-
PDF
- Split View
-
Views
-
Cite
Cite
Scott William Roy, Manuel Irimia, David Penny, Very Little Intron Gain in Entamoeba histolytica Genes Laterally Transferred from Prokaryotes, Molecular Biology and Evolution, Volume 23, Issue 10, October 2006, Pages 1824–1827, https://doi.org/10.1093/molbev/msl061
Close - Share Icon Share
Abstract
The evolution of spliceosomal introns remains intensely debated. We studied 96 Entamoeba histolytica genes previously identified as having been laterally transferred from prokaryotes, which were presumably intronless at the time of transfer. Ninety out of the 96 are also present in the reptile parasite Entamoeba invadens, indicating lateral transfer before the species' divergence ∼50 MYA. We find only 2 introns, both shared with E. invadens. Thus, no intron gains have occurred in ∼50 Myr, implying a very low rate of intron gain of less than one gain per gene per ∼4.5 billion years. Nine other predicted introns are due to annotation errors reflecting apparent mistakes in the E. histolytica genome assembly. These results underscore the massive differences in intron gain rates through evolution.
Although common to all eukaryotic species, spliceosomal intron number varies tremendously across eukaryotes, from only 3 characterized introns in Giardia lambia to more than 8 introns per gene in vertebrates (compiled in Jeffares et al. 2006; Roy and Gilbert 2006). Patterns of intron gain and loss also show striking variations and often perplexing patterns (e.g., Bon et al. 2003; Perumal et al. 2005; Rodríguez-Trelles et al. 2006 provide an excellent recent review). For instance, intron-rich taxa often show very low rates of intron gain and/or high rates of loss (Seo et al. 2001; Rogozin et al. 2003; Roy et al. 2003; Cho et al. 2004; Raible et al. 2005; Roy and Gilbert 2005b; Stajich and Dietrich 2006). Some groups show high degrees of both intron loss and gain; others exhibit almost no loss or gain over very long periods of time (Seo et al. 2001; Roy et al. 2003; Edvardsen et al. 2004; Roy and Hartl 2006; Stajich and Dietrich 2006). However, attempts to estimate rates of intron loss and gain and to infer the relative importance of the 2 processes have been thwarted by lack of consensus over appropriate evolutionary assumptions, with different groups sometimes reaching very different conclusions from the same data set (Rogozin et al. 2003; Babenko et al. 2004; Nielsen et al. 2004; Qiu et al. 2004; Csurös 2005; Nguyen et al. 2005; Roy and Gilbert 2005a, 2005b).
Here we take a novel approach. We studied 96 genes from the moderately intron-dense parasitic amoeba Entamoeba histolytica (0.3 introns per gene on average) that were previously identified by phylogenetic analysis to represent lateral gene transfers (LGTs) from prokaryotes to Entamoeba (Loftus et al. 2005). Such genes were presumably intronless at the time of LGT, allowing confident inferences about intron gain. Sequence searches showed that 90/96 LGTs are present in the reptile parasite Entamoeba invadens and thus predate the E. histolytica–E. invadens divergence ∼50.5 ± 13.5 MYA based on conservative assumptions (see Methods).
Strikingly, 11/90 (11.5%) of these LGTs were predicted to have introns (table 1). However, investigation showed that 9/11 predicted introns reflected annotation errors. In 7 cases, comparison between the genome assembly and individual sequence reads identified assembly errors (in each case a single basepair indel relative to sequence reads). In each case, correction yielded a single long open reading frame (ORF) between the predicted start and stop codons, suggesting against intron presence. In 6 out of 7 cases, homologous E. invadens sequences were obtained; in each case, the corresponding sequence also appeared exonic (multiple of 3 bases and no stop codons).
Probable LGTs with Annotated Introns and the Conclusions Drawn from the Analyses Reported Here
Gene Name . | Putative gene function . | Conclusion . |
|---|---|---|
| 13.m00321 | Nitroreductase family protein | Supposed intron is exonic (fig. 1B) |
| 13.m00327 | Prolyl oligopeptidase family protein | Assembly indel |
| 192.m00077 | Geranylgeranyl pyrophosphate synthase | Assembly indel |
| 22.m00291 | Aspartate ammonia-lyase | Assembly indel |
| 289.m00068 | Amidohydrolase | Assembly indel |
| 328.m00056 | Alcohol dehydrogenase | Assembly indel/Alt. ATG (fig. 1A) |
| 3.m00589 | Glutamate synthase beta chain related | Assembly indel |
| 555.m00024 | D-glycerate dehydrogenase | Assembly indel |
| 78.m00151 | Lysophospholipase L2 | Assembly indel |
| 87.m00169 | 2-phosphosulfolactate phosphatase | Apparent intron insertion (fig. 2) |
| 8.m00343 | Deoxycytidine triphosphate deaminase | Present in Dictyostelium discoideum (fig. 1C) |
Gene Name . | Putative gene function . | Conclusion . |
|---|---|---|
| 13.m00321 | Nitroreductase family protein | Supposed intron is exonic (fig. 1B) |
| 13.m00327 | Prolyl oligopeptidase family protein | Assembly indel |
| 192.m00077 | Geranylgeranyl pyrophosphate synthase | Assembly indel |
| 22.m00291 | Aspartate ammonia-lyase | Assembly indel |
| 289.m00068 | Amidohydrolase | Assembly indel |
| 328.m00056 | Alcohol dehydrogenase | Assembly indel/Alt. ATG (fig. 1A) |
| 3.m00589 | Glutamate synthase beta chain related | Assembly indel |
| 555.m00024 | D-glycerate dehydrogenase | Assembly indel |
| 78.m00151 | Lysophospholipase L2 | Assembly indel |
| 87.m00169 | 2-phosphosulfolactate phosphatase | Apparent intron insertion (fig. 2) |
| 8.m00343 | Deoxycytidine triphosphate deaminase | Present in Dictyostelium discoideum (fig. 1C) |
Probable LGTs with Annotated Introns and the Conclusions Drawn from the Analyses Reported Here
Gene Name . | Putative gene function . | Conclusion . |
|---|---|---|
| 13.m00321 | Nitroreductase family protein | Supposed intron is exonic (fig. 1B) |
| 13.m00327 | Prolyl oligopeptidase family protein | Assembly indel |
| 192.m00077 | Geranylgeranyl pyrophosphate synthase | Assembly indel |
| 22.m00291 | Aspartate ammonia-lyase | Assembly indel |
| 289.m00068 | Amidohydrolase | Assembly indel |
| 328.m00056 | Alcohol dehydrogenase | Assembly indel/Alt. ATG (fig. 1A) |
| 3.m00589 | Glutamate synthase beta chain related | Assembly indel |
| 555.m00024 | D-glycerate dehydrogenase | Assembly indel |
| 78.m00151 | Lysophospholipase L2 | Assembly indel |
| 87.m00169 | 2-phosphosulfolactate phosphatase | Apparent intron insertion (fig. 2) |
| 8.m00343 | Deoxycytidine triphosphate deaminase | Present in Dictyostelium discoideum (fig. 1C) |
Gene Name . | Putative gene function . | Conclusion . |
|---|---|---|
| 13.m00321 | Nitroreductase family protein | Supposed intron is exonic (fig. 1B) |
| 13.m00327 | Prolyl oligopeptidase family protein | Assembly indel |
| 192.m00077 | Geranylgeranyl pyrophosphate synthase | Assembly indel |
| 22.m00291 | Aspartate ammonia-lyase | Assembly indel |
| 289.m00068 | Amidohydrolase | Assembly indel |
| 328.m00056 | Alcohol dehydrogenase | Assembly indel/Alt. ATG (fig. 1A) |
| 3.m00589 | Glutamate synthase beta chain related | Assembly indel |
| 555.m00024 | D-glycerate dehydrogenase | Assembly indel |
| 78.m00151 | Lysophospholipase L2 | Assembly indel |
| 87.m00169 | 2-phosphosulfolactate phosphatase | Apparent intron insertion (fig. 2) |
| 8.m00343 | Deoxycytidine triphosphate deaminase | Present in Dictyostelium discoideum (fig. 1C) |
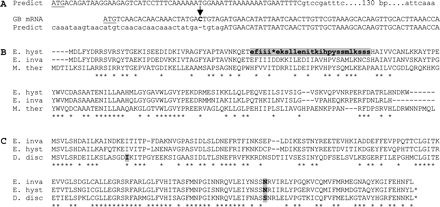
In another case, a homologous E. histolytica mRNA from GenBank (AAA81906.1) had a single basepair indel relative to the predicted gene (which fell within the predicted intron) and an intronless gene structure spanning the 3′ of the predicted intron terminus and the downstream exon (fig. 1A). In yet another case, amino acid–level similarity to homologous sequences from bacteria and E. invadens continues through the intron (fig. 1B), suggesting that the predicted intronic sequence is in fact exonic. Interestingly, the E. histolytica sequence but not the corresponding E. invadens sequence contains a single in-frame stop codon, which is confirmed by individual sequencing reads. Whether this apparent gene truncation occurred in natural populations or in the lab is unknown.
Three examples of LGTs predicted to contain introns. (A) 5′ alignment of predicted Entamoeba histolytica gene 328.00056 (“Predict”) and an E. histolytica GB mRNA (GenBank accession number AAA81906.1). The GB mRNA contains an extra cytosine (arrow) relative to the predicted gene, uses an alternative start codon (underlined), and does not reflect a splicing event. Upper/lowercase indicates exonic/intronic sequence. (B) E. histolytica gene 13.m00321 and homologs. The supposedly intronic sequence (lower case bold) shows strong coding-level sequence similarity to a bacterial homolog (43% amino acid identity; Morella thermoacetica gene, GenBank accession number ABC19526.1) and to the apparent Entamoeba invadens homolog (57% identity), suggesting that it is a coding sequence, not an intron. (C) E. histolytica gene 8.m00343 and homologs from D. discoideum (GenBank accession number XP_629020) and E. invadens. Gray boxes indicate intron positions.
Thus, only 2 genes showed evidence of intron presence. One has a close homolog in Dictyostelium discoideum, which shares the intron (fig. 1C). The D. discoideum–Entamoeba divergence represents a deep split within amoebozoa, thus this gene is either a very old LGT or is not an LGT at all (fig. 1C). This leaves a single intron in the 2-phosphosulfolactate phosphatase gene. The predicted 226-codon gene contains a single 53-bp intron with 79.2% AT content. The intronic sequence is not a multiple of 3 bases and contains 6 stop codons falling in all 3 frames and is thus almost certainly an intron (fig. 2A). Both upstream and downstream exons show coherent homology to bacterial homologs, suggesting that the intron was inserted into previously contiguous coding sequence (fig. 2B). The gene is absent from D. discoideum, available Acanthamoeba castellanii genomic sequence, and other eukaryotes in GenBank, supporting its lateral transfer. However, the intron is shared with E. invadens (fig. 2A and B), and thus the intron gain predates the E. histolytica–E. invadens divergence. This intron represents the first reported case of intron gain in an amoeba.
Apparent intron insertion in the Entamoeba histolytica 2-phosphosulfolactate phosphatase (87.m00169). (A) Intron and flanking exonic sequence for 4 Entamoeba species. Upper/lowercase indicates exonic/intronic sequence. Stop codons in the frame of the upstream and downstream coding sequences are shown (bold). For Entamoeba terrapinae, only the downstream exon and part of the intron sequence was available. E. hist, E. mosh, E. inva, and E. terr indicate E. histolytica, E. moshkovskii, E. invadens, and E. terrapinae, respectively. (B) Alignment with homologous bacterial genes (ClustalW, default parameters). Asterisks indicate positions at which there is identity between a bacterial gene and an Entamoeba gene. The gray box indicates the intron position. T. teng, T. meri, and C. perf indicate genes from Therobacter tengcongensis (GenBank accession number AAM25151.1), Thermotoga maritima (GenBank accession number AAD35879.1), and Clostridium perfringens (GenBank accession number BAB82262.1), respectively.
We found no intron gains in 90 LGTs in ∼50.5 ± 13.5 Myr, suggesting a rate of intron gain of less than 0.00022 ± 0.00006 intron gains per gene per Myr or one gain per gene per 4.5 ± 1.2 billion years. Importantly, this conclusion holds even if some of the genes are not actual LGTs because regardless of the genes' origin, no intron gains are found in ∼50 Myr. It is unlikely that many gained introns have been subsequently lost because even assuming the highest loss rates ever estimated (∼2.2 × 10−9 per year; Roy and Gilbert 2005b) only 10% of introns are expected to be lost over 50 Myr. This low rate of gain is not consistent with high intron numbers in diverse modern eukaryotes (e.g., 37.8 billion years would be required to reach the 8.4 introns per gene found in Homo sapiens) or with the apparently high intron numbers already present relatively early in eukaryotic evolution (Csurös 2005; Nguyen et al. 2005; Roy and Gilbert 2005a), implying that rates of intron creation have varied significantly through evolution (see Roy and Gilbert 2005b for a more thorough discussion).
Genome-wide studies of closely related species indicate very low rates of intron gain of less than one per gene per 1.5 billion years in animals, fungi, plants, apicomplexans (Roy et al. 2003; Coghlan and Wolfe 2004; Nielsen et al. 2004; Lin et al. 2006; Roy and Hartl 2006; Stajich and Dietrich 2006), and now amoebozoa. Only a single genome-wide study, in A. thaliana, shows a higher rate, though as the authors of that manuscript concede some reported gains may in fact represent losses, and their data warrant further study (Knowles and McLysaght 2006). These modern rates are too low to explain modern and estimated ancestral intron densities (Fedorov et al. 2002; Csurös 2005; Roy and Gilbert 2005b), implying much higher rates of intron creation during some earlier period(s) of evolution (Fedorov et al. 2003). To explain this pattern, we will need to better understand the evolutionary forces governing intron gain and loss.
Methods
We downloaded the E. histolytica genome gbk files (version 1) from National Center for Biotechnology Information (NCBI) (http://www.ncbi.nlm.nih.gov/) and extracted exon and intron sequences. For each of the 11 LGT genes that were predicted to contain introns, we performed BlastN searches of the corresponding genomic region against all E. histolytica reads in the NCBI Trace Archive and compared the assembled sequence with the best hit. For cases in which reads and assembly agreed, we performed TBlastN searches at NCBI for corresponding Entamoeba and A. castellanii sequences and searched NCBI and the D. discoideum genome project for corresponding sequences from other amoebae. TBlastN searches against available genome sequence from other Entamoeba species were performed online (http://www.sanger.ac.uk/Projects/Comp_Entamoeba/). A TBlastN search of the E. histolytica 2-phosphosulfolactate phosphatase sequence against all eukaryotic sequences in GenBank yielded no non-Entamoeba sequences. To estimate dS between E. invadens–E. histolytica, we downloaded available E. invadens mRNAs in GenBank and excluded those not beginning with “ATG” or ending with a stop codon. Reciprocal BlastP searches against the E. histolytica predicted proteome identified 10 putative ortholog pairs with strong amino acid–sequence identity (>40%). Sequences were aligned in ClustalX using default parameters, and average dS and confidence intervals (CI) across genes were calculated using PAUP*4.0 using a general time reversible substitution model estimated from the data set (Lanave et al. 1984). Although mutation rates for amoebae have not been estimated, conservatively assuming the highest estimates of which we are aware for any unicellular eukaryote (around 5 × 10−9 per year, in Plasmodium; Castillo-Davis et al. 2004; Tanabe et al. 2004; Neafsey et al. 2005) yields an estimate of 50.5 ± 13.5 Myr.
Martin Embley, Associate Editor
We thank Warwick Allen for help formatting the figures. MI was supported by funds from Fundacion Caixa Galicia.
References
Babenko VN, Rogozin IB, Mekhedov SL, Koonin EV.
Bon E, Casaregola S, Blandin G, et al. (11 co-authors).
Castillo-Davis CI, Bedford TB, Hartl DL.
Cho S, Jin SW, Cohen A, Ellis RE.
Coghlan A, Wolfe KH.
Csurös M.
Edvardsen RB, Lerat E, Maeland AD, Flat M, Tewari R, Jensen MF, Lehrach H, Reinhardt R, Seo HC, Chourrout D.
Fedorov A, Merican AF, Gilbert W.
Knowles DG, McLysaght A.
Lanave C, Preparata G, Saccone C, Serio G.
Lin H, Zhu W, Silva JC, Gu X, Buell CR.
Loftus B, Anderson I, Davies R, et al. (54 co-authors).
Neafsey DE, Hartl DL, Berriman M.
Nguyen HD, Yoshihama M, Kenmochi N.
Nielsen CB, Friedman B, Birren B, Burge CB, Galagan JE.
Perumal BS, Sakharhar KR, Chow VT, Pandjassarame K, Sakharkar MK.
Qiu WG, Schisler N, Stoltzfus A.
Raible F, Tessmar-Raible K, Osoegawa K, et al. (12 co-authors).
Rodríguez-Trelles F, Tarrío R, Ayala FJ.
Rogozin IB, Wolf YI, Sorokin AV, Mirkin BG, Koonin EV.
Roy SW, Fedorov A, Gilbert W.
Roy SW, Gilbert W.
Roy SW, Gilbert W.
Roy SW, Hartl DL.
Seo H-C, Kube M, Edvardsen RB, et al. (11 co-authors).
Stajich JE, Dietrich FS.