T.K., Wang, Z., Kovelman, R., and Roeder, R.G. (
Angus-Hill, M.L., Dutnall, R.N., Tafrov, S.T., Sternglanz, R., and Ramakrishnan, V. (
Lorch, Y., Beve, J., Gustafsson, C.M., Myers, L.C., and Kornberg, R.D. (
Kawasaki, H., Schiltz, L., Chiu, R., Itakura, K., Taira, K., Nakatani, Y., and Yokoyama, K.K. (
Scott, E.K., Lee, T., and Luo, L. (
Grienenberger, A., Miotto, B., Sagnier, T., Cavalli, G., Schramke, V., Geli, V., Mariol, M., Berenger, H., Graba, Y., and Pradel, J. (
Suzuki, T., Kimura, A., Nagai, R., Horikoshi, M. (
Miyamoto, S., Suzuki, T., Muto, S., Aizawa, K., Kimura, A., Mizuno, Y., Nagino, T., Imai, Y., Adachi, N., Horikoshi, M., and Nagai, R. (
Daitoku, H., Hatta, M., Matsuzaki, H., Aratani, S., Ohshima, T., Miyagishi, M., Nakajima, T., and Fukamizu, A. (
Ma, K., Chan, J.K., Zhu, G., and Wu, Z. (
Thevenet, L., Mejean, C., Moniot, B., Bonneaud, N., Galeotti, N., Aldrian-Herrada, G., Poulat, F., Berta, P., Benkirane, M., and Boizet-Bonhoure, B. (
Yanazume, T., Hasegawa, K., Morimoto, T., Kawamura, T., Wada, H., Matsumori, A., Kawase, Y., Hirai, M., and Kita, T. (
Kawamura, T., Ono, K., Morimoto, T., Wada, H., Hirai, M., Hidaka, K., Morisaki, T., Heike, T., Nakahata, T., Kita, T., and Hasegawa, K. (
Rausa, F.M., 3rd, Hughes, D.E., and Costa, R.H. (
Wang, R., Cherukuri, P., and Luo, J. (
Yang, X.J. (
Pray-Grant, M.G., Schieltz, D., McMahon, S.J., Wood, J.M., Kennedy, E.L., Cook, R.G., Workman, J.L., Yates, J.R. 3rd, and Grant, P.A. (
Daniel, J.A., Torok, M.S., Sun, Z.W., Schieltz, D., Allis, C.D., Yates, J.R., 3rd, and Grant, P.A. (
Sendra, R., Tse, C., and Hansen, J.C. (
Martinez, E., Palhan, V.B., Tjernberg, A., Lymar, E.S., Gamper, A.M., Kundu, T.K., Chait, B.T., and Roeder, R.G. (
Helmlinger, D., Hardy. S., Sasorith, S., Klein, F., Robert, F., Weber, C., Miguet, L., Potier, N., Van-Dorsselaer, A., Wurtz, J.M., Mandel, J.L., Tora, L., and Devys, D. (
Brand, M., Yamamoto, K., Staub, A., and Tora, L. (
Osada, S., Sutton, A., Muster, N., Brown, C.E., Yates, J.R. 3rd, Sternglanz, R., and Workman, J.L. (
Meijsing, S.H. and Ehrenhofer-Murray, A.E. (
Ikura, T., Ogryzko, V.V., Grigoriev, M., Groisman, R., Wang, J., Horikoshi, M., Scully, R., Qin, J., and Nakatani, Y. (
Cai, Y., Jin, J., Tomomori-Sato, C., Sato, S., Sorokina, I., Parmely, T.J., Conaway, R.C., and Conaway, J.W. (
Cai, Y., Jin, J., Florens, L., Swanson, S.K., Kusch, T., Li, B., Workman, J.L., Washburn, M.P., Conaway, R.C., and Conaway, J.W. (
Parthun, M.R., Widom, J., and Gottschling, D.E. (
Poveda, A., Pamblanco, M., Tafrov, S., Tordera, V., Sternglanz, R., and Sendra, R. (
Winkler, G.S., Petrakis, T.G., Ethelberg, S., Tokunaga, M., Erdjument-Bromage, H., Tempst, P., and Svejstrup, J.Q. (
Winkler, G.S., Kristjuhan, A., Erdjument-Bromage, H., Tempst, P., and Svejstrup, J.Q. (
Burley, S.K. and Roeder, R.G. (
Wu, R.C., Qin, J., Hashimoto, Y., Wong, J., Xu, J., Tsai, S.Y., Tsai, M.J., and O'Malley, B.W. (
Tora, L. (
Bourbon, H.M., Aguilera, A., Ansari, A.Z., Asturias, F.J., Berk, A.J., Bjorklund, S., Blackwell, T.K., Borggrefe, T., Carey, M., Carlson, M., Conaway, J.W., Conaway, R.C., Emmons, S.W., Fondell, J.D., Freedman, L.P., Fukasawa, T., Gustafsson, C.M., Han, M., He, X., Herman, P.K., Hinnebusch, A.G., Holmberg, S., Holstege, F.C., Jaehning, J.A., Kim, Y.J., Kuras, L., Leutz, A., Lis, J.T., Meisterernest, M., Naar, A.M., Nasmyth, K., Parvin, J.D., Ptashne, M., Reinberg, D., Ronne, H., Sadowski, I., Sakurai, H., Sipiczki, M., Sternberg, P.W., Stillman, D.J., Strich, R., Struhl, K., Svejstrup, J.Q., Tuck, S., Winston, F., Roeder, R.G, and Kornberg, R.D. (
Clarke, A.S., Lowell, J.E., Jacobson, S.J., and Pillus, L. (
Lahn, B.T., Tang, Z.L., Zhou, J., Barndt, R.J., Parvinen, M., Allis, C.D., and Page, D.C. (
Pivot-Pajot, C., Caron, C., Govin, J., Vion, A., Rousseaux, S., and Khochbin, S. (
Santos-Rosa, H., Schneider, R., Bannister, A.J., Sherriff, J., Bernstein, B.E., Emre, N.C., Schreiber, S.L., Mellor, J., and Kouzarides, T. (
Pray-Grant, M.G., Daniel, J.A., Schieltz, D., Yates, JR, 3rd, and Grant, P.A. (
Zhang, L., Schroeder, S., Fong, N., and Bentley, D.L. (
Rea, S., Eisenhaber, F., O'Carroll, D., Strahl, B.D., Sun, Z.W., Schmid, M., Opravil, S., Mechtler, K., Ponting, C.P., Allis, C.D., and Jenuwein, T. (
Lindroth, A.M., Shultis, D., Jasencakova, Z., Fuchs, J., Johnson, L., Schubert, D., Patnaik, D., Pradhan, S., Goodrich, J., Schubert, I., Jenuwein, T., Khorasanizadeh, S., and Jacobsen, S.E. (
Cao, R., Wang, L., Wang, H., Xia, L., Erdjument-Bromage, H., Tempst, P., Jones, R.S., and Zhang, Y. (
Czermin, B., Melfi, R., McCabe, D., Seitz, V., Imhof, A., and Pirrotta, V. (
Muller, J., Hart, C.M., Francis, N.J., Vargas, M.L., Sengupta, A., Wild, B., Miller, E.L., O'Connor, M.B., Kingston, R.E., and Simon, J.A. (
Huyen, Y., Zgheib, O., Ditullio, R.A., Jr., Gorgoulis, V.G., Zacharatos, P., Petty, T.J., Sheston, E.A., Mellert, H.S., Stavridi, E.S., and Halazonetis, T.D. (
Feng, Q., Wang, H., Ng, H.H., Erdjument-Bromage, H., Tempst, P., Struhl, K., and Zhang, Y. (
Lo, W.S., Duggan, L., Emre, N.C., Belotserkovskya, R., Lane, W.S., Shiekhattar, R., and Berger, S.L. (
Downs, J.A., Lowndes, N.F., and Jackson, S.P. (
Downs, J.A., Allard, S., Jobin-Robitaille, O., Javaheri, A., Auger, A., Bouchard, N., Kron, S.J., Jackson, S.P., and Côté, J (
Hudson, B.P., Martinez-Yamout, M.A., Dyson, H.J., and Wright, P.E. (
Owen, D.J., Ornaghi, P., Yang, J.C., Lowe, N., Evans, P.R., Ballario, P., Neuhaus, D., Filetici, P., and Travers, A.A. (
Jacobson, R.H., Ladurner, A.G., King, D.S., and Tjian, R. (
Kasten, M., Szerlong, H., Erdjument-Bromage, H., Tempst, P., Werner, M., and Cairns, B.R. (
Schiltz, R.L., Mizzen, C.A., Vassilev, A., Cook, R.G., Allis, C.D., and Nakatani, Y. (
Zhang, L., Eugeni, E.E., Parthun, M.K., and Freitas, M.A. (
Author notes
1Laboratory of Developmental Biology, Institute of Molecular and Cellular Biosciences, University of Tokyo, 1-1-1 Yayoi, Bunkyo-ku, Tokyo 113-0032; and 2Horikoshi Gene Selector Project, Exploratory Research for Advanced Technology (ERATO), Japan Science and Technology Agency (JST), 5-9-6 Tokodai, Tsukuba, Ibaraki 300-2635.