-
PDF
- Split View
-
Views
-
Cite
Cite
Ming-Hsiun Hsieh, Howard M. Goodman, A novel gene family in Arabidopsis encoding putative heptahelical transmembrane proteins homologous to human adiponectin receptors and progestin receptors, Journal of Experimental Botany, Volume 56, Issue 422, December 2005, Pages 3137–3147, https://doi.org/10.1093/jxb/eri311
Close - Share Icon Share
Abstract
A novel seven-transmembrane receptor family, that is comprised of human adiponectin receptors (AdipoRs) and membrane progestin receptors (mPRs) that share little sequence homology with all known G protein-coupled receptors (GPCRs), has been identified recently. Although a fish mPR has been suggested to be a GPCR, human AdipoRs seem to be structurally and functionally distinct from all known GPCRs. The identification of a novel gene family, the heptahelical protein (HHP) gene family, encoding proteins in Arabidopsis predicted to have a heptahelical transmembrane topology is reported here. There are at least five HHP genes in Arabidopsis whose encoded amino acid sequences have significant similarities to human AdipoRs and mPRs.The expression and regulation of the Arabidopsis HHP gene family has been studied here. The expression of the HHP gene family is differentially regulated by plant hormones. Steady-state levels of HHP1 mRNA are increased by treatments with abscisic acid and gibberellic acid, whereas levels of HHP2 mRNA are increased by abscisic acid and benzyladenine treatments. In addition, the expression of the HHP gene family is up-regulated by the presence of sucrose in the medium. Temperature and salt stress treatments also differentially affect the expression of the HHP genes. These novel seven-transmembrane proteins previously described in yeast and animals, and now identified in plants, may represent a new class of receptors that are highly conserved across kingdoms.
This paper is available online free of all access charges (see http://jxb.oxfordjournals.org/open_access.html for further details)
Introduction
A large number of protein receptors, characterized by a signature seven-transmembrane (7TM) motif, has been identified in animals. Members of this expanding superfamily include receptors for light, hormones, neurotransmitters, and sensory receptors for various odorants and taste (Bockaert and Pin, 1999; Pierce et al., 2002). These receptors are commonly referred to as G protein-coupled receptors (GPCRs), because most of them transmit the extracellular signals to the cytoplasm via heterotrimeric G proteins (Horn et al., 2003). Interestingly, a novel 7TM receptor family consisting of human adiponectin (also known as adipocyte complement-related protein of 30 kDa/ACRP30 or adipoQ) receptors (AdipoRs) and membrane progestin receptors (mPRs) that share little sequence similarity with all known GPCRs has been identified recently (Yamauchi et al., 2003; Zhu et al., 2003a, b). Similar sequences in mammals with homology to AdipoRs and mPRs have been deposited and annotated as the PAQR (progestin/adiponectin/adipoQ receptor) family in the GenBank database (http://www.ncbi.nlm.nih.gov/).
Adiponectin is an adipocyte-secreted hormone that plays an important role in regulating energy homeostasis and insulin sensitivity (Scherer et al., 1995; Hu et al., 1996; Berg et al., 2001, 2002; Fruebis et al., 2001; Yamauchi et al., 2001; Kubota et al., 2002; Maeda et al., 2002; Tsao et al., 2002). The insulin-sensitizing effect of adiponectin seems to be mediated by the recently discovered AdipoRs. It has been shown that interactions between adiponectin and AdipoRs increased AMP-activated protein kinase (AMPK) and peroxisome proliferators-activated receptor (PPAR)-α ligand activities, as well as fatty-acid oxidation and glucose uptake (Yamauchi et al., 2002, 2003). Human AdipoR1 and AdipoR2 are integral membrane proteins with an intracellular N terminus, a predicted 7TM domain and an extracellular C terminus (Yamauchi et al., 2003). The locations of the N terminus and the C terminus of human AdipoRs are opposite to the topology of all known GPCRs. In addition, the downstream signalling molecules of adiponectin, such as AMPK, PPAR-α, and p38MAPK, do not seem to be linked to the G protein-signalling pathway. Thus the 7TM containing AdipoRs may belong to a new receptor family, which are structurally and functionally distinct from GPCRs (Yamauchi et al., 2003).
By contrast, a new mPR has been proposed to be a novel GPCR (Zhu et al., 2003a, b). Usually, steroid signalling pathways are mediated by intracellular (nuclear) receptors to modulate the expression of target genes. However, there are increasing examples of steroid-induced responses that are too rapid to be mediated by transcription (Losel and Wehling, 2003). It is believed that these ‘non-genomic’ responses are mediated by specific steroid receptors localized on the plasma membranes. Progestin-induced oocyte maturation is one of the best-studied non-genomic responses (Maller, 2001). By expression screening, a novel mPR with predicted 7TM was identified in spotted seatrout (Cynoscion nebulosus) (Zhu et al., 2003a). The fish mPR has been shown to be a plasma membrane protein and the binding of progestin to mPR leads to inhibition of adenylyl cyclase in a pertussis toxin-sensitive manner, consistent with mPR being a GPCR (Zhu et al., 2003a). However, the location of the N terminus of mPR has yet to be confirmed. It is not clear whether mPRs are classical or novel type GPCRs.
In plants, signalling through heterotrimeric G proteins has been implicated in several phytohormone signal transduction pathways and in the control of cell division (Fujisawa et al., 2001; Ma, 2001; Assmann, 2002; Jones, 2002; Jones and Assmann, 2004). However, very few plant 7TM receptors or proteins involved in the G protein-coupled signal transduction pathway have been identified. Only a single gene, GCR1, encoding a putative GPCR with significant amino acid similarity to the Dictyostelium discoideum cAMP receptors has been identified in Arabidopsis (Josefsson and Rask, 1997; Plakidou-Dymock et al., 1998; Josefsson, 1999). The Arabidopsis GCR1 seems to have multiple functions. Plants overexpressing GCR1 have reduced seed dormancy and shortened flowering time (Colucci et al., 2002). The GCR1 protein has also been shown to regulate DNA synthesis, which is mediated by the phosphatidylinositol-specific phospholipase C (Apone et al., 2003). It has been shown that Arabidopsis GCR1 interacts with GPA1 (Gα) and regulates plant hormone abscisic acid (ABA) signalling (Pandey and Assmann, 2004). Arabidopsis GCR1 also has a role in seed germination that is, however, independent of the heterotrimeric G protein (Chen et al., 2004).
In addition to GCR1, a novel 7TM protein family related to barley MLO has been identified in Arabidopsis (Devoto et al., 1999, 2003). The barley Mlo gene encodes a novel 7TM protein that negatively regulates defense against powdery mildew and cell death (Buschges et al., 1997). It has been shown that the signalling activity of the 7TM protein MLO is enhanced by the interaction with calmodulin and that MLO-dependent defence suppression does not require heterotrimeric G proteins (Kim et al., 2002). The MLO-related protein family is the only abundant class of 7TM proteins unique to plants. There are at least 15 MLO genes in Arabidopsis (Devoto et al., 2003). It is not clear whether members of the MLO family are involved in other cellular activities mediated by heterotrimeric G proteins in plants. Interestingly, a novel seven-transmembrane regulator of G-protein signalling (RGS) protein has been identified in Arabidopsis (Chen et al., 2003). The AtRGS1 protein has an N-terminal seven transmembrane domain characteristic of GPCRs and a C-terminal RGS box. AtRGS1 modulates plant cell proliferation and functions in a hexokinase-independent glucose signalling pathway (Chen et al., 2003; Chen and Jones, 2004).
The identification is reported here of a novel 7TM-containing protein family, the heptahelical protein (HHP) family, in Arabidopsis that shows significant similarities to human AdipoRs and mPRs. In contrast to the previously identified plant 7TM proteins GCR1 and MLOs, the Arabidopsis HHP family and their homologues (the PAQR family) are highly conserved across kingdoms. There are at least five HHP genes in Arabidopsis that are differentially expressed in various organs. Moreover, the expression of the HHP gene family is induced by light or sucrose, but is differentially regulated by plant hormones, temperature, and salt stress. The evolutionary conservation and the regulation of the HHP gene expression suggest that these 7TM proteins may have important roles in plant developmental and physiological processes.
Materials and methods
Plant material and growth conditions
Arabidopsis thaliana ecotype Columbia-0 was used in all experiments. Plants grown in soil were placed in the greenhouse on a 16/8 h light/dark cycle at 23 °C. Total RNA extracted from roots, leaves, stems, flowers, and siliques from the same batch of 6-week-old soil-grown Arabidopsis plants was used for northern blot analysis. Two-week-old Arabidopsis seedlings grown in tissue culture on a 16/8 h light/dark cycle at 23 °C were used for HHP gene regulation studies. Arabidopsis seeds were sterilized and sown on nylon nets placed on MS (Murashige and Skoog salt) phytoagar plates [MS salts (GIBCO/BRL), pH adjusted to 5.7 with 1 N KOH, 0.8% (w/v) agar] containing 3% sucrose. For plant hormone treatments, the nylon nets with 2-week-old Arabidopsis seedlings were lifted and placed in 15×90×90 mm square plates with 15 ml H2O (control) or 50 μM plant hormones ABA, 1-aminocyclopropane-1-carboxylic acid (ACC) for ethylene, benzyladenine (BA), gibberellic acid (GA), and indole-3-acetic acid (IAA) for 4 h.
Sequence analysis
B. cereus Hly III, yeast YOL002c, and human AdipoR1 amino acid sequences were used to BLAST the Arabidopsis genome with default settings (http://www.ncbi.nlm.nih.gov/BLAST/). No significant similarity was found in the search with B. cereus Hly III. However, five expressed proteins At5g20270 (HHP1), At4g30850 (HHP2), At2g24150 (HHP3), At4g37680 (HHP4), and At4g38320 (HHP5), annotated as unknown proteins, shared significant similarities with yeast YOL002c and human AdipoR1 throughout their entire sequences. Two additional putative proteins At2g40710 (93 amino acids) and At4g38290 (170 amino acids) that share significant similarities with Arabidopsis HHPs and yeast YOL002c in the TM6 and TM7 regions were not considered further as members of the HHP family, because their predicted open reading frames are too short. In At4g30850 (HHP2), a difference resulting in an amino acid change was found between the genomic sequence and the cDNA sequence (AY074328) in Genbank. Reverse transcription-PCR was used to amplify full length HHP2 cDNA and three independent clones were sequenced. These cDNA sequences are identical to the genomic sequence, i.e. the 119th amino acid residue of HHP2 is methionine, instead of isoleucine as indicated in AY074328. The following programs were used for transmembrane predictions: TMHMM 2.0 (http://www.cbs.dtu.dk/services/TMHMM-2.0/), HMMTOP (http://www.enzim.hu/hmmtop/), TMPred (http://www.ch.embnet.org/software/TMPRED_form.html), TopPred 2.0 (http://www.sbc.su.se/∼erikw/toppred2/). A suite of algorithms for transmembrane preditions in the ARAMEMNON database (http://aramemnon.botanik.uni-koeln.de/) was also used to analyse the Arabidopsis HHPs with similar results (Schwacke et al., 2003).
The deduced amino acid sequences of Arabidopsis HHP proteins and their homologues in yeast and animals were aligned with the program ClustalX 1.8 with default settings. The sequence alignment was shaded with Macboxshade 2.15. Neighbor–Joining and a bootstrap analysis of 1000 replicates were performed to generate the phylogenetic tree (Kumar et al., 2001). The accession numbers for sequences are HHP1 (At5g20270, AY143975), HHP2 (AY267331), HHP3 (At2g24150, AF370179), HHP4 (At4g37680, AY056348), HHP5 (At4g38320, AY070414), rice AdipoR1 (BAD37427), rice AdipoR2 (BAD28011), rice AdipoR3 (AAM19133), yeast YOL002c (CAA99001), human AdipoR1 (NP_057083), human AdipoR2 (NP_078827), MPRA (NM_178422), MPRB (NP_588608), MPRG (NM_017705), Drosophila CG5315 (AAF56017), and C. elegans C43G2.1 (AAB09107). The accession number for B. cereus Hly III is P54176.
Northern blot analysis
Arabidopsis total RNA was isolated using a phenol extraction protocol (Lam et al., 1998). For the detection of HHP mRNA, digoxigenin (DIG)- or 32P-labelled single-stranded DNA probes were generated by PCR. The following primer pairs were used to make probes for RNA blot analysis. HHP1 (At5g20270): 5′-ATGGACCAAAATGGTCATAA-3′ and 5′-TTAACAACCAACGTGGTCACG-3′. HHP2 (At4g30850): 5′-GTGAAGGAATCTACGAAGAT-3′ and 5′-GTTGAACCGTCTGGAGTGAC-3′. HHP3 (At2g24150): 5′-TGCAATGGGATCATCTGCTG-3′ and 5′-GAAATAGCCATGGATTGTCC-3′. HHP4 (At4g37680): 5′-ATGGGTGATGAGGCAGAGAT-3′ and 5′-CAATCTTCATCTTCAACATC-3′. PCR products were sequenced to verify the identities of probes before labelling. DIG probe labelling, prehybridization, hybridization, wash conditions, and detection were performed according to the Boehringer-Mannheim Genius System User's Guide version 3.0. ULTRAhyb was used for 32P-labelled probes and prehybridization, hybridization, and wash conditions were performed according to the manufacturer's suggestion (Ambion, Austin, TX).
Results
Identification of a novel heptahelical transmembrane protein (HHP) family in Arabidopsis
Members of the uncharacterized protein family UPF0073 (http://pfam.wustl.edu) including Bacillus cereus haemolysin (Hly) III and Saccharomyces cerevisiae YOL002c are integral membrane proteins that are predicted to contain seven transmembrane (7TM) domains by several transmembrane prediction programs (data not shown). Interestingly, the yeast YOL002c protein is homologous to the recently cloned human adiponectin receptors (AdipoR1 and AdipoR2), which represent a novel type of 7TM receptors structurally and functionally distinct from the GPCRs (Yamauchi et al., 2003). To identify novel 7TM proteins in Arabidopsis, BLAST searches were conducted using the yeast YOL002c and human AdipoR1 as queries. Five Arabidopsis expressed proteins were identified, At2g24150, At4g30850, At4g37680, At4g38320, and At5g20270, which are highly similar to the yeast YOL002c and human adiponectin receptors. To determine whether these Arabidopsis proteins also contain seven transmembrane domains, several transmembrane prediction methods were used that rely on various algorithms (see Materials and methods). Most of the predictions suggest the existence of seven hydrophobic segments with the potential to form transmembrane domains in these Arabidopsis proteins. These Arabidopsis proteins have been named the HHP (HeptaHelical transmembrane Protein) family.
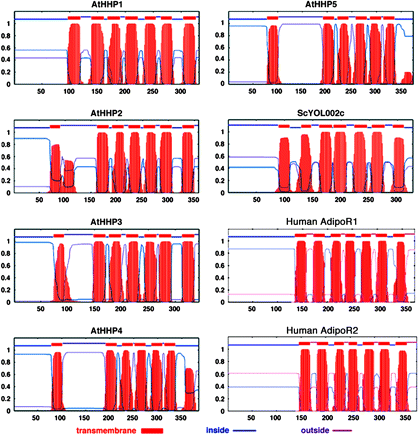
The topological predictions of Arabidopsis HHPs are shown in Fig. 1. The data presented here are from the transmembrane hidden Markov model (TMHMM) algorithm (Krogh et al., 2001). For Arabidopsis HHP1 to HHP3, the TMHMM program predicts a long intracellular N-terminal domain, 7TM domains, and a short extracellular C-terminal segment. The TMHMM program predicts a similar topology with 6TM domains for the Arabidopsis HHP4 and HHP5 proteins. While the TMHMM program predicts 6TM for HHP4 and HHP5, the HMMTOP program predicts 7TM for both HHP4 and HHP5 and the TMpred program predicts 7TM for HHP4 and 6TM for HHP5. Nonetheless, the real topology assignment of the Arabidopsis HHP protein family has to rely on further experimental data. For a comparison, the TMHMM predictions of yeast YOL002c, human AdipoR1 and AdipoR2 proteins are also shown in Fig. 1.
Predicted topology of Arabidopsis HHP1 to HHP5, yeast YOL002c, human AdipoR1, and human AdipoR2 proteins. The prediction is based on the TMHMM algorithm (Krogh et al., 2001). The numbers in each panel represent probability (y-axis) and amino acid position (x-axis). At, Arabidopsis thaliana; Sc, Saccharomyces cerevisiae.
Arabidopsis HHPs are homologous to human adiponectin receptors and membrane progestin receptors
Amino acid sequence alignment of the Arabidopsis HHPs, human AdipoR1, AdipoR2, MPRA, MPRB, and MPRG, and their homologues in yeast, Drosophila and C. elegans revealed that these novel proteins are highly conserved in both their primary sequences and their predicted 7TM topology (supplementary information can be found at JXB online). The Arabidopsis HHPs and human adiponectin receptors (AdipoR1 and AdipoR2) share 31.4% to 38.2% amino acid sequence similarities (Table 1). In addition, several unknown function proteins with significant sequence homology to Arabidopsis HHPs and human adiponectin receptors were found in a diverse set of eukaryotes including genetic model organisms Saccharomyces cerevisiae, Caenorhabditis elegans and Drosophila melanogaster. Interestingly, protein BLAST searches using Arabidopsis HHPs as queries revealed that the HHP protein family also shows amino acid sequence similarities (22.1–28.5%) to human membrane progestin receptors α, β, and γ (MPRA, MPRB, and MPRG) (Table 1). These recently discovered steroid membrane receptors are predicted to be 7TM proteins with characteristics of a G protein-coupled receptor (Zhu et al., 2003a, b). The amino acid sequence similarities among Arabidopsis HHPs, human adiponectin receptors (AdipoR1 and AdipoR2), human progestin receptors (MPRA, MPRB, and MPRG) and a representative member of the protein family from each organism (yeast YOL002c, Drosophila CG5315, and C. elegans C43G2.1) are shown in Table 1.
Amino acid similarities among Arabidopsis HHPs and their homologues in other species
. | AtHHP2 . | AtHHP3 . | AtHHP4 . | AtHHP5 . | Sc . | Hs . | Hs . | Dm . | Ce . | Hs . | Hs . | Hs . |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
. | . | . | . | . | YOL002c . | AdipoR1 . | AdipoR2 . | CG5315 . | C43G2.1 . | MPRG . | MPRA . | MPRB . |
| AtHHP1 | 52.2 | 53.0 | 46.3 | 44.0 | 39.9 | 35.8 | 35.2 | 39.5 | 29.3 | 25.4 | 26.0 | 27.7 |
| AtHHP2 | 77.3 | 46.0 | 44.7 | 36.4 | 34.6 | 31.4 | 37.6 | 29.1 | 24.6 | 26.2 | 25.5 | |
| AtHHP3 | 46.8 | 45.8 | 38.1 | 38.2 | 34.9 | 40.2 | 30.4 | 24.9 | 28.5 | 26.8 | ||
| AtHHP4 | 96.1 | 34.2 | 35.6 | 36.1 | 39.5 | 30.0 | 24.4 | 23.6 | 23.0 | |||
| AtHHP5 | 34.5 | 35.0 | 35.1 | 38.3 | 29.4 | 24.1 | 23.1 | 22.1 | ||||
| ScYOL002c | 42.0 | 39.9 | 49.1 | 38.0 | 29.7 | 30.1 | 28.2 | |||||
| HsAdipoR1 | 75.5 | 59.7 | 48.0 | 25.5 | 28.3 | 27.9 | ||||||
| HsAdipoR2 | 58.2 | 50.8 | 25.6 | 26.4 | 26.5 | |||||||
| DmCG5315 | 52.2 | 31.0 | 31.7 | 29.2 | ||||||||
| CeC43G2.1 | 22.1 | 22.9 | 21.5 | |||||||||
| HsMPRG | 35.2 | 39.2 | ||||||||||
| HsMPRA | 58.8 |
. | AtHHP2 . | AtHHP3 . | AtHHP4 . | AtHHP5 . | Sc . | Hs . | Hs . | Dm . | Ce . | Hs . | Hs . | Hs . |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
. | . | . | . | . | YOL002c . | AdipoR1 . | AdipoR2 . | CG5315 . | C43G2.1 . | MPRG . | MPRA . | MPRB . |
| AtHHP1 | 52.2 | 53.0 | 46.3 | 44.0 | 39.9 | 35.8 | 35.2 | 39.5 | 29.3 | 25.4 | 26.0 | 27.7 |
| AtHHP2 | 77.3 | 46.0 | 44.7 | 36.4 | 34.6 | 31.4 | 37.6 | 29.1 | 24.6 | 26.2 | 25.5 | |
| AtHHP3 | 46.8 | 45.8 | 38.1 | 38.2 | 34.9 | 40.2 | 30.4 | 24.9 | 28.5 | 26.8 | ||
| AtHHP4 | 96.1 | 34.2 | 35.6 | 36.1 | 39.5 | 30.0 | 24.4 | 23.6 | 23.0 | |||
| AtHHP5 | 34.5 | 35.0 | 35.1 | 38.3 | 29.4 | 24.1 | 23.1 | 22.1 | ||||
| ScYOL002c | 42.0 | 39.9 | 49.1 | 38.0 | 29.7 | 30.1 | 28.2 | |||||
| HsAdipoR1 | 75.5 | 59.7 | 48.0 | 25.5 | 28.3 | 27.9 | ||||||
| HsAdipoR2 | 58.2 | 50.8 | 25.6 | 26.4 | 26.5 | |||||||
| DmCG5315 | 52.2 | 31.0 | 31.7 | 29.2 | ||||||||
| CeC43G2.1 | 22.1 | 22.9 | 21.5 | |||||||||
| HsMPRG | 35.2 | 39.2 | ||||||||||
| HsMPRA | 58.8 |
Amino acid similarities among Arabidopsis HHPs and their homologues in other species
. | AtHHP2 . | AtHHP3 . | AtHHP4 . | AtHHP5 . | Sc . | Hs . | Hs . | Dm . | Ce . | Hs . | Hs . | Hs . |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
. | . | . | . | . | YOL002c . | AdipoR1 . | AdipoR2 . | CG5315 . | C43G2.1 . | MPRG . | MPRA . | MPRB . |
| AtHHP1 | 52.2 | 53.0 | 46.3 | 44.0 | 39.9 | 35.8 | 35.2 | 39.5 | 29.3 | 25.4 | 26.0 | 27.7 |
| AtHHP2 | 77.3 | 46.0 | 44.7 | 36.4 | 34.6 | 31.4 | 37.6 | 29.1 | 24.6 | 26.2 | 25.5 | |
| AtHHP3 | 46.8 | 45.8 | 38.1 | 38.2 | 34.9 | 40.2 | 30.4 | 24.9 | 28.5 | 26.8 | ||
| AtHHP4 | 96.1 | 34.2 | 35.6 | 36.1 | 39.5 | 30.0 | 24.4 | 23.6 | 23.0 | |||
| AtHHP5 | 34.5 | 35.0 | 35.1 | 38.3 | 29.4 | 24.1 | 23.1 | 22.1 | ||||
| ScYOL002c | 42.0 | 39.9 | 49.1 | 38.0 | 29.7 | 30.1 | 28.2 | |||||
| HsAdipoR1 | 75.5 | 59.7 | 48.0 | 25.5 | 28.3 | 27.9 | ||||||
| HsAdipoR2 | 58.2 | 50.8 | 25.6 | 26.4 | 26.5 | |||||||
| DmCG5315 | 52.2 | 31.0 | 31.7 | 29.2 | ||||||||
| CeC43G2.1 | 22.1 | 22.9 | 21.5 | |||||||||
| HsMPRG | 35.2 | 39.2 | ||||||||||
| HsMPRA | 58.8 |
. | AtHHP2 . | AtHHP3 . | AtHHP4 . | AtHHP5 . | Sc . | Hs . | Hs . | Dm . | Ce . | Hs . | Hs . | Hs . |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
. | . | . | . | . | YOL002c . | AdipoR1 . | AdipoR2 . | CG5315 . | C43G2.1 . | MPRG . | MPRA . | MPRB . |
| AtHHP1 | 52.2 | 53.0 | 46.3 | 44.0 | 39.9 | 35.8 | 35.2 | 39.5 | 29.3 | 25.4 | 26.0 | 27.7 |
| AtHHP2 | 77.3 | 46.0 | 44.7 | 36.4 | 34.6 | 31.4 | 37.6 | 29.1 | 24.6 | 26.2 | 25.5 | |
| AtHHP3 | 46.8 | 45.8 | 38.1 | 38.2 | 34.9 | 40.2 | 30.4 | 24.9 | 28.5 | 26.8 | ||
| AtHHP4 | 96.1 | 34.2 | 35.6 | 36.1 | 39.5 | 30.0 | 24.4 | 23.6 | 23.0 | |||
| AtHHP5 | 34.5 | 35.0 | 35.1 | 38.3 | 29.4 | 24.1 | 23.1 | 22.1 | ||||
| ScYOL002c | 42.0 | 39.9 | 49.1 | 38.0 | 29.7 | 30.1 | 28.2 | |||||
| HsAdipoR1 | 75.5 | 59.7 | 48.0 | 25.5 | 28.3 | 27.9 | ||||||
| HsAdipoR2 | 58.2 | 50.8 | 25.6 | 26.4 | 26.5 | |||||||
| DmCG5315 | 52.2 | 31.0 | 31.7 | 29.2 | ||||||||
| CeC43G2.1 | 22.1 | 22.9 | 21.5 | |||||||||
| HsMPRG | 35.2 | 39.2 | ||||||||||
| HsMPRA | 58.8 |
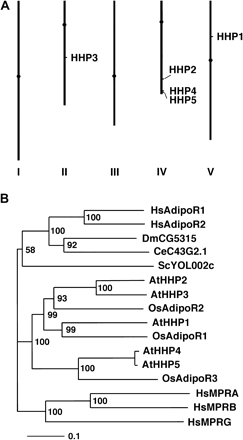
Among the Arabidopsis HHPs, HHP2 is most similar to HHP3 (69.8% identity and 77.3% similarity), and the amino acid sequences of HHP4 and HHP5 are almost identical (95.3% identity and 96.1% similarity). In addition, the genomic DNA sequences of HHP4 and HHP5 are also almost identical throughout the genes including exons, introns, and the putative 5′- and 3′-untranslated regions (data not shown). The chromosomal locations of the HHP gene family are shown in Fig. 2A. To identify HHP homologues in the other plant species, Arabidopsis HHPs were used as queries to search the database. There are at least three rice (Oryza sativa) proteins (accession numbers BAD37427, BAD28011, and AAM19133) that show significant similarity to the Arabidopsis HHPs. The sequences of BAD37427 and BAD28011 were annotated as ‘putative adiponectin receptor 1’ and the AAM19133 sequence was annotated as ‘hypothetical protein’ in the database. To avoid confusion, the BAD37427, BAD28011, and AAM19133 proteins were named rice AdipoR1, AdipoR2, and AdipoR3, respectively. A phylogenetic tree generated from alignment of the Arabidopsis HHP family, rice AdipoRs, yeast YOL002c, human AdipoR1, AdipoR2, MPRA, MPRB, and MPRG, Drosophila CG5315, and C. elegans C43G2.1 is shown in Fig. 2B. The amino acid sequence similarities between AdipoRs and mPRs in humans are only 25.5% to 28.3% (Table 1). In the phylogenetic tree, these two novel human receptor families belong to two different clades (Fig. 2B).
(A) Chromosomal locations of the Arabidopsis HHP genes. (B) Phylogenetic analysis of Arabidopsis HHPs, rice putative AdipoRs, yeast YOL002c, human AdipoRs, human mPRs, fly CG5315, and worm C43G2.1 proteins. The sequence alignment was generated from the ClustalX 1.8 program. Neighbor–Joining and bootstrap analyses were performed to generate the phylogenetic tree. A bootstrap value is provided for each node. These values indicate the percentage of bootstrap replicates that support this node out of 1000 samples. At, Arabidopsis thaliana; Os, Oryza sativa; Sc, Saccharomyces cerevisiae; Hs, Homo sapiens; Dm, Drosophila melanogaster; Ce, Caenorhabditis elegans.
Expression patterns of the Arabidopsis HHP gene family
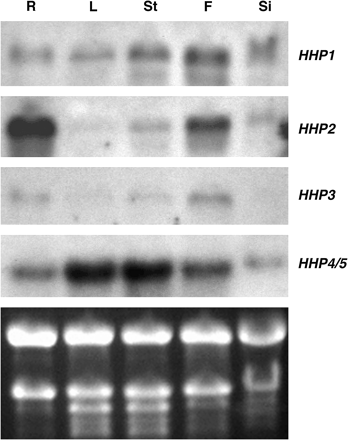
The conservation of this protein family across different kingdoms suggests that members of this uncharacterized protein family may have an important role during growth and development. To elucidate the function of Arabidopsis HHPs, the expression patterns of the HHP gene family was first examined in various organs by RNA gel-blot analysis. Total RNA extracted from roots, leaves, stems, flowers, and siliques of soil-grown 6-week-old Arabidopsis was used and distinct expression patterns for the HHP gene family were observed (Fig. 3). The HHP1 mRNA was detected in all organs tested with the highest levels in flowers, while the highest levels of HHP2 mRNA were detected in roots. The HHP2 mRNA also showed high levels in flowers, moderate levels in stems and siliques, and low levels in leaves. Levels of HHP3 mRNA were high in flowers, moderate in roots, low in leaves and stems, and undetectable in siliques. Because the nucleic acid sequences of HHP4 and HHP5 are almost identical, the signals detected by the HHP4-derived probe may represent both HHP4 and HHP5 transcripts. In contrast to the expression patterns of HHP2 and HHP3, high levels of HHP4/5 mRNA were detected in leaves and stems.
HHP gene family expression in various organs of 6-week-old Arabidopsis. Total RNA (10 μg) from roots (R), leaves (L), stems (St), flowers (F), and siliques (Si) was used for RNA gel-blot analysis and hybridized with gene-specific probes of HHP1 to HHP3. Because the DNA sequences of HHP4 and HHP5 genes are almost identical, the HHP4-derived probe may cross-hybridize to the HHP5 mRNA. Thus the signals detected in the RNA gel-blot by the HHP4 probe may represent the sum of HHP4 and HHP5 transcripts. The ethidium bromide-stained agarose gel of the same samples is shown at the bottom.
Effects of plant hormones on the expression of the Arabidopsis HHP gene family
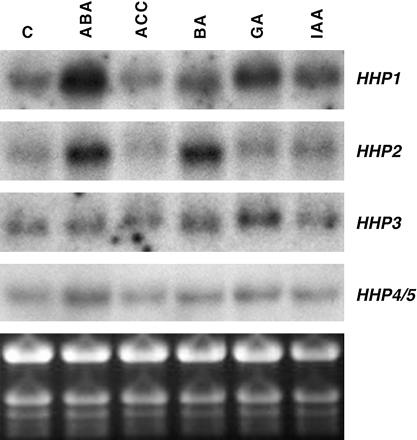
The effects of five classical plant hormones, ABA, ethylene, cytokinin, GA, and auxin, on the expression of the HHP gene family were examined. Two-week-old Arabidopsis seedlings grown in tissue culture were treated with plant hormones and total RNA extracted from these samples was used for RNA gel-blot analysis to detect the expression of HHP1 to HHP5 (Fig. 4). Compared with the untreated sample, steady-state levels of HHP1 mRNA are increased by treatments with ABA and GA, whereas ACC, BA, and IAA have little or no effects on the accumulation of HHP1 mRNA. While treatments with ABA and BA induce the expression of HHP2, ACC, GA, and IAA have no significant effects on the levels of HHP2 mRNA. The expression of HHP3 is slightly increased by GA, whereas ABA, ACC, BA, and IAA have no effects on the accumulation of HHP3 mRNA. By contrast, levels of HHP4/5 mRNA are slightly increased by ABA. Treatments with ACC, BA, GA, and IAA have little or no effects on the expression of HHP4/5.
Plant hormones differentially regulate steady-state levels of HHP mRNA. Total RNA (10 μg) from 2-week-old Arabidopsis seedlings treated with plant hormones for 4 h was used for RNA gel-blot analysis to detect levels of HHP1 to HHP4/5 mRNA. ABA and GA induced the expression of HHP1 (compared to C, control), whereas ABA and BA significantly increased the expression levels of HHP2. The ethidium bromide-stained gel is shown at the bottom.
Light/dark and sucrose effects on the expression of the HHP gene family
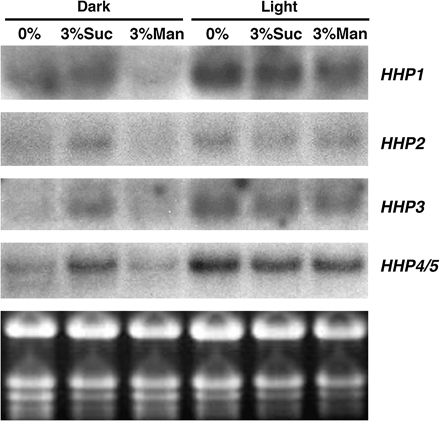
To test the effects of light/dark and sucrose treatments on the expression of the HHP gene family, Arabidopsis seedlings were grown under normal light conditions (16/8 h light/dark cycle) for 2 weeks and were subsequently placed in continuous light or in the dark for 48 h. During the 48 h dark or light treatments, plants were grown in media containing 0% sucrose, 3% sucrose, or 3% mannitol. The non-metabolizable sugar, mannitol, was included as an osmotic control. Total RNA extracted from these samples was used for RNA gel-blot analysis to detect the steady-state levels of HHP1 to HHP4/5 mRNA. The results are shown in Fig. 5. In dark-adapted seedlings, steady-state levels of HHP1 to HHP4/5 mRNA are low in the absence of sucrose. Supplementation of the media with 3% sucrose increases the levels of HHP1 to HHP4/5 mRNA. This sucrose effect is not due to an osmotic change, because the addition of 3% mannitol to the medium has no significant effect on the accumulation of HHP1 to HHP4/5 mRNA. In dark-adapted samples, especially dark plus 0% sucrose, an additional band smaller than the expected HHP4 or HHP5 mRNA was detected. The identity of this band is unknown. In light-treated plants, levels of HHP1 to HHP4/5 mRNA are high regardless of the amount of sucrose in the media. In the absence of sucrose, the light treatment significantly increases levels of HHP1 to HHP4/5 mRNA (compare dark 0% to light 0% sucrose). Light plus sucrose cannot induce the accumulation of HHP1 to HHP4/5 mRNA beyond the light-induced levels (compare dark 0%, light 0%, and light 3% sucrose). Moreover, the induction of HHP1 to HHP4/5 by light in the absence of sucrose is comparable to the sucrose-induced levels independent of light (compare dark 0%, dark 3%, and light 0% sucrose). These results suggest that the induction of HHP1 to HHP4/5 by light may primarily derive from photosynthetic products.
Sucrose induces the accumulation of HHP1 to HHP4/5 mRNA independent of light. Two-week-old Arabidopsis plants grown on MS medium plus 3% sucrose in a normal 16/8 h- light/dark cycle were transferred to fresh MS media with 0% sucrose, 3% sucrose, or 3% mannitol and then left in the dark or transferred to continuous light for 2 d before harvesting the whole plant for RNA extraction. Replicate blots of 10 μg of total RNA were hybridized to detect levels of HHP1 to HHP4/5 mRNA. The ethidium bromide-stained gel is shown at the bottom.
Effects of temperature and salt stress on the expression of the HHP gene family
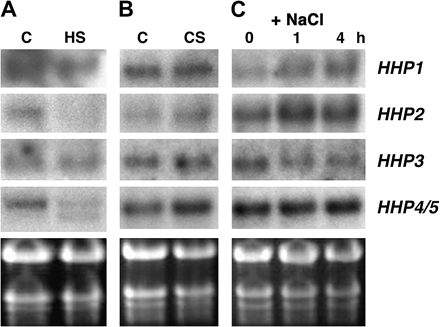
The effect of heat stress on the expression of the HHP gene family (Fig. 6A) was examined. After treatment at 37 °C for 2 h, the levels of HHP1, HHP2, and HHP4/5 mRNA decreased, whereas the steady-state levels of HHP3 mRNA were not affected. Similar to the effect of the dark treatment, an additional lower band was detected by the HHP4 probe in the heat-treated sample. When 2-week-old Arabidopsis plants were treated with 4 °C for 2 h, the accumulation of HHP2 mRNA increased slightly and little or no significant changes in steady-state levels of HHP1, HHP3, and HHP4/5 mRNA was observed (Fig. 6B). The effect of salt stress on the expression of the Arabidopsis HHP gene family was also examined. Treatments with 250 mM NaCl for 1 h or 4 h also differentially affect the expression of the HHP gene family. Steady-state levels of HHP1 and HHP2 mRNA are increased, whereas levels of HHP3 mRNA are slightly decreased and the accumulation of HHP4/5 mRNA is not affected by the salt treatment (Fig. 6C).
The expression of HHP genes is differentially regulated by heat, cold, and salt treatment. Total RNA (10 μg) from 2-week-old Arabidopsis seedlings treated with 37 °C for 2 h (A), 4 °C for 2 h (B), or 250 mM NaCl for 0 h, 1 h, or 4 h (C) was used for RNA gel-blot analysis to detect levels of HHP1 to HHP4/5 mRNA. C, control; HS, heat shock; CS, cold shock. The ethidium bromide-stained gel is shown at the bottom.
Expression patterns of the HHP gene family in the public microarray database
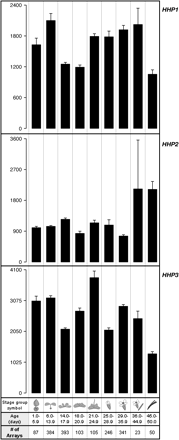
To gain a broader view regarding the transcriptional regulation of the Arabidopsis HHP gene family, the Genevestigator database (https://www.genevestigator.ethz.ch/) that archives publicly accessible microarray results (Zimmermann et al., 2004) was searched. Consistent with the northern blot analyses (Fig. 4), the expression of HHP1 and HHP2 was induced by the plant hormone ABA (Table 2). The treatments that affect the expression of HHP1, HHP2, and HHP3 genes in a ratio (treatment/control) greater than 2 (up-regulation) or less than 0.5 (down-regulation) have been summarized in Table 2. Because the probe sets used in the Affymetrix microarray cannot distinguish the signals for HHP4 and HHP5, only the data mining results for HHP1, HHP2, and HHP3 are listed. The expression of HHP1 is induced by ethylene inhibitors silver nitrate (AgNO3) and aminoethoxyvinyl glycine (AVG), salicyclic acid, and the auxin transport inhibitor 2,3,5-triiodobenzoic acid (TIBA). By contrast, the treatment of syringolin, a peptide elicitor, significantly reduced the expression of HHP1. The expression of HHP2 is up-regulated in the Agrobacterium tumefaciens-infected tumour tissue, by treatments of protein synthesis inhibitor cycloheximide, peptide elicitor syringolin, and senescence-induced programmed cell death. The expression of HHP3 is induced by senescence, whereas treatments of cycloheximide and syringolin repressed it. In addition to the effect of various treatments on the expression of HHPs, the Genevestigator database was also surveyed to examine the expression patterns of these genes at nine different growth stages (Fig. 7). These results provide some insights into the regulation of Arabidopsis HHP genes under various conditions. However, these data together with our studies cannot really define the functions of these novel 7TM proteins in plants. Several putative T-DNA insertion lines in the Arabidopsis HHP genes are available in the SALK ‘T-DNA express’ database (http://signal.salk.edu/cgi-bin/tdnaexpress) (Table 3). Further studies on the HHP mutants may unravel the functions of these novel proteins in plants.
Expression levels of HHP genes at different growth stages derived from the Genevestigator database (https://www.genevestigator.ethz.ch/). Data shown are means ±standard errors. y-axis represents the signal intensity detected in the microarray chips. The age and number of arrays are indicated at the bottom. Please note that the growth stage symbols at the bottom do not represent organs.
Response of Arabidopsis HHP genes to various treatments in the Genevestigator microarray database (https://www.genevestigator.ethz.ch/)
Treatmenta . | . | Ratio (Treatment/Control) . | . | . | ||
|---|---|---|---|---|---|---|
. | . | HHP1 . | HHP2 . | HHP3 . | ||
| Hormone: | ABA (1 h) | 2.27 | 1.52 | 1.2 | ||
| ABA (3 h) | 7.69 | 4.39 | 0.8 | |||
| Biotic: | A. tumefaciens | 0.79 | 8.19 | 0.66 | ||
| Chemical: | AgNO3 | 2.25 | 1.9 | 0.7 | ||
| AVG | 2.11 | 1.26 | 1.33 | |||
| Cycloheximide | 0.79 | 3.05 | 0.15 | |||
| Salicyclic acid | 2.15 | 0.95 | 0.77 | |||
| Syringolin | 0.17 | 5.22 | 0.38 | |||
| TIBA | 3.31 | 1.76 | 0.52 | |||
| PCD: | Senescence | 1.68 | 3.16 | 2.83 | ||
Treatmenta . | . | Ratio (Treatment/Control) . | . | . | ||
|---|---|---|---|---|---|---|
. | . | HHP1 . | HHP2 . | HHP3 . | ||
| Hormone: | ABA (1 h) | 2.27 | 1.52 | 1.2 | ||
| ABA (3 h) | 7.69 | 4.39 | 0.8 | |||
| Biotic: | A. tumefaciens | 0.79 | 8.19 | 0.66 | ||
| Chemical: | AgNO3 | 2.25 | 1.9 | 0.7 | ||
| AVG | 2.11 | 1.26 | 1.33 | |||
| Cycloheximide | 0.79 | 3.05 | 0.15 | |||
| Salicyclic acid | 2.15 | 0.95 | 0.77 | |||
| Syringolin | 0.17 | 5.22 | 0.38 | |||
| TIBA | 3.31 | 1.76 | 0.52 | |||
| PCD: | Senescence | 1.68 | 3.16 | 2.83 | ||
ABA, abscisic acid; A. tumefaciens, Agrobacterium tumefaciens; AVG, aminoethoxyvinyl glycine; TIBA, 2,3,5-triiodobenzoic acid; PCD, programmed cell death.
Response of Arabidopsis HHP genes to various treatments in the Genevestigator microarray database (https://www.genevestigator.ethz.ch/)
Treatmenta . | . | Ratio (Treatment/Control) . | . | . | ||
|---|---|---|---|---|---|---|
. | . | HHP1 . | HHP2 . | HHP3 . | ||
| Hormone: | ABA (1 h) | 2.27 | 1.52 | 1.2 | ||
| ABA (3 h) | 7.69 | 4.39 | 0.8 | |||
| Biotic: | A. tumefaciens | 0.79 | 8.19 | 0.66 | ||
| Chemical: | AgNO3 | 2.25 | 1.9 | 0.7 | ||
| AVG | 2.11 | 1.26 | 1.33 | |||
| Cycloheximide | 0.79 | 3.05 | 0.15 | |||
| Salicyclic acid | 2.15 | 0.95 | 0.77 | |||
| Syringolin | 0.17 | 5.22 | 0.38 | |||
| TIBA | 3.31 | 1.76 | 0.52 | |||
| PCD: | Senescence | 1.68 | 3.16 | 2.83 | ||
Treatmenta . | . | Ratio (Treatment/Control) . | . | . | ||
|---|---|---|---|---|---|---|
. | . | HHP1 . | HHP2 . | HHP3 . | ||
| Hormone: | ABA (1 h) | 2.27 | 1.52 | 1.2 | ||
| ABA (3 h) | 7.69 | 4.39 | 0.8 | |||
| Biotic: | A. tumefaciens | 0.79 | 8.19 | 0.66 | ||
| Chemical: | AgNO3 | 2.25 | 1.9 | 0.7 | ||
| AVG | 2.11 | 1.26 | 1.33 | |||
| Cycloheximide | 0.79 | 3.05 | 0.15 | |||
| Salicyclic acid | 2.15 | 0.95 | 0.77 | |||
| Syringolin | 0.17 | 5.22 | 0.38 | |||
| TIBA | 3.31 | 1.76 | 0.52 | |||
| PCD: | Senescence | 1.68 | 3.16 | 2.83 | ||
ABA, abscisic acid; A. tumefaciens, Agrobacterium tumefaciens; AVG, aminoethoxyvinyl glycine; TIBA, 2,3,5-triiodobenzoic acid; PCD, programmed cell death.
T-DNA insertion lines in the Arabidopsis HHP gene family
Gene . | T-DNA insertion line . |
|---|---|
| HHP1 | SALK_008343, SALK_056174 |
| HHP2 | SALK_048055, SALK_048056, SAIL_822_C02 |
| HHP3 | SALK_044573, SALK_151347, GABI_610E10, GABI_357D09 |
| HHP4 | SAIL_569_F11, SAIL_844_C10 |
| HHP5 | SALK_104717 |
Gene . | T-DNA insertion line . |
|---|---|
| HHP1 | SALK_008343, SALK_056174 |
| HHP2 | SALK_048055, SALK_048056, SAIL_822_C02 |
| HHP3 | SALK_044573, SALK_151347, GABI_610E10, GABI_357D09 |
| HHP4 | SAIL_569_F11, SAIL_844_C10 |
| HHP5 | SALK_104717 |
T-DNA insertion lines in the Arabidopsis HHP gene family
Gene . | T-DNA insertion line . |
|---|---|
| HHP1 | SALK_008343, SALK_056174 |
| HHP2 | SALK_048055, SALK_048056, SAIL_822_C02 |
| HHP3 | SALK_044573, SALK_151347, GABI_610E10, GABI_357D09 |
| HHP4 | SAIL_569_F11, SAIL_844_C10 |
| HHP5 | SALK_104717 |
Gene . | T-DNA insertion line . |
|---|---|
| HHP1 | SALK_008343, SALK_056174 |
| HHP2 | SALK_048055, SALK_048056, SAIL_822_C02 |
| HHP3 | SALK_044573, SALK_151347, GABI_610E10, GABI_357D09 |
| HHP4 | SAIL_569_F11, SAIL_844_C10 |
| HHP5 | SALK_104717 |
Discussion
The Arabidopsis HHPs share significant homology with the newly discovered AdipoRs and mPRs (the PAQR family) that are highly conserved from yeast to humans (Karpichev et al., 2002; Yamauchi et al., 2003; Zhu et al., 2003a, b). Members of this protein family are predicted to contain a 7TM domain, which is a signature motif for all known GPCRs. Beyond the conserved 7TM architecture, Arabidopsis HHPs and members of the PAQR family found in other organisms share little amino acid sequence homology with GPCRs or other known 7TM-containing proteins in the GenBank database. The conventional GPCRs possess an extracellular N-terminus, a 7TM domain, and a cytoplasmic C-terminus. However, despite the TMHMM prediction that the human AdipoR2 has an extracellular N-terminus (Fig. 1), the N-terminus of both human AdipoR1 and AdipoR2 has been demonstrated to be cytoplasmic (Yamauchi et al., 2003). Although the topology of human mPRs has been hypothesized to be similar to classical GPCRs, the proposed model was not derived from experimental data (Zhu et al., 2003a, b). Regardless of the TMHMM prediction of a cytoplasmic N-terminus for the Arabidopsis HHPs (Fig. 1), the real topology of these proteins has yet to be confirmed by further experiments.
G proteins have been implicated in ABA signalling, GA responses, and auxin-mediated cell proliferation in plants (Fujisawa et al., 2001; Ma, 2001; Assmann, 2002; Jones, 2002; Jones and Assmann, 2004). Interestingly, the expression of the Arabidopsis HHP gene family is differentially regulated by plant hormones (Fig. 4). The expression of both HHP1 and HHP2 is induced by ABA. In addition, GA and BA also induce the expression of HHP1 and HHP2, respectively. It will be interesting to test whether these Arabidopsis 7TM-containing proteins are involved in the ABA and GA signalling pathways. The yeast G protein-coupled receptor Gpr1 has been suggested to be involved in sugar sensing (Forsberg and Ljungdahl, 2001; Versele et al., 2001). The involvement of plant G proteins or GPCRs in sugar sensing and signalling is less clear (Rolland et al., 2002). The expression levels of HHP1 to HHP4/5 mRNA are increased by sucrose (Fig. 5). It will be interesting to examine if these 7TM proteins play a role in sensing intracellular or extracellular nutrients.
The PAQR homologues are broadly distributed across kingdoms. However, functions of these proteins are largely unknown. Of the few characterized PAQR family members, the yeast YOL002c protein has been shown to be involved in lipid and phosphate metabolism (Karpichev et al., 2002). Recently, it has been shown that the yeast YOL002c protein is also involved in zinc metabolism (Lyons et al., 2004) and osmotin-induced cell death via a RAS2 signalling pathway in yeast (Narasimhan et al., 2005). Human AdipoR1 and AdipoR2, which have been suggested to be functionally and structurally distinct from GPCRs, mediate the insulin-sensitizing effect of adiponectin by increasing fatty-acid oxidation and glucose uptake (Yamauchi et al., 2003). It is interesting that both yeast YOL002c protein and human AdipoRs have a role in fatty acid metabolism. The underlying molecular mechanisms of yeast YOL002c in fatty acid metabolism remain unknown. By contrast, the human AdipoRs, which were identified by specific binding to adiponectin, have been placed in the upstream of the adiponectin signal transduction pathway (Yamauchi et al., 2003). The paradigm of the adiponectin signalling pathway in mammals includes the binding of adiponectin to AdipoRs, which, in turn, increases the activities of AMPK and the nuclear receptor PPARα to activate the oxidation of fatty acids (Yamauchi et al., 2002, 2003). It is interesting to speculate that the plant homologues of AdipoRs such as Arabidopsis HHPs may also be involved in regulating fatty acid and glucose metabolism in order to maintain energy (i.e. ATP) homeostasis inside a cell.
In animals, classical steroid hormone signalling pathways are mediated by nuclear receptors. Interestingly, a fish mPR with characteristics of GPCRs has been shown to mediate progestin-induced oocyte maturation (Zhu et al., 2003a, b). The identification of vertebrate mPRs has further redefined the steroid-mediated signalling in the non-genomic aspect (Losel and Wehling, 2003; Zhu et al., 2003a, b). By contrast, plant steroid hormones mainly act through cell surface proteins, independently of the well-studied animal nuclear receptors (Thummel and Chory, 2002). For instance, the Arabidopsis brassinolide is perceived by the membrane leucine-rich repeat (LRR) kinase, BRI1, which then signals through a phosphorylation cascade to regulate the expression of downstream target genes (Li and Chory, 1997; He et al., 2000; Thummel and Chory, 2002). The identification of mPR homologues in plants raises the possibility that these 7TM proteins may also serve as steroid receptors to regulate some of the developmental and physiological processes in plants. If the Arabidopsis HHPs function as receptors, it will be interesting to identify the specific ligands, the downstream effectors and the relevant physiological responses mediated by these molecules.
Although a search of GenBank with the bacterial Hly III sequence failed to identify homologues in Arabidopsis, further sequence analysis revealed that Arabidopsis HHPs and B. cereus Hly III protein do share 21.7% to 25.3% amino acid sequence similarities (data not shown). A PSI-BLAST search with the B. cereus Hly III sequence revealed that its homologues with unknown function exist in a broad range of bacteria (data not shown). All the bacterial Hly III homologues are about 220 amino acids in length and are mainly composed of 7TM domains as revealed by topographic analysis with the TMHMM program (data not shown). It is conceivable that the PAQR family, Arabidopsis HHPs, B. cereus Hly III, and other bacterial Hly III homologues are evolved from a common ancestral 7TM protein. There is very limited knowledge about bacterial Hly III. The B. cereus Hly III is an extracellular protein that can be inserted into the membrane of erythrocytes and acts as an oligomeric pore-forming haemolysin (Baida and Kuzmin, 1995, 1996). It is possible that Arabidopsis HHPs may also function as channel proteins. Further studies of this gene family promise to add insight into the functional conservation or divergence of these novel 7TM proteins.
Supplementary material
The sequence alignment of Arabidopsis HHPs and their representative homologues from humans, yeast, Drosophila, and C. elegans is available at JXB online.
We thank Drs Lance Davidow and Julie Nardone for critical reading of the manuscript.
References
Apone F, Alyeshmerni N, Wiens K, Chalmers D, Chrispeels MJ, Colucci G.
Assmann SM.
Baida GE, Kuzmin NP.
Baida GE, Kuzmin NP.
Berg AH, Combs TP, Du X, Brownlee M, Scherer PE.
Berg AH, Combs TP, Scherer PE.
Bockaert J, Pin JP.
Buschges R, Hollricher K, Panstruga R, et al.
Chen JG, Willard FS, Huang J, Liang J, Chasse SA, Jones AM, Siderovski DP.
Chen J-G, Jones AM.
Chen J-G, Pandey S, Huang J, Alonso JM, Ecker JR, Assmann SM, Jones AM.
Colucci G, Apone F, Alyeshmerni N, Chalmers D, Chrispeels MJ.
Devoto A, Hartmann HA, Piffanelli P, et al.
Devoto A, Piffanelli P, Nilsson I, Wallin E, Panstruga R, von Heijne G, Schulze-Lefert P.
Forsberg H, Ljungdahl PO.
Fruebis J, Tsao TS, Javorschi S, Ebbets-Reed D, Erickson MR, Yen FT, Bihain BE, Lodish HF.
Fujisawa Y, Kato H, Iwasaki Y.
He Z, Wang ZY, Li J, Zhu Q, Lamb C, Ronald P, Chory J.
Horn F, Bettler E, Oliveira L, Campagne F, Cohen FE, Vriend G.
Hu E, Liang P, Spiegelman BM.
Jones AM.
Jones AM, Assmann SM.
Josefsson L, Rask L.
Josefsson LG.
Karpichev IV, Cornivelli L, Small GM.
Kim MC, Panstruga R, Elliott C, Muller J, Devoto A, Yoon HW, Park HC, Cho MJ, Schulze-Lefert P.
Krogh A, Larsson B, von Heijne G, Sonnhammer EL.
Kubota N, Terauchi Y, Yamauchi T, et al.
Kumar S, Tamura K, Jakobsen IB, Nei M.
Lam HM, Hsieh MH, Coruzzi G.
Li J, Chory J.
Losel R, Wehling M.
Lyons TJ, Villa NY, Regalla LM, Kupchak BR, Vagstad A, Eide DJ.
Maeda N, Shimomura I, Kishida K, et al.
Maller JL.
Narasimhan ML, Coca MA, Jin J, et al.
Pandey S, Assmann SM.
Pierce KL, Premont RT, Lefkowitz RJ.
Plakidou-Dymock S, Dymock D, Hooley R.
Rolland F, Moore B, Sheen J.
Scherer PE, Williams S, Fogliano M, Baldini G, Lodish HF.
Schwacke R, Schneider A, van der Graaff E, Fischer K, Catoni E, Desimone M, Frommer WB, Flugge UI, Kunze R.
Thummel CS, Chory J.
Tsao TS, Lodish HF, Fruebis J.
Versele M, Lemaire K, Thevelein JM.
Yamauchi T, Kamon J, Ito Y, et al.
Yamauchi T, Kamon J, Minokoshi Y, et al.
Yamauchi T, Kamon J, Waki H, et al.
Zhu Y, Bond J, Thomas P.
Zhu Y, Rice CD, Pang Y, Pace M, Thomas P.




Comments