-
PDF
- Split View
-
Views
-
Cite
Cite
Tsubasa Shoji, Yasuyuki Yamada, Takashi Hashimoto, Jasmonate Induction of Putrescine N-Methyltransferase Genes in the Root of Nicotiana sylvestris, Plant and Cell Physiology, Volume 41, Issue 7, 15 July 2000, Pages 831–839, https://doi.org/10.1093/pcp/pcd001
Close - Share Icon Share
Abstract
Nicotine alkaloids are synthesized in the root of Nicotiana species, and their synthesis increases after insect attack, wounding and jasmonate treatment of the leaf. Putrescine N-methyltransferase (PMT) catalyzes the first committed step in nicotine biosynthesis. The expression patterns of the three Nicotiana sylvestris PMT genes (NsPMT1, NsPMT2, and NsPMT3) are reported in this study. Transcripts of the NsPMT genes were detected only in the root, and were up-regulated by methyl jasmonate treatment. When the 5′-flanking regions of NsPMT1, NsPMT2, and NsPMT3 were fused independently to β-glucuronidase reporter gene and introduced into N. sylvestris by Agrobacterium-mediated transformation, all introduced transgenes were expressed in the cortex, endodermis, and xylem in the root, as well as upregulated by methyl jasmonate treatment. These qualitatively similar patterns of expression for the NsPMT genes are achieved with only 0.25 kb of their conserved 5′-flanking regions, which contained no known jasmonate-responsive elements.
The nucleotide sequences reported in this paper have been submitted to the DDBJ, EMBL and GenBank under accession numbers AB004322 (NsPMT1), AB004323 (NsPMT2) and AB004324 (NsPMT3).
(Received January 7, 2000; Accepted April 11, 2000).
Introduction
Plants have evolved several defense strategies against herbivores. Toxic proteins and secondary metabolites accumulating in damaged and neighboring plant tissues are believed to prevent excessive herbivory (Harborne 1997, Baldwin and Preston 1999). In many cases, defensive proteins and chemicals increase after herbivore attack both in local and remote tissues. Herbivory and mechanical wounding initiate signal transduction pathways which ultimately up-regulate plant genes involved in inducible defenses. Extensive studies on tomato proteinase inhibitor (PI) genes revealed that systemic induction of PI genes after insect feeding or mechanical wounding are mediated by signal transduction pathways which possibly involve systemin, oligosaccharides, jasmonates, and electric signal (Koiwa et al. 1997). Jasmonic acid and its methyl ester, collectively referred to jasmonates, are fatty acid derivatives synthesized from linolenic acid via the octadecanoid pathway (Creelman and Mullet 1997). Endogenous levels of jasmonates increase rapidly after wounding, which is followed by induction of the defense-related genes such as the PI genes (Wasternack and Parthier 1997). Jasmonates also induce accumulation of several secondary metabolites possibly involved in plant defense (Gundlach et al. 1992, Blechert et al. 1995). Tomato and Arabidopsis mutants impaired in jasmonate biosynthesis are also compromised in their resistance to insect predators (Howe et al. 1996, McConn et al. 1997). These studies suggest that jasmonates are ubiquitous regulatory signals involved in herbivory- and wound-induced gene expression (Creelman and Mullet 1997, Reymond and Farmer 1998).
Nicotine is the predominant alkaloid in a number of Nicotiana species and may function as a defensive toxin by acting on the nervous system of herbivores (Wink 1998). After leaf damage by insects or wounding, the nicotine content of leaves increases dramatically within several days, lending support to its protective role (Baldwin 1989, Baldwin 1998, Baldwin 1999). Nicotine is synthesized in the root and is then transported to aerial parts of the plant via the xylem (Dawson 1942a, Wink and Roberts 1998). Thus, tissue damage is recognized in the leaf and the wound signal is transmitted to the root where the genes for nicotine biosynthesis are activated. One candidate for this long-distance signal is jasmonate (Baldwin et al. 1994, Zhang and Baldwin 1997). Endogenous jasmonate levels in Nicotiana sylvestris root increase within 2 h after leaf damage and positively correlate with the damage-induced accumulation of nicotine (Baldwin et al. 1994, Baldwin et al. 1997, Ohnmeiss et al. 1997). When jasmonate biosynthesis is inhibited by prior treatment of N. sylvestris leaves with inhibitors of the octadecanoid pathway, increases in nicotine accumulation in response to wounding is blocked (Baldwin et al. 1996). Conversely, exogenous application of jasmonates to N. sylvestris leaves or roots induces nicotine biosynthesis (Baldwin et al. 1994, Baldwin et al. 1996). Consistent with the jasmonate-induced accumulation of nicotine, several genes involved in nicotine biosynthesis have been isolated as jasmonate-inducible genes in tobacco BY-2 suspension cell cultures (Imanishi et al. 1998).
Nicotine and tropane alkaloids, some of which have been used for their medicinal properties, are synthesized from putrescine, a diamine derived from ornithine and/or arginine in several Solanaceae species (Hashimoto and Yamada 1994, Kutchan 1995). Putrescine N-methyltransferase (PMT) catalyzes the S-adenosylmethionine-dependent N-methylation of putrescine at the first committed step in such putrescine-derived alkaloid biosynthesis. The PMT cDNA was first isolated from tobacco (Nicotiana tabacum) and has been shown to be expressed in the root (Hibi et al. 1994). In tobacco, the PMT gene(s) is under genetic control by the Nic regulatory loci, repressed by auxin, and induced by jasmonates (Hibi et al. 1994, Imanishi et al. 1998). PMT gene(s) in Atropa belladonna, a tropane alkaloid-producing medicinal plant, is expressed specifically in the root pericycle cells adjacent to xylem, as shown by expression analysis of a promoter::GUS fusion transgene (Suzuki et al. 1999). N. sylvestris, a diploid progenitor of tobacco, contains three PMT genes (NsPMT1, NsPMT2, and NsPMT3) which encode proteins which are highly conserved, except for varying numbers of tandem repeats of an 11 amino acid motif in their N-terminal regions (Hashimoto et al. 1998a). In this report, we analyzed the expression patterns of the three members of the N. sylvestris PMT gene family by RT-PCR and promoter::GUS fusion gene analyses. We demonstrated that a highly conserved NsPMT 5′-flanking region of only 0.25 kb is sufficient for tissue-specific and jasmonate-responsive expression of all three PMT genes.
Materials and Methods
Plant materials and jasmonate treatment
Sterile plants of Nicotiana tabacum cv. Burley21 and N. sylvestris were grown in Agripots (Kirin, Tokyo) on half strength B5 medium (Gamborg et al. 1968) containing 1.5% sucrose and 0.3% gellan gum, under continuous illumination (ca. 40 µmol Em–2 s–1) at 26°C. For methyl jasmonate (MeJA) treatment of plants, a small cotton ball soaked with 0.5 ml of a 100 µM MeJA solution, was placed beside the plant in a tightly sealed Agripot for the indicated time periods. N. sylvestris hairy roots were subcultured at 2-week intervals in liquid MS medium (Murashige and Skoog 1962) with shaking at 90 r.p.m. at 26°C in the dark. MeJA solution was directly added to 3-day-old root cultures to a final concentration of 20 µM.
RNA gel blot analysis
Total RNA was isolated using an RNeasy kit (Qiagen), separated by electrophoresis on 1.2% formaldehyde agarose gels, and blotted onto a Hybond-N+ nylon membrane (Amersham). Equal loading for each lane was confirmed by staining the gels with ethidium bromide before blotting. The blots were hybridized in 50% (v/v) formamide, 5 × SSC, 2 × Denhardt’s solution, 0.1% (w/v) SDS, and 0.1 mg ml–1 salmon sperm DNA at 42°C for 16 h with probes labeled with 32P by using a Bcabest labeling kit (Takara, Kyoto). A tobacco PI-II cDNA (Balandin et al. 1995), a tobacco spermidine synthasecDNA (Hashimoto et al. 1998b), and the GUS gene were used as DNA probes. Washing was performed three times in 0.2 × SSC and 0.1% (w/v) SDS at 65°C. For the PMT probe, an antisense RNA probe was synthesized with a Riboprobe kit (Promega) from tobacco PMT cDNA (Hibi et al. 1994). Hybridization and washing with the RNA probe was carried out according to the manufacturer’s instructions.
RT-PCR analysis
First-strand cDNAs were synthesized from total RNA (5 µg) by using a cDNA synthesis kit (Takara), and quantified based on the efficiency of cDNA synthesis according to the manufacture’s instructions. Aliquots of first-strand cDNA were amplified using two PCR primers (5′-CATATCTACCAACACAAATGG and 5′-AATGCGC-TAAA-CT-C-T-G-AAAACC) for 3 min at 94°C, followed by 25 cycles of 1 min at 94°C, 1 min at 55°C, and 1 min at 72°C, and finally a 10 min incubation at 72°C. The regions to which these PCR primers anneal, flank the NsPMT repeat elements of variable length, and are conserved among the three NsPMT genes (Hashimoto et al. 1998a). To quantify the PCR products, [α-32P]dCTP was added to the PCR reaction mixtures. Amplified PMT cDNA fragments were separated on a 5% polyacrylamide gel and the gel was dried and exposed to X-ray film. As a positive control, N. sylvestris genomic DNA (50 ng) was also used as template for PCR.
DNA cloning and plasmid construction
Genomic clones containing the NsPMT genes had been subcloned into pBluescript II vectors (Hashimoto et al. 1998a). In this study, the nucleotide sequences of the 5′-flanking regions of the NsPMT genes were determined using an ABI DNA sequencer (Perkin Elmer). DNA sequences were aligned with a GeneWorks program (Intelligenetics, Campbell, U.S.A.).
The 5′-flanking regions of the NsPMT genes were fused in frame upstream of the GUS coding sequence in the pLM9 vector (Mlynarova et al. 1995). ClaI and NcoI sites were added to the ends of the DNA fragments (−254 to +34 in NsPMT1, −236 to +34 in NsPMT2, and −257 to +34 in NsPMT3; +1 denotes the adenine in the translation initiation ATG) by PCR with appropriate primers, and the digested PCR products were inserted into the ClaI-NcoI site of pLM9 to give pMT102, pMT202, and pMT302 vectors. After the NcoI site at −1,231 of NsPMT3 was changed to a ClaI site by linker ligation, the ClaI-XhoI fragment (−1,231 to −203) was used to replace the ClaI-XhoI sequence on pMT302 (−257 to −203) to give pMT301. Likewise, after the HindIII site at −1,743 of NsPMT2 was changed to a ClaI site by linker ligation, the ClaI-HindIII fragment (−1,231 to −54) was used to replace the ClaI-HindIII sequence on pMT202 (−236 to −54) to give pMT201.
Plant transformation
Binary vectors were used to transform Agrobacterium rhizogenes strain 15834 and A. tumefaciens strain LBA 4404 by electroporation. Leaf-disc infection and plant regeneration were done according to Kanegae et al. (1994). Transgenic plants were regenerated from leaf discs, grown in a greenhouse to maturity, and selfed. The seeds were germinated on B5 agar plates, and the seedlings were used for histochemical GUS analysis.
Histochemical and fluorometric GUS assays
GUS activity was stained histochemically according to Jefferson (1987) with some modifications. Plant tissues were incubated in 1 mM 5-bromo-4-chloro-3-indolyl-β-d-glucuronide, 50 mM potassium phosphate buffer (pH 7.0), 0.1% (v/v) Tween-20, 20% (v/v) methanol, and 5 mM dithiothreitol at 37°C for 1 h to overnight. For aerial tissues, pigments were removed by incubation in ethanol after histochemical staining. Stained tissues were then sliced into sections of 50 to 75 µm in thickness with a DTK-1500 microslicer (DohanEM, Kyoto).
GUS activity was quantified fluorometrically according to Jefferson (1987) with some modifications. Hairy roots were homogenized in GUS assay buffer (phosphate-buffered saline containing 1 mM phenylmethylsulfonyl fluoride and 5 mM dithiothreitol). After centrifugation of the homogenate, proteins in the supernatant were precipitated at 80% (w/v) saturation ammonium sulfate, and then desalted with a PD-10 column (Amersham Pharmacia Biotech). The desalted enzyme solution was incubated at 37°C for 5 min in GUS assay buffer containing 0.05 mM 4-methylumbeliferryl-β-d-glucuronide. The amount of 4-methylumbelliferone formed by the GUS reaction was determined using a fluorescence spectrophotometer (F-3010; Hitachi, Ibaraki). Protein concentration was determined with a Coomassie protein assay reagent (Pierce, Rockford, U.S.A.).
Results
PMT genes are induced by jasmonate treatment in tobacco roots
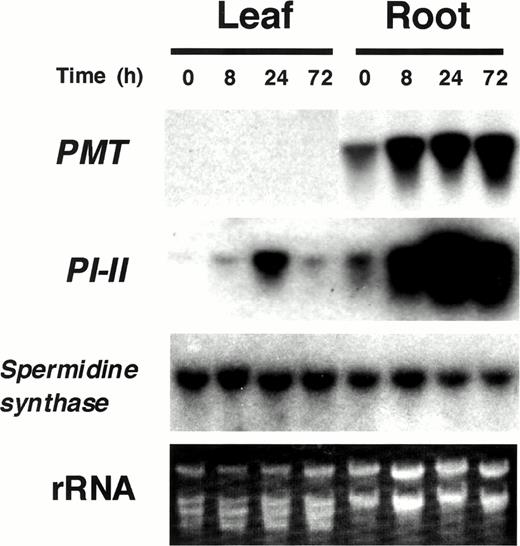
Tobacco PMT genes have been previously shown to be up-regulated upon jasmonate treatment in tobacco BY-2 suspension cultured cells (Imanishi et al. 1998). Thus, we first examined whether exogenous application of jasmonates induces PMT gene expression in intact tobacco plants. Axenically grown tobacco plantlets were treated with MeJA vapor in air-tight containers. RNA gel blot analysis using a tobacco PMT probe revealed that PMT mRNA in the root, which was present even in the untreated root, began to increase within 8 h after MeJA treatment, and this elevated level was maintained until 72 h (Fig. 1). In contrast, no PMT mRNA was detectable in the leaf even after MeJA treatment. The same RNA gel blots were also probed with the tobacco proteinase inhibitor-II (PI-II) gene (Balandin et al. 1995). The PI-II mRNA levels increased strongly in the leaf and, although smaller in magnitude, in the root after MeJA treatment. Both hybridization signals started to increase within 8 h, reached a maximum at 24 h, and already were in decline at 72 h. We also analyzed the expression of the spermidine synthase gene, which encodes an enzyme of polyamine biosynthesis and is the presumed ancestor gene of PMT (Hibi et al. 1994, Hashimoto et al. 1998b). The spermidine synthase mRNA was present in both untreated root and leaf, and the RNA level was not affected by MeJA application.
These results demonstrated that exogenous application of MeJA to tobacco plantlets induces PMT gene expression in the root, but not in the leaf, and that the organ-specificity and induction kinetics of PMT are distinct from those of PI-II.
Conserved regions within 5′-flanking sequences of three NsPMT genes
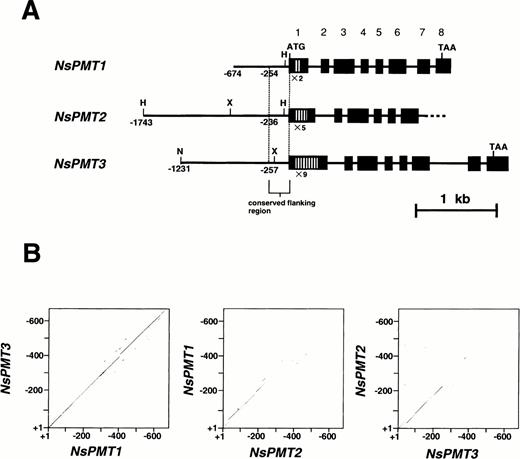
Cultivated tobacco (N. tabacum) is a natural amphidiploid and contains five PMT genes (Riechers and Timko 1999). To facilitate gene expression analysis, PMT genes in its diploid progenitor were studied. N. sylvestris is believed to have contributed one of the parental genomes to tobacco, and possesses three PMT genes, NsPMT1, NsPMT2, and NsPMT3 (Fig. 2A, Hashimoto et al. 1998a). For this study, we determined the sequences of their 5′-flanking regions, which had previously been isolated and subcloned into plasmid vectors (Hashimoto et al. 1998a). The 5′-flanking sequences of 674 bp in NsPMT1, 1,743 bp in NsPMT2, 1,231 bp in NsPMT3 (nucleotide positions were determined by setting the adenine of the first translational start ATG to position 1) were deposited into the GenBank database. Putative TATA-boxes were found about 40–50 bp upstream from a position corresponding to the 5′-end of the tobacco PMT cDNA, although the presumed TATA-box sequence of NsPMT2 (5′-TATTTAAA) was slightly different from the reported consensus sequence (5′-TATA(T/A)A(T/A)A; Yamaguchi et al. 1998).
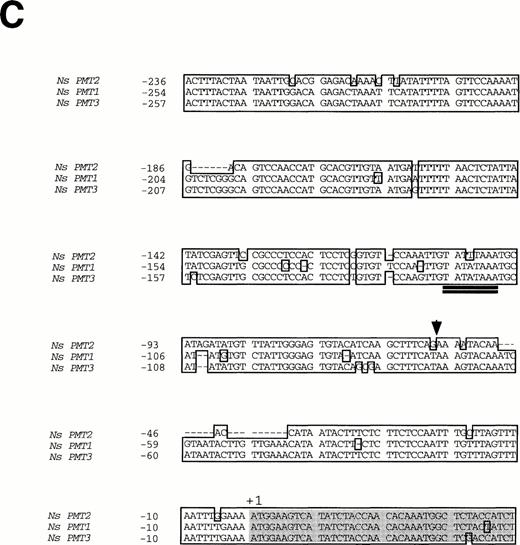
Sequence alignments between the 5′-flanking regions of NsPMTs revealed that a streach of approximately 250 bp of DNA sequence upstream from the first ATG is highly conserved among all three genes (Fig. 2B, C). Further upstream from these conserved regions, the NsPMT2 sequence diverges from the NsPMT1 and NsPMT3 sequences, and there is no conserved element longer than 6 bp between NsPMT2 and either of the other two NsPMTs. On the other hand, NsPMT1 and NsPMT3 are very similar in their sequences along the entire lenght of the sequenced fragments.
Expression patterns of three NsPMT genes
To study the expression patterns of individual members of the PMT gene family, we amplified the Exon 1 regions of NsPMT cDNAs by RT-PCR. The Exon 1 regions have been reported to contain different length of tandem repeat domains from one another (Fig. 2A, Hashimoto et al. 1998a), and thus gave rise to different lengths of PCR products which are specific to individual members of the PMT gene family.
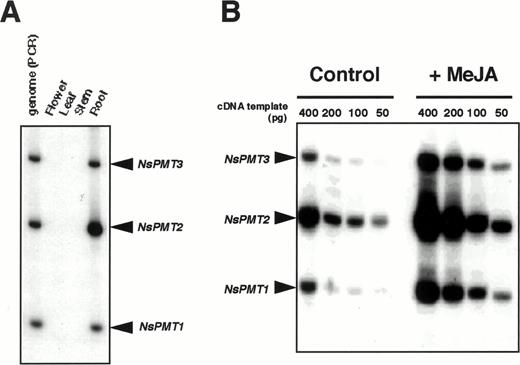
When NsPMT cDNA fragments were amplified from total RNA isolated from various organs, RT-PCR products were obtained only from root RNA (Fig. 3A). PCR products from genomic DNA served as amplification controls for the respective NsPMT genes. Quantification of the RT-PCR signals relative to the genomic PCR signals indicated that the mRNA levels of NsPMT1, NsPMT2, and NsPMT3 in the root are 1 : 6 : 1. To study whether jasmonate treatment induces NsPMT gene family members differentially, RT-PCR was performed with total RNA isolated from N. sylvestris hairy roots treated with or without 20 µM MeJA for 4 h (Fig. 3B). To rigorously compare the amplified signal levels between the two treatments, four different amounts of cDNA template were used and the signals quantified before they reached their plateaus. The transcripts from all three NsPMT genes were upregulated in MeJA-treated hairy roots, and their induction levels were approximately 4-fold in all three NsPMT genes.
The 5′-flanking regions of NsPMTs mediate tissue-specific and jasmonate-inducible expression
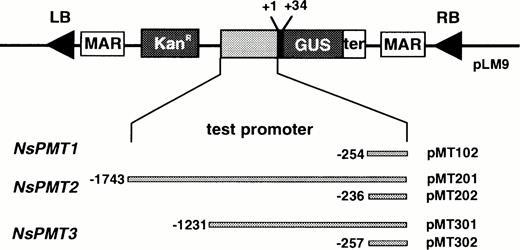
To study the function of the 5′-flanking regions of the NsPMT genes, the promoter fragments were translationally fused to a GUS reporter gene in the pLM9 binary vector (Fig. 4). The matrix attachment regions in pLM9 sandwich transgenes so that the expression levels of the transgenes are believed to be less variable among transgenic clones (Mlynarova et al. 1995). The flanking sequences which are highly conserved among the NsPMT promoters were used to construct pMT102, pMT202, and pMT302 vectors. All five constructs were introduced into N. sylvestris by Agrobacterium rhizogenes to produce hairy root clones. In addition, pMT301 and pMT302 were also used to produce N. sylvestris transgenic plants by A. tumefaciens.
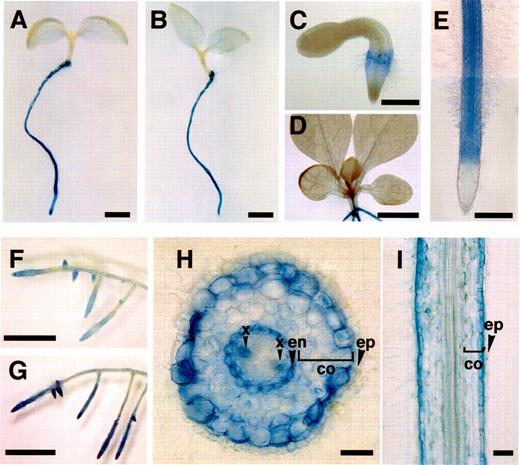
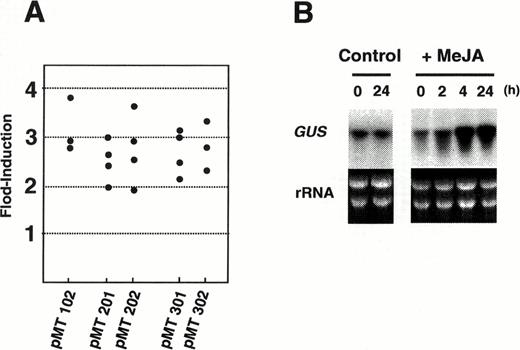
Transgenic seedlings and hairy roots were stained for GUS activities (Fig. 5). Transgenic seedlings transformed with either pMT301 or pMT302 gave essentially identical staining patterns: GUS activities were detected only in the roots and not in the aerial organs (A–D). In root tissues, root tip, epidermis, and root hairs were not stained (E). Transgenic hairy roots transformed with any of the five constructs gave the same tissue-specific GUS expression pattern (data not shown). Representative staining patterns from the pMT201-transformed hairy root clones are shown in F–I. GUS staining was restricted mainly to young root tissues (F). A cross-sectional view of a strongly stained root region, showed that cortex, endodermis, and xylem were stained blue (H). Of the three cell layers in the root cortex, the outermost layer was stained most strongly, as clearly observed in a longitudinal section (I). GUS activity was not detected in the epidermis and root hairs. Treatment of transgenic hairy root clones with 20 µM MeJA did not change the cell types of GUS staining, but did increase the overall level of staining (compare F and G). We further quantified GUS activities fluorometrically in MeJA-treated and untreated control roots. Several independent transgenic hairy root clones were analyzed for each construct (Fig. 6A). GUS activities increased 2–4 fold with all promoter constructs after treatment with 20 µM MeJA for 1 d. In a transgenic hairy root clone transformed with pMT201, expression of the transgene was analyzed by RNA gel blot analysis (Fig. 6B). GUS mRNA started to increase within 2 h and continued to increase until 24 h after MeJA application.
These results indicate that the conserved, first 0.25 kb of the NsPMT promoter regions is sufficient to confer a cell-specific expression pattern in N. sylvestris roots and to respond positively to exogenously supplied MeJA.
Discussion
Structure of NsPMT genes
The nucleotide sequences of the three PMT genes of N. sylvestris show a high degree of similarity to one another both in their coding and noncoding regions, which includes introns, untranslated regions, and the 5′-flanking region (Hashimoto et al. 1998a, this study). This sequence conservation suggests a recent generation of the NsPMT gene family by gene duplication from a common ancestor gene (Maeda and Smithies 1986). The 5′-flanking sequence of NsPMT2 diverges suddenly from the sequences of NsPMT1 and NsPMT3 upstream of −236. This point of divergence might represent one end of a gene duplication event.
One possible advantage of possessing duplicated genes is greater flexibility in temporal and spacial control of gene expression. In branched metabolic pathways, members of small gene families which encode rate-limiting enzyme are often differentially regulated to meet the distinct metabolic demands of the downstream branches. Thus, individual members of small gene families, such as phenylalanine ammonia-lyase (Liang et al. 1989), chalcone synthase (Martin 1993), terpenoid synthase (Bohlmann et al. 1998), and 3-hydroxy-3-methylglutaryl-CoA reductase (Yang et al. 1991, Enjuto et al. 1994), are expressed in distinct cell types, and respond differentially to various regulatory signals. However, all three NsPMT genes are expressed in the same cell types and are activated similarly by jasmonate treatment. A possible explanation for their similarity in expression patterns is that nicotine and its immediate metabolic products are the only known compounds derived from the linear biosynthetic pathway in which PMT is involved (Leete 1980), and that consequentially there is no advantage in controlling PMT gene family members differentially. In this regard, the gene duplication events, which presumably involved the ancestral PMT gene of N. sylvestris, might have been fixed by chance. Incidentally, other diploid Nicotiana species, such as N. tomentosiformis and N. otophora possess only a single copy of the PMT gene (Hashimoto et al. 1998a, Riechers and Timko 1999).
Cell-specific expression of NsPMT genes
RNA gel blot and RT-PCR analyses showed that NsPMT genes are specifically expressed in the root. This root-specific expression pattern was also evident for the NsPMT::GUS transgenes, indicating that the 5′-flanking regions which were used most likely contain imformation sufficient for the spatial regulation of NsPMT genes. Expression of NsPMT::GUS transgenes was detected in cortex, endodermis, and xylem in the root (Fig. 5). Since nicotine is transported from the root to the aerial parts of the plant through the xylem (Wink and Roberts 1998), PMT expression in the xylem tissue may facilitate nicotine loading into the xylem. If nicotine is produced in the cortex, the alkaloid must be transported symplastically to the stele since the Casparian strip of the endodermis will block apoplastic transport (Clarkson 1996). The cellular lipid bilayer is effectively impermeable to charged or polar alkaloids, which are therefore postulated to require a transport carrier for movement between cells (Mende and Wink 1987). In contrast, the weakly basic nicotine molecule was shown to pass through the tonoplast by simple diffusion (Kurkdjian 1982), and may not require a specific carrier for cell-to-cell translocation. Thus, nicotine in the root cortex might simply diffuse into the stele and enter the upward-flowing xylem stream.
It is also possible that part of the nicotine synthesized in the cortex may also move out towards the epidermis and then out of the root. A significant proportion of the nicotine produced in Nicotiana root cultures is known to be secreted into the culture medium (e.g. Dawson 1942b, Hamill et al. 1986, Hibi et al. 1994). The strong expression of the PMT promoters in the sub-epidermal cortex correlates well with the secretion of nicotine from the root. Although the distribution of nicotine in root cell types and in the surrounding soil is not known, the secretion of nicotine may play a defensive role in warding off soil insects in the rhizosphere.
The recent immunohistochemical localization of several enzymes involved in the biosynthesis of monoterpenoid indole alkaloids (St-Pierre et al. 1999) and tropane alkaloids (Nakajima and Hashimoto 1999) showed that enzyme subsets were present in different, non-overlapping cell types, suggesting intercellular translocation of pathway intermediates. In order to determine the cell types involved in nicotine biosynthesis more definitively, the spatial expression patterns of several other enzymes in the nicotine pathway must be analyzed.
Jasmonate-inducible PMT gene expression
Jasmonates play key roles as global regulators of plant gene expression in growth, development and responses to environmental stresses, including responses against herbivory and wounding (Staswick 1995). Consistent with the pleiotropic effects of jasmonates, various genes involved in different cellular functions have been reported as jasmonate-inducible genes (Wasternack and Parthier 1997). As expected from their diverse functions, these jasmonate-inducible genes may be regulated by jasmonates in combination with other regulatory cues to accomplish specific functions at the right time and place. Thus, like other biological systems, gene activation by jasmonates may be mediated through combinatorial signal transduction pathways. As an initial steps in the understanding of jasmonate-signaling networks, several cis promoter elements in jasmonate-inducible genes have been characterized in relation to their jasmonate response. Analysis of the potato PI-II promoter has identified a G-box sequence (CACGTGG) as an essential element for the jasmonate response (Kim et al. 1992), and this G-box-type motif has also been identified in the jasmonate-responsive promoters of several other genes (Mason et al. 1993, Kim et al. 1993, Rouster et al. 1997). Recently, a novel jasmonate- and elicitor-responsive element containing a GCC motif was identified in a terpenoid indole alkaloid biosynthetic gene, and was shown to bind cognate interacting factors belonging to AP2-domain proteins (Menke et al. 1999). In this study, we demonstrated that approximately 0.25 kb of the conserved 5′-flanking regions of the NsPMT genes confers jasmonate-responsive expression to a downstream reporter gene (Fig. 6). However, our computer-assisted search was not successful in revealing known jasmonate-responsive elements in the three 0.25 kb PMT promoter regions. Future analysis of the NsPMT promoters may lead to the identification of a novel jasmonate-responsive cis-element. It would also be of interest to study other nicotine biosynthetic genes to see whether their promoters respond to jasmonates, and whether their jasmonate-responsive regions show any similarity to the NsPMT promoters.
Acknowledgements
We thank Dr. H. Sano for the tobacco PI-II cDNA, Dr. J.-P. Nap for pLM9, and Dr. R. Winz for critical reading of the manuscript. This research was supported in part by a grant (“Molecular Breeding of Plants for Foods, Resource and Environment in the 21st Century”, JSPS-RFTF9616001, in “Research for the Future” Program) from the Japan Society for the Promotion of Science to T.H.
Corresponding author: E-mail, hashimoto@bs.aist-nara.ac.jp.
Fig. 1 RNA gel blot analysis of tobacco plants treated with MeJA. Total RNA was prepared from leaves and roots of tobacco plantlets treated with MeJA vapor for 0, 8, 24, or 72 h. Ten micrograms of total RNA was loaded onto each lane. RNA gel blots were probed with tobacco PMT, PI-II, and spermidine synthase genes.
Fig. 2 Structure of NsPMT genes and nucleotide sequence alignments of their 5′-flanking regions. (A) Schematic representation of NsPMT gene structures. Nucleotides are numbered from each translational start site. Several restriction sites are indicated: H (HindIII), X (XhoI), and N (NcoI). Solid boxes indicate exons and open boxes indicate tandem repeats used in RT-PCR analysis. The 5′-flanking region and introns are shown as solid bars. (B) Homology plots between the nucleotide sequences of the 5′-flanking regions. Each dot marks the midpoint of ten consecutive nucleotides in which more than seven residues were identical between two sequences. (C) Nucleotide sequence alignment of the 5′-flanking regions. Identical nucleotides are boxed, and protein coding regions are shaded. An arrowhead corresponds to the 5′-end of tobacco PMT cDNA. Putative TATA boxes are double-underlined.
Fig. 3 RT-PCR expression analysis of PMT genes in N. sylvestris. (A) Expression of NsPMT genes in various organs. 32P-labeled PCR products were amplified from 100 ng of cDNA derived from flower, leaf, stem, or root tissue, and separated on a 5% polyacrylamide gel. Genomic DNA was also amplified. Positions of the respective NsPMT cDNA fragments are indicated on the right. (B) Expression of NsPMTs genes in hairy roots treated with 20 µM MeJA for 4 h. Variable amounts of cDNA were used as template for PCR amplification.
Fig. 4 Binary vectors used for Agrobacterium-mediated transformation. PMT promoters were translationally fused to GUS reporter genes in the T-DNA region of the pLM9 binary vector. KanR, neomycin phosphotransferase gene; MAR, matrix attachment region derived from a chicken lysozyme gene; LB, left border sequence; RB, right border sequence; ter, terminator sequence of nopaline synthase gene.
Fig. 5 Histochemical localization of GUS activity in transgenic seedlings and transgenic hairy roots of N. sylvestris. (A and E) 5-day-old transgenic seedlings transformed with pMT301. (B) 5-day-old transgenic seedling transformed with pMT302. (C) 2-day-old transgenic seedling transformed with pMT301. (D) 14-day-old transgenic seedling transformed with pMT301. (F, G, H, and I) transgenic hairy root transformed with pMT201. (G) The same transgenic hairy root clone as in F, but treated with 20 µM MeJA for 3 d. Bar=2 mm in A and B, 0.5 mm in C and E, 5 mm in D, F, and G, 0.1 mm in H and I. co, cortex; en, endodermis; x, xylem; ep, epidermis.
Fig. 6 Effects of MeJA treatment on GUS gene expression in transgenic N. sylvestris hairy roots. (A) Induction of GUS activity by MeJA. Hairy roots were treated with 20 µM MeJA for 1 d, and increased GUS activity was divided by the activity of the untreated control. Each black dot represents fold-induction of GUS activity in each independent hairy root clone transformed by the binary vector shown at the bottom. (B) RNA gel blot analysis of the GUS gene in transgenic hairy roots transformed with pMT201. Total RNA was isolated from roots after treatment with 20 µM MeJA for 0, 2, 4, or 24 h. Ten micrograms of total RNA was loaded onto each lane. RNA gel blot was probed with the GUS gene.
Abbreviations
- GUS
β-glucuronidase
- MeJA
methyl jasmonate
- NsPMT
Nicotiana sylvestris putrescine N-methyltransferase
- PI
proteinase inhibitor
- PMT
putrescine N-methyl-transferase.
References
Balandin, T., Does, C.V.D., Albert, J-M.B., Bol, J.F. and Linthorst, H.J.M. (
Baldwin, I.T. (
Baldwin, I.T. (
Baldwin, I.T. (
Baldwin, I.T. and Preston, C.A. (
Baldwin, I.T., Schmelz, E.A. and Ohnmeiss, T.E. (
Baldwin, I.T., Schmelz, E.A. and Zhang, Z-P. (
Baldwin, I.T., Zhang, Z.-P., Diab, N., Ohnmeiss, T.E., McCloud, E.S., Lynds, G.Y. and Schmelz, E.A. (
Blechert, S., Brodschelm, W., Hoelder, S., Kammerer, L., Kutchan, T.M., Mueller, M.T., Xia, Z-Q. and Zenk M.H. (
Bohlmann, J., Meyer-Gauen, G. and Croteau, R. (
Clarkson, D.T. (
Creelman, R.A. and Mullet, J.E. (
Dawson, R.F. (
Enjuto, M., Balcells, L., Campos, N., Caelles, C., Arro, M. and Boronat, A. (
Gamborg, O.L., Miller, R.A. and Ojima, K. (
Gundlach, H., Mueller, M.J., Kutchan, T.M. and Zenk, M.H. (
Hamill, J.D., Parr, A.J., Robins, R.J. and Rhodes, M.J.C. (
Harborne, J.B. (
Hashimoto, T., Shoji, T., Mihara, T., Oguri, H., Tamaki, K., Suzuki, K. and Yamada, Y. (
Hashimoto, T., Tamaki, K., Suzuki, K. and Yamada, Y. (
Hashimoto, T. and Yamada, Y. (
Hibi, N., Higashiguchi, S., Hashimoto, T. and Yamada, Y. (
Howe, G.A., Lightner, J., Browse, J. and Ryan, C.A. (
Imanishi, S., Hashizume, K., Nakakita, M., Kojima, H., Matsubayashi, Y., Hashimoto, T., Sakagami, Y., Yamada, Y. and Nakamura, K. (
Jefferson, R.A. (
Kanegae, T., Kajiya, H., Amano, Y., Hashimoto, T. and Yamada, Y. (
Kim, S.-R., Choi, J.-L., Costa, M.A. and An, G. (
Kim, S.-R., Kim, Y. and An, G. (
Koiwa, H., Bressan, R.A. and Hasegawa, P.M. (
Kurkdjian, A. (
Kutchan, T.M. (
Leete, E. (
Liang, X., Dron, M., Cramer, C.L., Dixon, R.A. and Lamb, C.J. (
Maeda, N. and Smithies, O. (
Martin, C.R. (
Mason, H.S., DeWald, D.B. and Mullet, J.E. (
McConn, M., Creelman, R.A., Bell, E., Mullet, J.E. and Browse, J. (
Mende, P. and Wink, M. (
Menke, F.L.H., Champion, A., Kijne, J.W. and Memelink, J. (
Mlynarova, L., Jansen, R.C., Conner, A.J., Stiekema, W.J. and Nap, J-P. (
Murashige, T. and Skoog, F. (
Nakajima, K. and Hashimoto, T. (
Ohnmeiss, T.E., McCloud, E.S., Lynds, G.Y. and Baldwin, I.T. (
Reymond, P. and Farmer, E.E. (
Riechers, D.E. and Timko, M.P. (
Rouster, J., Leah, R., Mundy, J. and Cameron-Mills, V. (
Staswick, P.E. (
St-Pierre, B., Vazquez-Flota, F.A. and De Luca, V. (
Suzuki, K.-I., Yamada, Y. and Hashimoto, T. (
Wasternack, C. and Parthier, B. (
Wink, M. (
Wink, M. and Roberts, M.F. (
Yamaguchi, Y., Itoh, Y., Takeda, Y. and Yamazaki, K. (
Yang, Z., Park, H., Lacy, G.H. and Cramer, C.L. (