-
PDF
- Split View
-
Views
-
Cite
Cite
Rogier B. Donker, Jean-François Mouillet, D.Michael Nelson, Yoel Sadovsky, The expression of Argonaute2 and related microRNA biogenesis proteins in normal and hypoxic trophoblasts, Molecular Human Reproduction, Volume 13, Issue 4, April 2007, Pages 273–279, https://doi.org/10.1093/molehr/gam006
Close - Share Icon Share
Abstract
Endogenous microRNAs (miRNAs) post-transcriptionally regulate mRNA and protein expression during tissue development and function. Whereas adaptation to environmental insults are tightly regulated in human tissues, the role of miRNAs and miRNA biogenesis proteins in this context is inadequately explored. We sought to analyse the expression of the key RNAi enzyme Argonaute2 (Ago2) and other miRNA biogenesis proteins in human trophoblasts during differentiation and in hypoxic environment. Using an in vitro analysis of primary term human trophoblasts, we identified the expression of the core miRNA biogenesis proteins in human villous trophoblasts, with expression levels unaffected by cellular differentiation. We found that the miRNA biosynthetic pathway was functional and produced miRNAs, with miR-93 up-regulated and miR-424 down-regulated in hypoxic environment. In contrast, hypoxia did not alter the expression of key miRNA machinery proteins. The pivotal miRNA processing enzyme Ago2, along with its interacting protein DP103, were expressed in normal placentas as well as in placentas from pregnancies complicated by placental hypoperfusion that resulted in fetal growth restriction. Ago2 and DP103 co-immunoprecipitated, and did not limit trophoblast response to hypoxic stress. We concluded that the core miRNA machinery proteins are expressed and functional in human trophoblasts. The influence of hypoxia on the expression of a subset of placental miRNA species is unlikely to reflect altered expression of key miRNA biogenesis proteins.
Introduction
MicroRNAs (miRNAs) are endogenous, non-coding, short RNAs that post-transcriptionally regulate gene expression through sequence-specific base-pairing with target mRNAs. Hundreds of miRNAs, which are predicted to comprise ∼2% of known genes, have been identified, with a fraction of them demonstrating differential expression during development and across diverse tissues (Bentwich et al., 2005). In animals, miRNAs usually inhibit gene expression through partially complementary elements in the 3′ untranslated regions of their target mRNAs. Mammalian miRNAs repress protein target expression through inhibition of translational initiation or elongation by ribosomes (Pillai et al., 2005; Petersen et al., 2006) or by diverting mRNA targets away from the translational machinery into cytoplasmic processing bodies (P- or GW-bodies) (Jakymiw et al., 2005; Liu et al., 2005b). In addition, miRNAs are capable of degrading their mRNA targets (Jing et al., 2005; Rehwinkel et al., 2006). A single miRNA may target numerous mRNAs that are involved in diverse cellular functions, including development, cellular differentiation, apoptosis, oncogenesis, metabolism and endocrine function (Alvarez-Garcia and Miska, 2005; Esquela-Kerscher and Slack, 2006; Krutzfeldt and Stoffel, 2006). Furthermore, miRNAs may contribute to cell lineage determination and maintenance of cell-type specific transcriptome profiles (Sood et al., 2006). It is presently estimated that miRNA can modify the expression of ∼30% of protein-coding transcripts, implicating miRNAs as one of the largest classes of gene regulators (Lim et al., 2005).
Biogenesis of miRNAs begins with RNA polymerase II transcription of miRNA genes into several hundred-nucleotide long primary miRNAs (pri-miRNAs) (Bartel, 2004). Nuclear pri-miRNAs are processed into ∼70 nucleotide precursor miRNAs (pre-miRNAs) by the microprocessor complex that includes the RNase III endonuclease Drosha together with the double-stranded RNA-binding domain (dsRBD) containing protein Pasha (DGCR8) (Gregory et al., 2004; Han et al., 2004). Folded into imperfect stem-loop structures, pre-miRNAs are exported from the nucleus into the cytoplasm by the nuclear transmembrane Ran GTP-dependent transporter Exportin-5 (Exp5) (Lund et al., 2004). Cytoplasmic miRNA is processed by a second RNase III endonuclease, Dicer, to liberate 20–22 nucleotide miRNA duplexes, of which the mature miRNA strands are preferentially retained in the RNA-induced silencing complex (RISC) (Chendrimada et al., 2005; Gregory et al., 2005). The key components of RISC are Argonaute (Ago) proteins (Meister and Tuschl, 2004), which in humans include four miRNA-binding members. Only Ago2 is associated with the required catalytic activity for mRNA cleavage (Liu et al., 2004; Meister et al., 2004). Human Ago2 is associated with other RISC proteins, which include the DEAD-box helicase protein DP103 (Gemin3 and Ddx20) (Hutvagner and Zamore, 2002; Mourelatos et al., 2002; Yan et al., 2003; Shpargel and Matera, 2005), Gemin4, MOV10, PRMT5 (Meister et al., 2005) and RCK/p54 (Chu and Rana, 2006), as well as the P-body components GW182, dcp1a and dcp2 (Jakymiw et al., 2005; Liu et al., 2005a,b).
Intact differentiation and adaptive responses of the placenta are essential for embryonic development. Differentiated trophoblasts at the surface of human placental villi play a pivotal role in gas exchange, nutrition, waste removal, endocrine function and immunological support for the developing fetus. Early placental development occurs in an environment of relative hypoxia, which promotes trophoblast invasion and angiogenesis (Genbacev et al., 1997). In contrast, placental hypoxia beyond the first trimester has been associated with villous trophoblast injury, and impacts the trophoblastic transcriptome, differentiation, metabolic function, cell survival and apoptosis, which likely contribute to the pathophysiology of fetal growth restriction (FGR) or pre-eclampsia (Rodesch et al., 1992; Levy et al., 2000; Roh et al., 2005; Vaiman et al., 2005). Whereas several species of miRNA were previously detected in the human placenta (Barad et al., 2004), none has been associated with normal placental function or disease. Moreover, the expression of miRNA biogenesis enzymes in the human placenta has not been previously examined. miRNA are involved in cellular stress (Bhattacharyya et al., 2006; Sunkar et al., 2006), yet the expression of miRNA machinery proteins in hypoxia has not been previously examined, we sought to elucidate the expression of Ago2 and related core miRNA biosynthetic proteins in villous trophoblast and their regulation in a hypoxic environment. Our data indicate that Ago2 and pivotal miRNA machinery proteins are expressed in human villous trophoblast. Although hypoxia modulates the expression of discrete miRNA species, the expression and interaction of Ago2 and its related miRNA biogenesis elements is stable in trophoblasts under standard or hypoxic conditions.
Materials and methods
Tissue procurement
The study was approved by the Institutional Review Board of Washington University School of Medicine in St. Louis. Human placentas for isolation and culture of trophoblasts were obtained from normal singleton pregnancies after vaginal or abdominal delivery. All pregnancies were completed between 37 and 40 weeks by reliable obstetrical dates. Placentas were excluded if the pregnancy was complicated by maternal diabetes, hypertensive disorder, tobacco or substance abuse, infection, prolonged rupture of membranes, multi-fetal gestation, meconium-stained amniotic fluid, placental abruption, previa, accreta or placentas harbouring visible abnormalities, known fetal chromosomal abnormalities or structural anomaly. Placental villous biopsies were obtained from dizygotic twin pairs at 32–38-week gestation where one fetus exhibited normal growth and the other FGR, attributed to placental hypoperfusion based on Doppler flow studies and placental pathology as we previously described (Roh et al., 2005). Placental villous biopsies, obtained immediately after delivery, were ∼0.5 cm3 and were taken midway between the medial region to lateral edge and midway from the basal plate to the chorionic plate of the placenta, as we previously described (Wyatt et al., 2005).
Cell culture
Primary human cytotrophoblasts were prepared from normal term placentas, using the trypsin-deoxyribonuclease-Dispase/Percoll method as described by Kliman with previously published modifications (Kliman et al., 1986; Nelson et al., 1999). Cultures were plated at a density of 350 000 cells cm−2 and maintained in Dulbecco's modified Eagle's medium (DMEM, Sigma, St. Louis, MO, USA) containing 10% fetal bovine serum (FBS, Hyclone, Logan, UT, USA) and antibiotics as described (Nelson et al., 1999) at 37°C in a 5% carbon dioxide-air atmosphere. Where indicated, cells were cultured in Ham's/Waymouth (H/W) medium containing 10% FBS and antibiotics. After 4 h, designed to allow cell attachment, the culture plates were allocated to standard or hypoxic growth conditions. Cultures at hypoxic conditions were maintained in an incubator (Thermo Electron, Marietta, OH, USA) that provided hypoxic atmosphere, defined as < 1% O2 (5% CO2, 10% H2, and 85% N2) with continuous computerized monitoring of atmospheric oxygen using a sensor connected to a data acquisition module and displayed as a strip chart on a PC laptop. All media were pre-equilibrated to the gas mixture prior to addition to the culture plate and refreshed every 24 h. Cultures in standard or hypoxic conditions continued for 48 h, or as indicated in the figures. Hormonal differentiation was routinely monitored by medium hCG levels, performed using enzyme-linked immunosorbent assay (DRG International, Mountainside, NJ, USA), showing a characteristic increase in medium hCG as trophoblasts differentiate, with attenuation of this process in hypoxic cells (Nelson et al., 1999; Humphrey et al., 2005). JEG3 human choriocarcinoma cells were grown and maintained in minimal essential medium (Hyclone) containing 10% FBS and antibiotics. Cells were passaged when confluent using a 0.05%Trypsin/0.02%EDTA solution.
Reverse transcription and quantitative PCR
Total cellular RNA was purified from cells and tissue biopsies using TriReagent (Sigma) or the mirVana™ miRNA isolation kit (Ambion, Austin, TX, USA), according to the manufacturer's instructions. Purified RNA was treated for 20 min at 37°C with DNAse I (TURBO DNA-free, Ambion) to remove contaminating DNA. RT was performed using TaqMan Reverse Transcription Reagents Kit (Applied Biosystems, Foster City, CA, USA) in a 50-µl reaction mix that contained 1–1.5 µg total RNA, 1 × RT buffer, 5.5 mM MgCl2, deoxy-NTP mixture (500 µM of each dNTP), 2.5 µM random hexamers, 0.4 U µl−1 RNase Inhibitor and 1.25 U µl−1 MultiScribe reverse transcriptase at 25°C for 10 min, 48°C for 30 min and 95°C for 5 min. The RT product (3 µl) was used for real-time PCR as described previously (Schaiff et al., 2005). Primer sequences used were (5′ → 3′, with accession number): Ago2 F GCACCGGCAGGAGATCATAC, R AGACACCGTCGCGGTAGAA (NM_012154); Dicer F TTCCTCACCAATGGGTCCTTT, R GCTTCAACGAGTTCAACCTGT (NM_030621); DP103 F GGAAGGCTTAGAGTGTCATGTC, R TGAGTTGCTTAATTCTGCCAGG (NM_007204); Drosha F TGTCACAGAATGTCGTTCCAC, R GGGCCTAAAGGATGGTGCT (BC054003), Exp5 F TAGGCATGGCGATGGATCAAG, R GGGGACACAGATAGGACACTTT (AB003117), β-hCG F CTACTGCCCCACCATGACC, R GCAGAGTGCACATTGACAGCT (BC126460); p53 F CCTATGGAAACTACTTCCTGAAAACAA, R ACAGCATCAAATCATCCATTGC (NM_000546); VEGF F CTGGCGCTGAGCCTCTCTA, R CCGGTGTCCTCATCCCTGTA (NM_003376); YWHAZ F ACTTTTGGTACATTGTGGCTTCAA, R CCGCCAGGACAAACCAGTAT (NM_145690); 18S F CGCCGCTAGAGGTGAAATTCT, R CGAACCTCCGACTTTCGTTCT (M10098). All primer sequences were BLAST-checked for specificity. Dissociation curves were run on all reactions to ensure amplification of a single product with the appropriate melting temperature. Control samples of H2O were included in each experiment. Samples were normalized to parallel reactions using primers specific for the trophoblast housekeeping gene YWHAZ [Figure 1B and C; Wyatt et al. (2005)] or for 18S rRNA. The fold increase relative to control samples was determined by the 2-ΔΔCT method (Livak and Schmittgen, 2001). Reverse transcription and quatitatives PCR (RT–qPCR) for miRNA was performed using the mirVana™ qRT–PCR miRNA Detection Kit (Ambion) with Klentaq-LA in RT reactions and primers specific for hsa-miR-93, hsa-miR-424 (Ambion), as well as 5S rRNA, used for miRNA normalization.
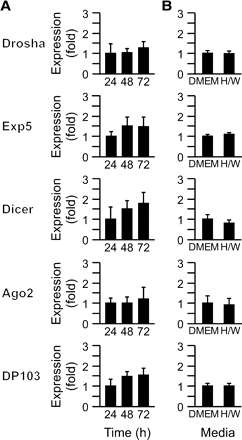
The expression of miRNA biogenesis pathways during differentiation of human trophoblasts. (A) The expression of core microRNA (miRNA) machinery transcripts in human trophoblasts, which differentiate into syncytiotrophoblast during a 72-h period. Expression was determined in duplicate as described in Materials and methods (a representative experiment, n = 5). Expression levels are expressed as fold overexpression at 24 h, defined as one. (B) The expression of miRNA machinery transcripts in human trophoblasts cultured for 72 h in either Dulbecco's modified Eagle's medium (DMEM), which promotes differentiation, or H/W medium, which hinders differentiation (a representative experiment, n = 3). Expression levels in DMEM were defined as one. None of the differences were significant.
miRNA northern blotting
For confirmation of miRNA levels, 20 µg of total RNA was denatured at 65°C for 10 min and subsequently resolved using 7 M urea/15% polyacrylamide gel electrophoresis (PAGE). The gel was stained using SYBR-gold (Invitrogen/Molecular Probes, Eugene, OR, USA) and RNA was transferred to a nylon Hybond N + membrane (GE Healthcare, Piscataway, NJ, USA). Hybridization was performed using hsa-miR-93 and hsa-miR-424 specific antisense StarFire probes (Integrated DNA Technologies, Coralville IA, USA), which were labelled with [α-32P]-dATP. After washing, membranes were exposed for 16–24 h to Kodak film for visualization.
Antibodies, protein expression using western immunoblotting and immunofluorescence
For western analysis, cells were lysed in 175 µl cold lysis buffer [1% Nonidet P-40, 0.5% deoxycholate and 0.1% sodium dodecyl sulphate (SDS) in phosphate-buffered saline (PBS)] containing a protease inhibitor cocktail (all from Sigma). The plates were scraped, and the lysate sonicated three times for 1 s each on ice and centrifuged. Proteins (10–15 µg) were resolved using 10% SDS–PAGE at 90 V, 4°C for 3 h. The proteins were transferred to polyvinylidene difluoride membranes (Immobilon-P, Millipore, Bedford, MA, USA) at 150 mA, 4°C for 18 h. The blot was blocked by incubation in 10% non-fat dry milk in PBST (0.01 M PBS with 0.05% Tween-20) for 2 h. The membrane was then incubated for 1.5 h at room temperature in 10% non-fat dry milk in PBST with either polyclonal rabbit anti-human Ago2 antibody (1:750 dilution; 07–590; Upstate USA, Charlottesville, VA, USA) or polyclonal rabbit anti-human DP103 antibody (0.9 mg ml−1, 1:900 dilution). For production of anti-DP103 polyclonal antibody, rabbits were injected with a MAP-linked peptide spanning aa 594–612 of DP103 (Research Genetics, Huntsville, AL, USA). The antibody was affinity-purified from the antisera using AminoLink Plus kit (Pierce, Rockford, IL, USA). Control membranes were also incubated with polyclonal goat anti-human actin antibody (1:900 dilution; sc-1616, Santa Cruz Biotech, Santa Cruz, CA, USA). Either horse-radish peroxidase-conjugated goat anti-rabbit immunoglobulin G (IgG) or rabbit anti-goat IgG antibody (1:2500; Santa Cruz) was used for secondary antibodies. The blots were washed and processed for chemiluminescence using enhanced chemiluminescence (PerkinElmer, Wellesley, MA, USA) and densitometrically quantified with Epichemi-3 software (UVP BioImaging, Upland, CA, USA).
For immunofluorescent staining, 5 mm thickness sections were cut from paraffin-embedded biopsies of placental villi and mounted onto glass slides, deparaffinized and subjected to heat-induced antigen retrieval. Sections were blocked with 10% bovine serum albumin, 0.1% Tween-20 in 0.01 M PBS at room temperature for 2 h. Subsequently, sections were incubated at room temperature with anti-human Ago2 or anti-human DP103 for 1.5 h. After washing, sections were incubated for 1 h with an AlexaFluor 488-conjugated F(ab′)2 goat anti-rabbit antibody (2 mg ml−1, Invitrogen/Molecular Probes). Nuclei were stained using TOPRO-3 iodide (Invitrogen/Molecular Probes).
Co-immunoprecipitation
Primary trophoblasts were cultured for 48 h in standard or hypoxic conditions. Cells were lysed using non-denaturing lysis buffer (1% Triton X-100, 50 mM Tris-HCl, 300 mM NaCl and 5 mM EDTA) containing a protease inhibitor cocktail (Sigma). Lysates were cleared by centrifugation and an aliquot was examined for protein concentration (BioRad). Lysates were incubated with 7.5 µg anti-human Ago2 or non-specific donkey anti-mouse antibody while rotating at 4°C for 14 h. Subsequently, 50 µl of 50% protein-A bead slurry was added to each sample for 2 h. The beads were collected by centrifugation and washed three times in a buffer containing 0.1% Triton X-100, 50 mM Tris-HCl, 300 mM NaCl and 5 mM EDTA. The beads were then suspended in 80 µl elution buffer containing 0.8% SDS and 2% 2-mercaptoethanol and heated at 100°C for 8 min to release bead-bound proteins. The supernatant was collected by centrifugation, and 20 µl was used for western immunoblotting, performed as described above.
DNA plasmids and transfection
JEG3 cells (10 000 cells cm−2) were plated 16 h before transfection. The modified calcium–phosphate co-precipitation method was used for transfection, as we previously described (Yan et al., 2003), using 3 µg of DNA plasmid expressing human Ago2 or enhanced green fluorescent protein (EGFP) (pIRESneo-FLAG/HA-Ago2, pIRESneo-FLAG/HA EGFP, Addgene, Cambridge, MA, USA). Cells were washed 18 h after transfection and cultured in standard or hypoxic conditions for 30 h. Subsequently, RNA and protein were harvested.
Statistics
Data were expressed as mean ± SD. qPCR data were computed as fold using geometric means and normalized to the mean value of the control sample in each paradigm, defined as one. All experiments were repeated at least three times in duplicates, and comparisons were analysed using Student's t-test (Primer of Biostatistics, McGraw-Hill NY, NY, USA). Significance was defined as P < 0.05.
Results
Expression of miRNA biogenesis proteins in human trophoblasts
We initially sought to determine the expression of key miRNA biogenesis proteins in primary term human trophoblasts. Using RT–qPCR, we found that transcripts for Drosha, Exp5, Dicer, Ago2 and DP103 were expressed in third-trimester human villous trophoblasts, cultured under standard conditions (Figure 1A). The expression level of these transcripts was not significantly different over 72 h in culture, a time period characterized by cytotrophoblast differentiation into multinucleated syncytia (Nelson et al., 1999). To ensure that the expression of miRNA biogenesis elements was not influenced by differentiation of trophoblasts, we also examined transcript expression in cells cultured in either standard DMEM or in H/W medium, which hinders trophoblast differentiation (Douglas and King, 1990). As shown in Figure 1B, mRNA levels for all five transcripts were similar between the two culture conditions, supporting the uniform expression level of pivotal miRNA biogenesis elements in cytotrophoblasts and syncytiotrophoblasts.
Regulation of miRNA species in hypoxic human trophoblasts
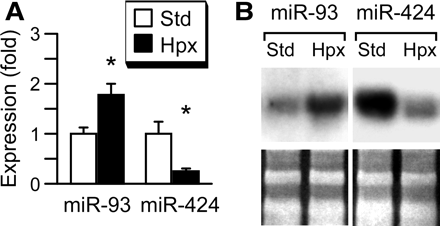
To assess the functionality of the miRNA biogenesis proteins under hypoxic conditions, we examined the expression of two miRNA species that we recently identified in human primary trophoblasts using miRNA microarray, miR-93 and miR-424 (JFM and YS, unpublished observation). The microarray screen, performed using GenoExplorer Biochips (Genosensor, Tempe, AZ, USA), demonstrated an increase in miR-93 expression (2.4–4.4-fold, n = 2) and a decrease in miR-424 expression (0.16–0.5-fold, n = 2) in hypoxic trophoblasts, compared to control. As shown in Figure 2, both quantitative PCR and northern analysis confirmed the expression of the two miRNA species in human trophoblasts. Importantly, as predicted by our miRNA microarray assay, we found that the expression of miR-93 was up-regulated in hypoxic trophoblasts, whereas the expression of miR-424 was down-regulated in hypoxic trophoblasts. These results support the notion that miRNA biogenesis pathways are active even in hypoxic trophoblasts and that hypoxia regulates the expression level of miRNA in these cells.
Hypoxia regulates the expression of miR-93 and miR-424 in primary human trophoblast. (A) Reverse transcription and quantitative PCR, with expression in standard conditions defined as one (a representative experiment, n = 4). * denotes P < 0.05. (B) Northern blot, with the lower panel depicting stable expression of 5S/tRNA, detected with SYBR-gold and used for loading control (a representative experiment, n = 3).
Constant expression of Ago2 and miRNA machinery enzymes in hypoxic trophoblasts
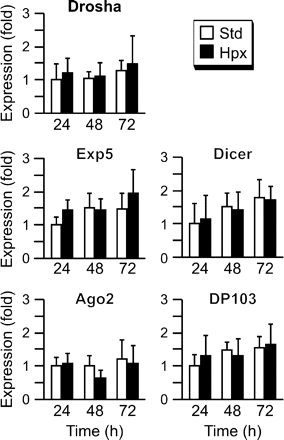
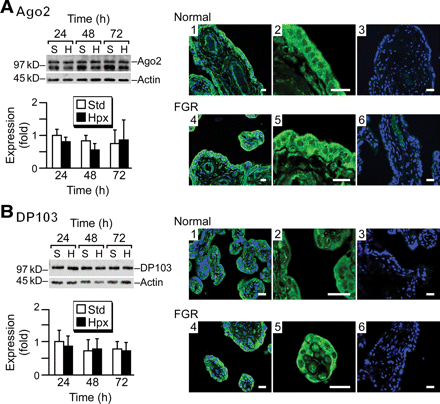
As hypoxic stress regulates the expression of discrete miRNA species in trophoblasts, we sought to determine if hypoxia alters the expression of key miRNA biogenesis components. To test for this possibility, we examined the expression of Drosha, Exp5, Dicer, Ago2 and DP103 in trophoblasts cultured in standard or hypoxic conditions. We found that hypoxia had an insignificant influence on the expression of the five transcripts during the culture period (Figure 3). As Ago2 is a pivotal component of RISC and therefore governs the final steps of mature miRNA function, we furthered our studies and focused on Ago2 expression in trophoblasts. We also examined the expression of the Ago2–interacting protein DP103. Using western immunoblotting we detected Ago2 and DP103 proteins in cultured trophoblasts (Figure 4), with stable expression during differentiation in vitro and in response to hypoxia. In addition, we used immunofluorescence to confirm the expression pattern of Ago2 and DP103 in villous trophoblasts in vivo (Figure 4A and B). Expression was localized primarily to the cytoplasm, consistent with the role of Ago2 in assembly of cytoplasmic RISC. Notably, the expression pattern of Ago2 and DP103 was similar between placental villi derived from the two placentas of dizygotic twin pregnancies in which one fetus exhibited placental hypoperfusion and FGR.
The influence of hypoxia on trophoblast expression of key miRNA machinery enzymes. The expression of five miRNA machinery transcripts in human trophoblasts was determined in duplicate and expressed as fold overexpression at 24 h in standard conditions, defined as one (a representative experiment, n = 5). None of the differences were significant.
The expression of Argonaute2 (Ago2) and DP103 proteins in hypoxic trophoblasts. The expression of Ago2 (A) and DP103 (B) in trophoblasts. Protein expression was determined by western blot (left panels) in trophoblasts cultured for 72 h in standard (S) or hypoxic (H) conditions. The results were quantified using densitometry, with expression at 24 h in standard conditions, defined as one. None of the differences were statistically significant. Note that a non-specific band was detected below the Ago2 band in all paradigms. This band has been described by the manufacturer and disappeared by immunoprecipitation (Figure 5). Protein expression was also determined by immunofluorescence (right panels) of placental villi from normal (1–3) or fetal growth restriction (FGR) (4–6) placentas from dizygotic twins (representative experiments of three pairs of twins). Panels 1 and 4 depict staining for Ago2 or DP103 (green) with TOPRO-3 staining (blue) for nuclei. Panels 2 and 5 are a higher magnification for Ago2 or DP103, and panels 3 and 6 are controls in which the primary antibody was omitted (bar = 20 µm).
Ago2 physically interacts with DP103 within RISC complexes (Hutvagner and Zamore, 2002; Mourelatos et al., 2002); therefore, we expanded our studies and examined Ago2-DP103 interaction in primary trophoblasts in standard and hypoxic conditions. Using co-immunoprecipitation, we found that Ago2 and DP103 form a complex in trophoblasts (Figure 5). We further found that a comparable amount of DP103 co-immunoprecipitated with Ago2 in standard or hypoxic culture conditions, indicating that hypoxia does not alter Ago2–DP103 interaction in trophoblasts (Figure 5). Together, these data support the notion that the expression level of the RISC proteins Ago2 and DP103 and the stability of Ago2–DP103 complexes are not modulated by hypoxia in trophoblasts.
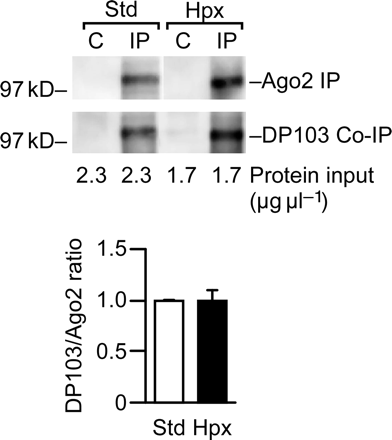
DP103 forms a complex with Ago2 in trophoblasts cultured in standard or hypoxic conditions. Protein complexes were precipitated using protein-A bead bound anti-Ago2 antibodies and detected using anti-DP103. Co-immunoprecipitation (a representative experiment, n = 3) was performed as described in Materials and methods, with the total amount of protein input indicated below the gel. The lower graph depicts the ratio between DP103 and Ago2 in standard or hypoxic conditions, with band intensity quantified using densitometry.
The expression level of Ago2 does not alter the expression of hypoxic trophoblast-related transcripts
Altered expression of a diverse set of trophoblast genes characterizes trophoblast response to hypoxia and mediates trophoblast response to hypoxic injury (Roh et al., 2005). We previously determined that the expression of trophoblast vascular endothelial growth factor (VEGF) is markedly enhanced during hypoxia. In contrast, the expression of β-hCG is dramatically reduced in hypoxic trophoblasts (Nelson et al., 1999). The expression of miRNA species in trophoblasts is regulated by hypoxia, yet the expression of miRNA biogenesis enzymes such as Ago2 is unchanged; therefore, we surmised that Ago2 is not rate-limiting for the expression of hypoxia-induced trophoblast transcripts. To examine this possibility, we used the JEG3 placental cell line. We initially confirmed that JEG3 cells exhibit hypoxia-induced change in transcript expression in a similar manner to primary trophoblasts (Figure 6A). In addition, these cells express Ago2 endogenously (Figure 6B and data not shown) and are suitable for overexpression of Ago2, as demonstrated using RT–qPCR and western analysis (Figure 6B). Importantly, overexpression of Ago2 did not alter the expression of a selected set of hypoxic trophoblast signature transcripts in cells cultured under standard or hypoxic conditions (Figure 6C). Together, these data support the notion that cellular Ago2 level is not limiting for trophoblast response to hypoxia.
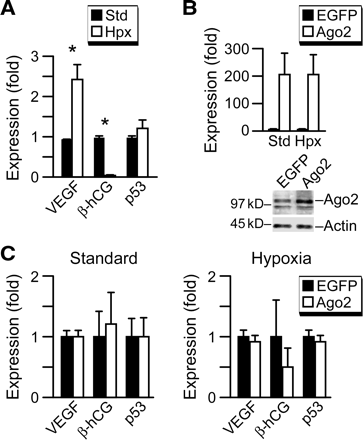
Ago2 expression level is not a limiting factor in trophoblast response to hypoxia. (A) The influence of hypoxia on vascular endothelial growth factor (VEGF), β-hCG and p53 transcript levels in JEG3 cells. Transcript levels were determined in duplicates as described in materials and methods (representative experiments, n = 3). Expression levels are expressed as fold over expression at 24 h, defined as one. *Denotes P < 0.05. (B) Transient over-transfection of Ago2 effectively increased Ago2 mRNA and protein expression levels in JEG3 cells cultured in standard or hypoxic conditions when compared with transfection with an enhanced green fluorescent protein (EGFP)-expressing plasmid control. Ago2 protein overexpression (right panel) resulted in 2–12-fold overexpression of Ago2 (n = 3). The non-specific band described in Figure 4 was again detected below the Ago2 band. (C) The effect of Ago2 overexpression in JEG3 cells on mRNA levels for VEGF, β-hCG and p53 in standard or hypoxic conditions (n = 3). The expression of each transcript was determined in duplicate and expressed as fold overexpression at 24 h in EGFP control overexpression, defined as one for standard or for hypoxia. The differences were not statistically significant.
Discussion
Regulation of gene expression by miRNA pathways has recently emerged as a major determinant of diverse cellular functions (Alvarez-Garcia and Miska, 2005). As exquisite regulation of trophoblast transcript and protein expression is essential for human placental function in health and disease, this system is particularly well suited for analysis of miR biogenesis. The trophoblastic miRNA biosynthetic pathway is functional and produces diverse mature miRNA species (Barad et al., 2004); J.F.M. and Y.S., unpublished observation). We initially determined the expression of central miRNA machinery proteins in human villous trophoblasts and found that expression levels are unaffected by cellular differentiation. Whereas hypoxia is an important and relevant type of trophoblast injury and is implicated in clinical diseases such as FGR and pre-eclampsia (Levy et al., 2000; Roh et al., 2005; Vaiman et al., 2005), the role of miRNA in this process remains unknown. We found that hypoxia has no significant influence on expression of the key miRNA processing proteins. These data indicate that miRNA biogenesis pathways are active even in hypoxic cells and suggest that hypoxia exerts its effect on miRNA pathways by regulating individual trophoblastic miRNA genes, but not the miRNA machinery proteins. Although predictable, these findings are important, as regulation of miRNA biogenesis components substantially alters miRNA maturation and its regulatory effects. Knockout of the pivotal RISC component Ago2 in mice is embryonically lethal (Liu et al., 2004), demonstrating the indispensable functions of these key RISC components. Similarly, siRNA inhibition of core miRNA biosynthetic proteins in human cells demonstrate their rate-limiting role for miRNA biosynthesis, resulting in accumulation of pri-miRNAs and pre-miRNAs and diminished expression of mature miRNAs (Hutvagner et al., 2001; Lee et al., 2003; Ohrt et al., 2006). Lastly, siRNA knockdown of Ago2 and Drosha in Drosophila melanogaster S2 cells affects expression levels of ∼20% of the transcriptome (Rehwinkel et al., 2006).
Ago2 is the catalytic engine of mammalian RISC complexes (Liu et al., 2004). In contrast, the role of DP103 in miRNA processing is currently unknown. We and other investigators have cloned DP103 and examined its function and its interaction with the Ago2-containing human miRNP complexes (Hutvagner and Zamore, 2002; Mourelatos et al., 2002; Yan et al., 2003). A function in RISC assembly may be postulated by the observations that DP103 contains dsRBDs and ATPase domains, exhibits helicase activity (Yan et al., 2003) and is involved in assembly of human RNP complexes (Shpargel and Matera, 2005). We demonstrated that hypoxia does not impact trophoblast expression and distribution of Ago2 and DP103 in vitro and in vivo. Moreover, the interaction between these two pivotal RISC components is not modulated by hypoxia, bolstering the notion that RISC components are functional in human trophoblasts even during hypoxic stress. Finally, we showed that overexpression of the critical miRNA maturation regulator, Ago2, does not alter the anticipated expression pattern of a selected set of trophoblast transcripts, suggesting that Ago2 is not limiting for trophoblast hypoxic response.
Our results cannot rule out the possibility that hypoxia regulates other miRNA biogenesis proteins, such as Pasha GW182 or Rck/p54. Stress might also modulate the kinetics of miRNA biogenesis enzymes or induce differential regulation of discrete miRNA species via selective miRNA biosynthetic co-factors or (yet unidentified) trophoblast- or hypoxia-specific regulators. As miRNA genes are regulated by RNA Polymerase II (Pol II) and Pol III (Lee et al., 2004; Borchert et al., 2006), it is likely that promoter-specific DNA binding proteins modulate the transcription rate of individual miRNA genes, which comprise the cellular adaptive response to stress. Furthermore, we did not study effects down-stream of the miRNA biosynthetic pathway, which may affect miRNA actions. Intriguingly, it was recently shown that stress conditions other than hypoxia can alter miRNA actions via relief of miRNA-mediated translational repression (Bhattacharyya et al., 2006), an effect down-stream of miRNA biosynthesis. Elucidating mechanisms of biosynthesis, stability and function of miRNAs during cellular function and stress may shed light on the contributions of miRNAs to the development and function of vital tissues such as the placenta.
Acknowledgements
We are grateful to Elena Sadovsky, Lynne Collins and Aaron Barton for their technical assistance during these studies, to Baosheng Chen and Timothy Schaiff for discussions and to Lori Rideout for assistance during manuscript preparation. This work was supported by the Jan Kornelis de Cock Foundation, Foundation De Drie Lichten and the Groningen University Institute for Drug Exploration (to R.B.D.) and NIH R21-HD053878 and R01-HD045675 (to Y.S.). R.B.D. was also supported by a Scholarship for Talented Students from the Dutch Ministry of Education.
References
Author notes
Presented, in part, at the 53rd Annual Scientific Meeting of the Society for Gynecologic Investigation, Toronto, Canada, March 2006.