-
PDF
- Split View
-
Views
-
Cite
Cite
Adrian J. Hill, Hiroki Teraoka, Warren Heideman, Richard E. Peterson, Zebrafish as a Model Vertebrate for Investigating Chemical Toxicity, Toxicological Sciences, Volume 86, Issue 1, July 2005, Pages 6–19, https://doi.org/10.1093/toxsci/kfi110
Close - Share Icon Share
Abstract
Zebrafish (Danio rerio) has been a prominent model vertebrate in a variety of biological disciplines. Substantial information gathered from developmental and genetic research, together with near-completion of the zebrafish genome project, has placed zebrafish in an attractive position for use as a toxicological model. Although still in its infancy, there is a clear potential for zebrafish to provide valuable new insights into chemical toxicity, drug discovery, and human disease using recent advances in forward and reverse genetic techniques coupled with large-scale, high-throughput screening. Here we present an overview of the rapidly increasing use of zebrafish in toxicology. Advantages of the zebrafish both in identifying endpoints of toxicity and in elucidating mechanisms of toxicity are highlighted.
Zebrafish have been used predominantly in developmental biology and molecular genetics, but their value in toxicology as well as drug discovery has been recognized. To evaluate the toxicity of a chemical, it is essential to identify the endpoints of toxicity and their dose-response relationships, elucidate the mechanisms of toxicity, and determine the toxicodynamics of the chemical. In addition to detailed toxicological investigations of a single chemical, there also is a need for high-throughput large-scale screening for toxicity of several hundreds of chemicals at a time. In both cases, the zebrafish has numerous attributes.
More is probably known about “what is normal” in the zebrafish than any other fish species. This includes morphological, biochemical, and physiological information at all stages of early development and in juveniles and adults of both sexes. This makes using the zebrafish ideal for toxicology research where the objective is to identify adverse effects of chemical exposure.
ZEBRAFISH: A SUITABLE VERTEBRATE MODEL FOR TOXICITY ASSESSMENT
Although reliable data for extrapolating toxicant effects to humans are obtained through laboratory rodent studies, these are expensive, time consuming, and more restricted by law. Because genes, receptors, and molecular processes are highly conserved across animal phyla, studies with other animal species could be representative for “higher,” more-complex animals. Examples of model invertebrate organisms are the fruit fly and nematode worm, while model vertebrates include the medaka, zebrafish, frog, chick, mouse, and rat. In particular, gene programming and development in the early life stages of all vertebrates are highly conserved, to the extent that there are significant similarities in the morphology of all vertebrate embryos. In addition, just as there is transplacental transfer of chemicals from the maternal body to the embryo and fetus of mammals (Jacobson et al., 1984), chemicals are transferred from the female to the eggs of fish, amphibians, and birds prior to being laid. The egg then becomes the source of embryonic exposure to chemicals.
Advantages of the Zebrafish
There are numerous advantages for the use of zebrafish as a toxicological model species (Spitsbergen and Kent, 2003; Teraoka et al., 2003a) as well as for other disciplines. This is evident by the increasing number of publications which have used this organism in the recent past. In the early 1990s there were less than 100 zebrafish-related publications annually submitted. This rose to ∼1,000 at the turn of the century and now averages around 3,500 per year.
Small size, big value.
The main benefits of using zebrafish as a toxicological model over other vertebrate species are with regards to their size, husbandry, and early morphology. Unlike other fish species such as trout, zebrafish adults are only approximately 1–1.5 inches long. This greatly reduces housing space and husbandry costs, and there are now several companies specializing in zebrafish aquaria capable of supporting several thousand fish. Also zebrafish have been utilized as a laboratory species for quite some time so the optimum breeding and maintenance conditions have been well determined (Westerfield, 1995). In contrast to larger species, the minute size of the larval and adult zebrafish minimizes costs through low quantities of dosing solutions (experimental chemicals, drugs, pollutants) and thereby creates limited volumes of waste for disposal and minimizes quantities of labware and chemicals, both for treating and maintaining live fish and for performing various assays (low quantities of reagents) and histological assessments (small amount of embedding materials and microscope slides) (Hill et al., 2002). In addition, small embryos allow reasonable sample sizes to be tested together using a single cell-culture plate or series of Petri dishes to provide several experimental replicates at one time. This allowed the creation of high-throughput screens for toxicity testing, small-molecule screening, and drug discovery, in which zebrafish grow and develop in small microformat screening plates. From the egg stage, zebrafish embryos can survive for several days in a single well of a 384-well plate through the absorption of yolk and can be visually assessed for malformation (MacRae and Peterson, 2003). It is therefore possible, with the aid of automated systems and possibly fluorescent transgenics, to rapidly treat and screen large libraries of molecules for toxicity or therapeutic value. In one screen the cardiovascular system, central nervous system, ear and skin were assessed with a dissection microscope (Peterson et al., 2000); 1100 small molecules were screened; and several were found to disturb organ development. Another screen assessed 100 molecules that cause cardiac QT prolongation in humans, but manifested as bradycardia and AV block in zebrafish (Milan et al., 2003). If certain human diseases can be modeled in zebrafish, it has been proposed that these screens could be used to ameliorate the disease phenotype (MacRae and Peterson, 2003).
It is clear to see why zebrafish embryos are so useful.
Besides their size, this species is invaluable because of their high fecundity and transparent embryos. One pair of adult fish is capable of laying 200–300 eggs in one morning, and if appropriately maintained, they can provide this yield every 5–7 days. Additionally, as numerous fish are generally established for each genetic line, several pairs can be rotated to provide thousands of eggs daily and all year round. This can be maximized by using newly matured fish that are between 3 and 6 months old (sexual maturation occurs around 100 days; Skidmore, 1965). The rapid maturation of zebrafish also allows easy experimentation for transgenerational endpoints required for mutagenesis screening, establishing transgenic lines, and assessing chemicals for teratogenicity.
Their optical clarity allows for easy developmental staging, identification of phenotypic traits during mutagenesis screening, and assessment of endpoints of toxicity during toxicity testing. This proves even more valuable when used in concert with in situ hybridization (ISH) and immunochemistry (IHC) techniques. As with other vertebrates, ISH (Oxtoby and Jowett, 1989) and whole mount IHC (Dent et al., 1989; Klymkowsky & Hanken, 1991) assays can be performed on zebrafish. In situ digoxigenin-labeled RNA probes are now common in virtually every laboratory due to the ease of manufacture, enabling toxicologists to screen for chemical-induced abnormalities in the expression of specific genes. There are a vast amount of histochemical markers available, allowing assessments of aberrant morphology or activation of certain signaling pathways by toxicants through the staining of specific tissues and cells types. For example, immunohistochemical assessment of CYP1A expression following exposure of zebrafish to halogenated aromatic hydrocarbons and polyaromatic hydrocarbons has shown endothelial cells of the cardiovasculature system to be a cellular site of action (Andreasen et al., 2002b; Dong et al., 2002; Yamazaki et al., 2002). Whole-mount larval staining can be performed rather than first having to dissect the tissue or stain sections. The visualization of gene expression throughout the larvae again is possible due to the optical clarity of the zebrafish tissues, which allows successful penetration for light microscopy. This approach has been used to show colocalization of different aryl hydrocarbon receptor (AHR) and aryl hydrocarbon receptor nuclear translocator (ARNT) mRNAs in the zebrafish cardiovascular system (Andreasen et al., 2002b). Fluorescence labeling coupled with confocal microscopy can also enhance expression imagery and allow assessments in older, thicker larvae (MacDonald, 1999). Likewise, along with various other markers, fluorescent dyes can also be utilized for cell lineage tracing (Cooper et al., 1999a,b; Kimmel, 1989; Kimmel and Warga, 1988).
Assessments for toxicological endpoints.
Zebrafish development has been well characterized (Kimmel, 1989; Kimmel et al., 1995). Because zebrafish eggs remain transparent from fertilization to when the tissues become dense and pigmentation is initiated (at approximately 30–72 h post fertilization (hpf)), this allows unobstructed observations of the main morphological changes up to and beyond pharyngulation. Pigmentation can be prevented in vivo by treatment with 0.003% phenylthiourea or removed by bleaching after fixation, thereby extending this period of unobstructed assessment. Therefore, using little magnification, adverse effects of chemical exposure on development of the brain, notochord, heart, and jaw, trunk segmentation, and measurements of size can be assessed quantitatively. Also, unlike rodents, embryological development can be continually followed in live individuals rather than harvested embryos and fetuses. In addition, zebrafish embryos that are malformed, missing organs, or displaying organ dysfunction, can usually survive substantially past the time in which those organs start to function in healthy individuals. For example, mutant zebrafish such as silent heart, still heart, and slow mo (Chen et al., 1996), and toxicant-exposed embryos with heart abnormalities (Antkiewicz et al., in press; Incardona et al., 2004;) survive well beyond 24 hpf when the heart normally begins to beat (Kimmel et al., 1995). This is in contrast to rodent embryos with malformed hearts that tend to die in utero.
THE ZEBRAFISH GENOME
To the layperson, the zebrafish, as well as other fish species, may be considered a relatively simple organism when compared to mammals. However this is not the case with respect to the zebrafish genome. It is more complex than the human genome, because zebrafish have two more pairs of chromosomes than the twenty-three pairs of human chromosomes. This difference occurred because, at some time in teleost evolution, there was a whole-genome duplication event that did not occur in mammals. It resulted in numerous duplicate genes (paralogs) of those found in mammals. However, only a small proportion of these gene duplications remain today (Beier, 1998; Fishman, 1999; Metscher and Ahlberg, 1999; Postlethwait et al., 1999). Some of these duplicated genes have a new function, and others no longer express in the same tissues as the original genes (orthologs) (Force et al., 1999). The significance of this finding is that, where a mutation in a mammalian ortholog may cause embryonic lethality, a mutation in one of the zebrafish paralogs may show a less severe phenotype with the embryo remaining viable. This may permit analyses of gene function in mutant zebrafish that would be difficult to achieve in mutant mammals due to the associated embryo mortality (Spitsbergen and Kent, 2003). The obvious disadvantage of the gene duplication, however, is that certain changes in gene expression caused by toxic insult or a mutation may be hard to extrapolate to other vertebrates such as mammals.
The main features of embryogenesis (Kimmel, 1989) and the genetic basis of development have been extensively studied in zebrafish (Driever et al., 1994). The zebrafish research community has developed a range of resources useful to the toxicologist including mutant strains, cDNA clone collections, a physical map (Phillips and Reed, 2000), and a near-completed genome sequence (http://www.ensembl.org/Danio_rerio, http://www.sanger.ac.uk/Projects/D_rerio, http://www.ncbi.nlm.nih.gov/genome/guide/zebrafish). The genome sequencing project was started in 2001 and is scheduled to be finished in 2005. To accomplish this task, a bacterial clone physical map was constructed from BAC libraries using restriction enzyme fingerprinting (Marra et al., 1997), and a whole genome shotgun was produced from DNA derived from Tübingen embryos. Both techniques have led to the current assembly designated Zv4 comprising a total length of 1,560,480,686 bp in 21,333 fragments in which 69% has been placed on chromosomes 1–25. Confirmed gene sequences from this assembly are listed on the Vertebrate Genome Annotation (VEGA) database (http://vega.sanger.ac.uk) that is frequently updated.
GENETIC AND TISSUE MANIPULATION
Transient Gene Expression and Stable Transgenic Lines
Injection of DNA or RNA constructs at the 1- to 2-cell stage can yield a transient expression of the gene (Holder and Xu, 1999). This allows genes to be easily visualized with fluorescent markers and can restore functional gene expression in mutant embryos. Expression of a construct can also be initiated in precise cell types if the construct was created with a certain promoter sequence, such as that from a heat shock gene that can be activated by a laser (Halloran et al., 2000; Krone et al., 1997). This offers opportunities to investigate effects of toxicants on gene expression that produce developmental toxicity by acting at a particular target organ at a critical stage of development.
The use of transgenics as a toxicological tool has increased with improved, more successful techniques for creating stable lines of fish. In particular they can be used in two ways. First, once a specific gene has been identified either as a marker for specific tissues or as an essential part of a developmental pathway, these genes can be assessed for disruption after chemical exposure. Second, when a chemical has disrupted gene expression or morphology, recovery of normal gene expression can be assessed after application of therapeutic agents or morpholinos. Instead of performing lengthy staining methods over several days to identify the spatial and temporal expression pattern of the gene, a line of zebrafish can be created to express a transgene with a fluorescent reporter such as green fluorescent protein (GFP), thereby enabling assessments at any stage of early development easily with fluorescence microscopy. Examples include gfp:ngn1 (Blader et al., 1997, 2003) and gfp:islet1 (Higashijima et al., 2000), allowing visualization of neurons for use in neurotoxicity testing, and gfp:fli1 transgenics (Fig. 1), which permit visualization of endothelial cells of different vascular beds for the study of toxicants that disrupt cardiovascular development (Bello et al., 2004; Lawson and Weinstein, 2002).
Visualization of the zebrafish embryo vasculature by gfp:fli1 expressing endothelial cells. Light and fluorescence microscopy of transgenic zebrafish embryos at 50 hfp in lateral and dorsal orientation. BA: basilar artery, CCV: common cardinal vein, IOC: inner optic circle, MCeV: middle cerebral vein, MtA: mesencephalic artery, OV: optic vein, PHBV: primordial hindbrain channel, PLA: palatocerebral artery, and PrA: prosencephalic artery.
Pioneering transgenic studies utilized linear bacterial reporter plasmids (Stuart et al., 1988) and a pseudotyped virus construct (Lin et al., 1994). More recently the SceI meganuclease has been utilized to aid the integration of the transgene and increase the success rate for creating founder fish (Thermes et al., 2002). When injecting transgenes (or morpholinos), timing and developmental stage has proved to be important. Zebrafish eggs should be collected immediately after spawning and can be transferred to 4°C in embryo media to arrest development (Westerfield, 1995) and allow more time to perform injections at the appropriate developmental stages.
Transgenic zebrafish embryos expressing an AHR-dependent GFP reporter gene have been utilized to identify 2,3,7,8-tetrachlorodibenzo-p-dioxin (TCDD)-susceptible target tissues (Mattingly et al., 2001). Transgenics have also been used to help determine TCDD-induced neurotoxicity via assessing changes in sonic hedgehog and neurogenin expression in the zebrafish brain (Hill et al., 2003) and as an in vivo system that uses hsp70 gene activation (with an eGFP reporter) as a measure of cadmium toxicity (Blechinger et al., 2002).
A method of controlling temporal and spatial gene expression of an injected construct is RNA caging (Ando et al., 2001). This technique uses compounds such as 6-bromo-4-diazomethyl-7-hydroxycoumarin (Bhc-diazo) to reduce translational activity by forming covalent bonds with the phosphate moiety of the sugar-phosphate backbone. Because Bhc-caged mRNA can be reactivated via illumination with UV light of a specific wavelength, which causes photolysis (uncaging), a partial recovery of translational activity can be obtained on demand. Therefore, the expression of gene constructs in specific tissues would be dependent on the time and accuracy of illumination. This was demonstrated by the photo-illumination-induced ectopic expression of engrailed2a (Ando et al., 2001) at the shield stage of zebrafish embryos, that led to a severe reduction in eye size.
Morpholino strategies
Knockdown technology (reverse genetics) has improved greatly from the initial antisense oligos that had nonspecific toxic side effects and required continued treatment to ensure mRNA degradation (Cazenave et al., 1989; Heasman et al., 1991). Morpholino oligonucleotides (MOs) are antisense nucleic acid analogs that have ribosides converted to morpholines (C4H9NO) and a phosphorodiamidate intersubunit linkage instead of phosphorodiester linkage (Summerton and Weller, 1997). They work by binding to, and blocking translation of specific mRNA. MOs are one of the most powerful modern techniques and valuable assets for zebrafish as a model organism. MOs have been shown to successfully knockdown gene expression in zebrafish embryos (Nasevicius and Ekker, 2001) by injecting complementary sequences to either the 25 bases 3′ to the AUG translational start site or the 5′ leader sequence of a gene of interest, at the 1- to 4-cell stages into the yolk or cytoplasm. In fact, the majority of studies with MO assays have been carried out in the zebrafish, and together with the vast catalog of mutant zebrafish, have proven invaluable for the functional definition of genes required for normal development. However, although an even distribution of MO through the zebrafish has been observed after injection (Nasevicius and Ekker, 2000), it is important to stress that phenotypes induced by MOs can vary considerably in severity, and injection of the oligo can cause nonspecific adverse effects (Heasman, 2002). The appropriate threshold for MO concentration must be determined to effectively block translation without causing overt toxicity.
Targeting MOs against specific genes essential in development or detoxification can create morphants that are phenotypically similar to those of mutant zebrafish embryos and embryos exhibiting signs of toxicity or disease (Guyon et al., 2003; Nasevicius and Ekker, 2000). This may help elucidate which gene pathways have been affected before intensive molecular investigations are performed. Likewise, a MO directed against a gene that mediates toxicity of a particular chemical can be used to prevent that toxicity.
Mutant Zebrafish and Their Use in Studying Toxicity and Human Diseases
Phenotypic comparisons between zebrafish manifesting chemical-specific endpoints of toxicity and zebrafish mutants may provide insight into the specific genes affected by the toxicant. Likewise, zebrafish carrying a certain mutation may be resistant to a particular toxicant and hence may help demonstrate the necessity for a specific developmental or detoxification pathway in mediating the toxic response.
One main advantage of mutagenesis is the selection of genes that have at least partially nonredundant functions. In addition, genes identified in the zebrafish model, especially those from gastrulation onwards, are well conserved among other vertebrates, enabling easy access to their homologs (Haffter et al., 1996). There are however, limitations to using mutants as diagnostics tools. Generally mutant screens in the past only detected mutations that exhibited a clearly visible phenotype. As a result, many forward genetic screens now invest more time in using ISH and IHC to aid the discovery of more subtle effects.
N-ethyl-N-nitrosurea (ENU) is effective in inducing random point mutations in zebrafish and has led to a plethora of mutations in genes essential for numerous processes including morphogenesis, pattern formation, organogenesis, and differentiation. Hundreds of zebrafish mutants have been identified (Haffter et al., 1996), and the list continues to grow. After ENU mutagenesis, instead of assessing each mutant, specific genes can be targeted via a high-throughput reverse genetic screen know as TILLING (Targeted Induced Local Lesions IN Genomes) (Wienholds et al., 2002). Further, insertional mutagenesis, in which retroviral vectors disrupt gene sequences, has also contributed significantly to zebrafish screening (Amsterdam et al., 1999; Golling et al., 2002). Many of these characterized mutations can be located on the Zebrafish Information Network (ZFIN) website.
As well as being useful to identify genes affected by the exposure to various toxicants, mutant models are useful for studying human diseases (Amatruda et al., 2002; Barut and Zon, 2000; Dodd et al., 2000; Dooley and Zon, 2000; Knapik, 2000; Ward and Lieschke, 2002; van Heyningen, 1997; Vascotto et al., 1997; Zon, 1999). Although a full list of mutants and associated diseases is beyond the scope of this review, the kind and extent of the growing resources available can be highlighted by focusing on the mutants available for a single organ. The heart is the first organ to develop and function in zebrafish (Lee et al., 1994; Stainier et al., 1993). The zebrafish heart has been used as a model for human cardiovascular diseases and also is commonly affected during toxicity studies, usually manifesting as bradycardia or an arrhythmia. The following includes a variety of mutant phenotypes that demonstrate altered heart development. Some of these mutations have been associated with human diseases, and others may show similarities with zebrafish exposed to cardiotoxic chemicals. Lonely atrium has no ventricle; bypass, sonic-you, you-too, you, kurzschluss, and chameleon show circulatory system defects (Chen et al., 1996; van Eeden et al., 1996); viper, weak atrium, heart attack, slop, slinky, and still heart have weakened contraction in one or both heart chambers (Chen et al., 1996; Granato et al., 1996; Jiang et al., 1996; Stainier et al., 1996); and other mutants such as breakdance, hiphop, leglong and slip jig have an abnormal heart rhythm (Chen et al, 1996). Milesapart, faust, casanova, and natter, and one-eyed pinhead have a cardio bifida (two hearts) phenotype (Jiang et al., 1996; Stainier et al., 1996); pikwik has a heart contraction defect with the mutated gene titin that is found in human cardiomyopathy (Gerull et al., 2002; Stainier et al., 1996; Xu et al., 2002); silent heart has no heartbeat, with a mutation in the gene tnnt2, also involved in cardiomyopathy (Chen et al., 1996; Sehnert et al., 2002); and heartstrings has differentiation defects with a mutation in the tbx5 gene found in Holt-Oram syndrome (Garrity et al., 2002). Chemicals adversely affecting heart formation have been screened on a large scale (Peterson et al., 2001). One such chemical, concentramide, showed the heart phenotype of the ventricle forming inside the atrium, just as in the heart and soul mutant.
CELL CULTURE AND MICROARRAY TECHNOLOGY
Primary and immortal cell lines have been established from adult and embryonic zebrafish (Collodi et al., 1992a, 1992b; Ghosh and Collodi, 1994; Ghosh et al., 1994; Helmrich and Barnes, 1999). In addition, non-zebrafish cell lines such as COS-7 cells transfected with different zebrafish AHRs, ARNTs, and a dioxin response element-driven luciferase reporter have been used to investigate different halogenated aromatic hydrocarbons for AHR agonist activity in transactivation assays (Andreasen et al., 2002a; Tanguay et al., 1999).
Microarrays have been used most powerfully when applied to genomically characterized model species. As the zebrafish genome is near completion, this should soon become a more common and valued resource to provide a comprehensive view of regulated gene expression in response to a given toxicant. The potential also exists for uncovering novel systems. Several companies including NimbleGen, Affymetrix, MWG, and Compugen have created microarray chips that are now commercially available. Large-scale genomic screens for chemically-induced transcriptional disruption can therefore already be performed in zebrafish.
ACUTE TOXICITY AND ADVERSE EFFECTS TO THE JUVENILE AND ADULT ZEBRAFISH
Acute toxicity studies using zebrafish are limited. Examples of toxicants investigated include lead and uranium (Labrot et al., 1999), malathion (Kumar and Ansari, 1986), metronidazole (Lanzky and Halling-Sorensen, 1997), anilines (Zok et al., 1991), and colchicines (Roche et al., 1994).
Apparently the main reason few studies have utilized juvenile and adult zebrafish is because the overarching value of zebrafish lies in its genetics and developmental biology. Several studies have exposed embryos to graded concentrations of a chemical and assessed endpoints of toxicity as juveniles. For example, hatching, morphological abnormalities and survival were assessed for up to 35 days post fertilization (dpf) in zebrafish exposed to lindane, deltamethrin, and atrazine (Gorge and Nagel, 1990). Effects on sex differentiation were also assessed for 17α-ethinylestradiol (Andersen et al., 2003). Likewise 17β-ethinylestradiol altered sex differentiation in the juvenile, caused modified sexual characteristics in the adult male, and inhibited egg production (Brion et al., 2004). Many other studies have been performed to investigate carcinogenicity during later stages (review, Spitsbergen and Kent, 2003).
THINGS TO CONSIDER BEFORE USING THE ZEBRAFISH MODEL
The types of toxicity studies in which zebrafish have been used are shown in Table 1. The studies include acute, subchronic, and chronic toxicity studies as well as hypothesis-driven studies on mechanisms of target organ toxicity.
Types of Toxicity Investigations Using Zebrafish
Reproductive toxicity |
| Developmental toxicity |
| Acute toxicity |
| Neurotoxicity |
| Cardiotoxicity |
| Ocular toxicity |
| Endocrine disruption |
| Neurobehavioral toxicity |
| Vascular toxicity |
| Carcinogenicity |
Reproductive toxicity |
| Developmental toxicity |
| Acute toxicity |
| Neurotoxicity |
| Cardiotoxicity |
| Ocular toxicity |
| Endocrine disruption |
| Neurobehavioral toxicity |
| Vascular toxicity |
| Carcinogenicity |
Types of Toxicity Investigations Using Zebrafish
Reproductive toxicity |
| Developmental toxicity |
| Acute toxicity |
| Neurotoxicity |
| Cardiotoxicity |
| Ocular toxicity |
| Endocrine disruption |
| Neurobehavioral toxicity |
| Vascular toxicity |
| Carcinogenicity |
Reproductive toxicity |
| Developmental toxicity |
| Acute toxicity |
| Neurotoxicity |
| Cardiotoxicity |
| Ocular toxicity |
| Endocrine disruption |
| Neurobehavioral toxicity |
| Vascular toxicity |
| Carcinogenicity |
Mechanisms of developmental toxicity have been partially explained for only a few toxicants, and there is no chemical for which it is fully explained. There have been several individual and groups of chemicals investigated for causing developmental toxicity in zebrafish. Most studies have concentrated on polyaromatic chemicals, pesticides, endocrine disruptors, retinoic acid, and cyclopamine (Table 2), but research investigating the adverse developmental effects of 2,3,7,8-tetrachlorodibenzo-p-dioxin (TCDD) has been most comprehensive. As a guide for using the zebrafish to investigate the mechanism of toxicity of other chemicals, the process used to validate the zebrafish model for investigating TCDD toxicity is instructive.
Classes of Chemicals Evaluated for Toxicity in Zebrafish
Chemical classes . | Type of toxicity study . | Reference . |
|---|---|---|
| Metals | ||
| Copper and zinc | Susceptibily to infection by listeria after exposure to metals | Rougier et al., 1996 |
| Copper, nickel, mercury, colbalt, lead | Toxicity: dose response, effects on hatching and survival | Dave and Xiu, 1991 |
| Methylmercury | Functional impairment and delayed mortality syndrome | Samson et al., 2001 |
| Aluminium, cadmium, iron | Toxicity: dose response, effects on hatching and survival | Dave, 1985 |
| Lead and uranium | Acute toxicity and toxicokinetics | Labrot et al., 1999 |
| Cadmium | Ectopic apoptosis induction | Chan and Cheng, 2003 |
| Cadmium | Abnormal somite patterning and defects in axonogenesis | Hen Chow and Cheng, 2003 |
| Tributyltin | Reproductive toxicity | McAllister and Kime, 2003 |
| PCBs/PAHs | ||
| PCBs | Transactivation activity of aryl hydrocarbon receptors in COS-7 | Abnet et al., 1999 |
| PCBs | CYP1A induction in a zebrafish liver cell line | Henry et al., 2001 |
| PCBs | Reproduction | Orn et al., 1998 |
| PCBs | Kinetics of bioconcentration clearance | Fox et al., 1994 |
| PCBs | Bioaccumulation with different routes of exposure | Andersson et al., 2001 |
| PAHs | Morphological abnormalities occurring after cardiac dysfunction | Incardona et al., 2004 |
| Retinoic acid | ||
| Retinoic acid | Abnormal pectoral fin bud morphology and ectopic shh expression | Akimenko and Ekker, 1995 |
| Retinoic acid | Abnormal development of the caudal midbrain and anterior hindbrain | Hill et al., 1995 |
| Retinoic acid | RA-mediated gene expression in transgenic reporter zebrafish | Perz-Edwards et al., 2001 |
| Retinoic acid | Pectoral fin duplications | Vandersea et al., 1998 |
| Cyclopamine (inhibitor of hedghog signaling) | ||
| Cyclopamine | Elimination of primary motoneurons | Chen et al., 2001 |
| Cyclopamine | Role of shh in the induction and patterning of the pituitary | Sbrogna et al., 2003 |
| Cyclopamine | Inhibition of fin outgrowth | Quint et al., 2002 |
| Cyclopamine | Role of hedgehog signaling in eye development | Stenkamp and Frey, 2003 |
| Fragrances/nitrated benzenes | ||
| Nitro musk–ketones and xylene | Effects on reproduction, mortality, and growth | Carlsson et al., 2000 |
| Nitro musk–ketones, xylene, AHTN, HHCB | Toxicity and mortality | Carlsson and Norrgren, 2004 |
| Nitro musk–AHTN, HHCB | Antiestrogenic effects | Schreurs et al., 2004 |
| Pesticides and Herbicides | ||
| Lindane, atrazine, and deltamethrin | Deformations, mortality, growth retardation and hatching rate | Gorge and Nagel, 1990 |
| Toxaphene | Toxicity, reproductive success and oviposition | Ree. and Payne, 1997 |
| Parathion | Acetylcholinesterase inhibition and food consumption rate | Roex et al., 2003 |
| Endosulfan | Primordial germ cell migration and distribution | Willey and Krone, 2001 |
| Sevin | Effects on reproduction and hatching | Todd and Van Leeuwen, 2002 |
| Chlorpyrifos | Effects on survival, response latency and spatial discrimination | Levin et al., 2003 |
| Atrazine (2-chloro-4-ethylamino-6-isopropylamine-s-triazine) | Morphological and functional abnormalities | Wiegand et al., 2001 |
| 3,4-dichloroaniline, lindane | Toxicity and effects on reproduction | Ensenbach and Nagel, 1997 |
| 4-chloroaniline | Effects on hatching and ultrastructural changes in liver and kidney | Oulmi and Braunbeck, 1996 |
| Estrogenics | ||
| 17-beta estradiol, diethylstilbestrol | Effects on mortality and hatching, consequences for CNS | Kishida et al., 2001 |
| Nonylphenol, ethinylestradiol, benzo[a]pyrene | CYP19 expression induction | Kazeto et al., 2004 |
| Nonylphenol | Primordial germ cell migration and distribution | Willey and Krone, 2001 |
| Nonylphenol, 17alpha-ethinylestradiol | Effects on sex ratio and breeding success | Hill and Janz, 2003 |
| Nonylphenol, 17beta-estradiol | Vitellogenin as an estrogenic biomarker | Van den Belt et al., 2003 |
| Phytosterols were isolated from wood and soy beans | Reproduction/altered sexual ratio | Nakari and Erkomaa, 2003 |
| Other investigations | ||
| Saxitoxin | Morphological abnormalities and sensorimotor deficits | Lefebvre et al., 2004 |
| 1,2,3-trichlorobenzene | Reproductive impairment by non-polar narcosis | Roex et al., 2001 |
| Ammonium perchlorate | Reproductive performance and thyroid follicle histology | Patino et al., 2003 |
| Flavopiridol, Brefeldin A, Neomycin, and caspase inhibitors | Bioassays for assessing toxicity, angiogenesis, and apoptosis | Parng et al., 2002 |
| 7,12-dimethylbenz[a]anthracene (DMBA) | Neoplasia | Spitsbergen et al., 2000 |
| Triphenyltin acetate | Effects on survival, hatching success, and liver ultrastructure | Strmac and Braunbeck, 1999 |
Chemical classes . | Type of toxicity study . | Reference . |
|---|---|---|
| Metals | ||
| Copper and zinc | Susceptibily to infection by listeria after exposure to metals | Rougier et al., 1996 |
| Copper, nickel, mercury, colbalt, lead | Toxicity: dose response, effects on hatching and survival | Dave and Xiu, 1991 |
| Methylmercury | Functional impairment and delayed mortality syndrome | Samson et al., 2001 |
| Aluminium, cadmium, iron | Toxicity: dose response, effects on hatching and survival | Dave, 1985 |
| Lead and uranium | Acute toxicity and toxicokinetics | Labrot et al., 1999 |
| Cadmium | Ectopic apoptosis induction | Chan and Cheng, 2003 |
| Cadmium | Abnormal somite patterning and defects in axonogenesis | Hen Chow and Cheng, 2003 |
| Tributyltin | Reproductive toxicity | McAllister and Kime, 2003 |
| PCBs/PAHs | ||
| PCBs | Transactivation activity of aryl hydrocarbon receptors in COS-7 | Abnet et al., 1999 |
| PCBs | CYP1A induction in a zebrafish liver cell line | Henry et al., 2001 |
| PCBs | Reproduction | Orn et al., 1998 |
| PCBs | Kinetics of bioconcentration clearance | Fox et al., 1994 |
| PCBs | Bioaccumulation with different routes of exposure | Andersson et al., 2001 |
| PAHs | Morphological abnormalities occurring after cardiac dysfunction | Incardona et al., 2004 |
| Retinoic acid | ||
| Retinoic acid | Abnormal pectoral fin bud morphology and ectopic shh expression | Akimenko and Ekker, 1995 |
| Retinoic acid | Abnormal development of the caudal midbrain and anterior hindbrain | Hill et al., 1995 |
| Retinoic acid | RA-mediated gene expression in transgenic reporter zebrafish | Perz-Edwards et al., 2001 |
| Retinoic acid | Pectoral fin duplications | Vandersea et al., 1998 |
| Cyclopamine (inhibitor of hedghog signaling) | ||
| Cyclopamine | Elimination of primary motoneurons | Chen et al., 2001 |
| Cyclopamine | Role of shh in the induction and patterning of the pituitary | Sbrogna et al., 2003 |
| Cyclopamine | Inhibition of fin outgrowth | Quint et al., 2002 |
| Cyclopamine | Role of hedgehog signaling in eye development | Stenkamp and Frey, 2003 |
| Fragrances/nitrated benzenes | ||
| Nitro musk–ketones and xylene | Effects on reproduction, mortality, and growth | Carlsson et al., 2000 |
| Nitro musk–ketones, xylene, AHTN, HHCB | Toxicity and mortality | Carlsson and Norrgren, 2004 |
| Nitro musk–AHTN, HHCB | Antiestrogenic effects | Schreurs et al., 2004 |
| Pesticides and Herbicides | ||
| Lindane, atrazine, and deltamethrin | Deformations, mortality, growth retardation and hatching rate | Gorge and Nagel, 1990 |
| Toxaphene | Toxicity, reproductive success and oviposition | Ree. and Payne, 1997 |
| Parathion | Acetylcholinesterase inhibition and food consumption rate | Roex et al., 2003 |
| Endosulfan | Primordial germ cell migration and distribution | Willey and Krone, 2001 |
| Sevin | Effects on reproduction and hatching | Todd and Van Leeuwen, 2002 |
| Chlorpyrifos | Effects on survival, response latency and spatial discrimination | Levin et al., 2003 |
| Atrazine (2-chloro-4-ethylamino-6-isopropylamine-s-triazine) | Morphological and functional abnormalities | Wiegand et al., 2001 |
| 3,4-dichloroaniline, lindane | Toxicity and effects on reproduction | Ensenbach and Nagel, 1997 |
| 4-chloroaniline | Effects on hatching and ultrastructural changes in liver and kidney | Oulmi and Braunbeck, 1996 |
| Estrogenics | ||
| 17-beta estradiol, diethylstilbestrol | Effects on mortality and hatching, consequences for CNS | Kishida et al., 2001 |
| Nonylphenol, ethinylestradiol, benzo[a]pyrene | CYP19 expression induction | Kazeto et al., 2004 |
| Nonylphenol | Primordial germ cell migration and distribution | Willey and Krone, 2001 |
| Nonylphenol, 17alpha-ethinylestradiol | Effects on sex ratio and breeding success | Hill and Janz, 2003 |
| Nonylphenol, 17beta-estradiol | Vitellogenin as an estrogenic biomarker | Van den Belt et al., 2003 |
| Phytosterols were isolated from wood and soy beans | Reproduction/altered sexual ratio | Nakari and Erkomaa, 2003 |
| Other investigations | ||
| Saxitoxin | Morphological abnormalities and sensorimotor deficits | Lefebvre et al., 2004 |
| 1,2,3-trichlorobenzene | Reproductive impairment by non-polar narcosis | Roex et al., 2001 |
| Ammonium perchlorate | Reproductive performance and thyroid follicle histology | Patino et al., 2003 |
| Flavopiridol, Brefeldin A, Neomycin, and caspase inhibitors | Bioassays for assessing toxicity, angiogenesis, and apoptosis | Parng et al., 2002 |
| 7,12-dimethylbenz[a]anthracene (DMBA) | Neoplasia | Spitsbergen et al., 2000 |
| Triphenyltin acetate | Effects on survival, hatching success, and liver ultrastructure | Strmac and Braunbeck, 1999 |
Classes of Chemicals Evaluated for Toxicity in Zebrafish
Chemical classes . | Type of toxicity study . | Reference . |
|---|---|---|
| Metals | ||
| Copper and zinc | Susceptibily to infection by listeria after exposure to metals | Rougier et al., 1996 |
| Copper, nickel, mercury, colbalt, lead | Toxicity: dose response, effects on hatching and survival | Dave and Xiu, 1991 |
| Methylmercury | Functional impairment and delayed mortality syndrome | Samson et al., 2001 |
| Aluminium, cadmium, iron | Toxicity: dose response, effects on hatching and survival | Dave, 1985 |
| Lead and uranium | Acute toxicity and toxicokinetics | Labrot et al., 1999 |
| Cadmium | Ectopic apoptosis induction | Chan and Cheng, 2003 |
| Cadmium | Abnormal somite patterning and defects in axonogenesis | Hen Chow and Cheng, 2003 |
| Tributyltin | Reproductive toxicity | McAllister and Kime, 2003 |
| PCBs/PAHs | ||
| PCBs | Transactivation activity of aryl hydrocarbon receptors in COS-7 | Abnet et al., 1999 |
| PCBs | CYP1A induction in a zebrafish liver cell line | Henry et al., 2001 |
| PCBs | Reproduction | Orn et al., 1998 |
| PCBs | Kinetics of bioconcentration clearance | Fox et al., 1994 |
| PCBs | Bioaccumulation with different routes of exposure | Andersson et al., 2001 |
| PAHs | Morphological abnormalities occurring after cardiac dysfunction | Incardona et al., 2004 |
| Retinoic acid | ||
| Retinoic acid | Abnormal pectoral fin bud morphology and ectopic shh expression | Akimenko and Ekker, 1995 |
| Retinoic acid | Abnormal development of the caudal midbrain and anterior hindbrain | Hill et al., 1995 |
| Retinoic acid | RA-mediated gene expression in transgenic reporter zebrafish | Perz-Edwards et al., 2001 |
| Retinoic acid | Pectoral fin duplications | Vandersea et al., 1998 |
| Cyclopamine (inhibitor of hedghog signaling) | ||
| Cyclopamine | Elimination of primary motoneurons | Chen et al., 2001 |
| Cyclopamine | Role of shh in the induction and patterning of the pituitary | Sbrogna et al., 2003 |
| Cyclopamine | Inhibition of fin outgrowth | Quint et al., 2002 |
| Cyclopamine | Role of hedgehog signaling in eye development | Stenkamp and Frey, 2003 |
| Fragrances/nitrated benzenes | ||
| Nitro musk–ketones and xylene | Effects on reproduction, mortality, and growth | Carlsson et al., 2000 |
| Nitro musk–ketones, xylene, AHTN, HHCB | Toxicity and mortality | Carlsson and Norrgren, 2004 |
| Nitro musk–AHTN, HHCB | Antiestrogenic effects | Schreurs et al., 2004 |
| Pesticides and Herbicides | ||
| Lindane, atrazine, and deltamethrin | Deformations, mortality, growth retardation and hatching rate | Gorge and Nagel, 1990 |
| Toxaphene | Toxicity, reproductive success and oviposition | Ree. and Payne, 1997 |
| Parathion | Acetylcholinesterase inhibition and food consumption rate | Roex et al., 2003 |
| Endosulfan | Primordial germ cell migration and distribution | Willey and Krone, 2001 |
| Sevin | Effects on reproduction and hatching | Todd and Van Leeuwen, 2002 |
| Chlorpyrifos | Effects on survival, response latency and spatial discrimination | Levin et al., 2003 |
| Atrazine (2-chloro-4-ethylamino-6-isopropylamine-s-triazine) | Morphological and functional abnormalities | Wiegand et al., 2001 |
| 3,4-dichloroaniline, lindane | Toxicity and effects on reproduction | Ensenbach and Nagel, 1997 |
| 4-chloroaniline | Effects on hatching and ultrastructural changes in liver and kidney | Oulmi and Braunbeck, 1996 |
| Estrogenics | ||
| 17-beta estradiol, diethylstilbestrol | Effects on mortality and hatching, consequences for CNS | Kishida et al., 2001 |
| Nonylphenol, ethinylestradiol, benzo[a]pyrene | CYP19 expression induction | Kazeto et al., 2004 |
| Nonylphenol | Primordial germ cell migration and distribution | Willey and Krone, 2001 |
| Nonylphenol, 17alpha-ethinylestradiol | Effects on sex ratio and breeding success | Hill and Janz, 2003 |
| Nonylphenol, 17beta-estradiol | Vitellogenin as an estrogenic biomarker | Van den Belt et al., 2003 |
| Phytosterols were isolated from wood and soy beans | Reproduction/altered sexual ratio | Nakari and Erkomaa, 2003 |
| Other investigations | ||
| Saxitoxin | Morphological abnormalities and sensorimotor deficits | Lefebvre et al., 2004 |
| 1,2,3-trichlorobenzene | Reproductive impairment by non-polar narcosis | Roex et al., 2001 |
| Ammonium perchlorate | Reproductive performance and thyroid follicle histology | Patino et al., 2003 |
| Flavopiridol, Brefeldin A, Neomycin, and caspase inhibitors | Bioassays for assessing toxicity, angiogenesis, and apoptosis | Parng et al., 2002 |
| 7,12-dimethylbenz[a]anthracene (DMBA) | Neoplasia | Spitsbergen et al., 2000 |
| Triphenyltin acetate | Effects on survival, hatching success, and liver ultrastructure | Strmac and Braunbeck, 1999 |
Chemical classes . | Type of toxicity study . | Reference . |
|---|---|---|
| Metals | ||
| Copper and zinc | Susceptibily to infection by listeria after exposure to metals | Rougier et al., 1996 |
| Copper, nickel, mercury, colbalt, lead | Toxicity: dose response, effects on hatching and survival | Dave and Xiu, 1991 |
| Methylmercury | Functional impairment and delayed mortality syndrome | Samson et al., 2001 |
| Aluminium, cadmium, iron | Toxicity: dose response, effects on hatching and survival | Dave, 1985 |
| Lead and uranium | Acute toxicity and toxicokinetics | Labrot et al., 1999 |
| Cadmium | Ectopic apoptosis induction | Chan and Cheng, 2003 |
| Cadmium | Abnormal somite patterning and defects in axonogenesis | Hen Chow and Cheng, 2003 |
| Tributyltin | Reproductive toxicity | McAllister and Kime, 2003 |
| PCBs/PAHs | ||
| PCBs | Transactivation activity of aryl hydrocarbon receptors in COS-7 | Abnet et al., 1999 |
| PCBs | CYP1A induction in a zebrafish liver cell line | Henry et al., 2001 |
| PCBs | Reproduction | Orn et al., 1998 |
| PCBs | Kinetics of bioconcentration clearance | Fox et al., 1994 |
| PCBs | Bioaccumulation with different routes of exposure | Andersson et al., 2001 |
| PAHs | Morphological abnormalities occurring after cardiac dysfunction | Incardona et al., 2004 |
| Retinoic acid | ||
| Retinoic acid | Abnormal pectoral fin bud morphology and ectopic shh expression | Akimenko and Ekker, 1995 |
| Retinoic acid | Abnormal development of the caudal midbrain and anterior hindbrain | Hill et al., 1995 |
| Retinoic acid | RA-mediated gene expression in transgenic reporter zebrafish | Perz-Edwards et al., 2001 |
| Retinoic acid | Pectoral fin duplications | Vandersea et al., 1998 |
| Cyclopamine (inhibitor of hedghog signaling) | ||
| Cyclopamine | Elimination of primary motoneurons | Chen et al., 2001 |
| Cyclopamine | Role of shh in the induction and patterning of the pituitary | Sbrogna et al., 2003 |
| Cyclopamine | Inhibition of fin outgrowth | Quint et al., 2002 |
| Cyclopamine | Role of hedgehog signaling in eye development | Stenkamp and Frey, 2003 |
| Fragrances/nitrated benzenes | ||
| Nitro musk–ketones and xylene | Effects on reproduction, mortality, and growth | Carlsson et al., 2000 |
| Nitro musk–ketones, xylene, AHTN, HHCB | Toxicity and mortality | Carlsson and Norrgren, 2004 |
| Nitro musk–AHTN, HHCB | Antiestrogenic effects | Schreurs et al., 2004 |
| Pesticides and Herbicides | ||
| Lindane, atrazine, and deltamethrin | Deformations, mortality, growth retardation and hatching rate | Gorge and Nagel, 1990 |
| Toxaphene | Toxicity, reproductive success and oviposition | Ree. and Payne, 1997 |
| Parathion | Acetylcholinesterase inhibition and food consumption rate | Roex et al., 2003 |
| Endosulfan | Primordial germ cell migration and distribution | Willey and Krone, 2001 |
| Sevin | Effects on reproduction and hatching | Todd and Van Leeuwen, 2002 |
| Chlorpyrifos | Effects on survival, response latency and spatial discrimination | Levin et al., 2003 |
| Atrazine (2-chloro-4-ethylamino-6-isopropylamine-s-triazine) | Morphological and functional abnormalities | Wiegand et al., 2001 |
| 3,4-dichloroaniline, lindane | Toxicity and effects on reproduction | Ensenbach and Nagel, 1997 |
| 4-chloroaniline | Effects on hatching and ultrastructural changes in liver and kidney | Oulmi and Braunbeck, 1996 |
| Estrogenics | ||
| 17-beta estradiol, diethylstilbestrol | Effects on mortality and hatching, consequences for CNS | Kishida et al., 2001 |
| Nonylphenol, ethinylestradiol, benzo[a]pyrene | CYP19 expression induction | Kazeto et al., 2004 |
| Nonylphenol | Primordial germ cell migration and distribution | Willey and Krone, 2001 |
| Nonylphenol, 17alpha-ethinylestradiol | Effects on sex ratio and breeding success | Hill and Janz, 2003 |
| Nonylphenol, 17beta-estradiol | Vitellogenin as an estrogenic biomarker | Van den Belt et al., 2003 |
| Phytosterols were isolated from wood and soy beans | Reproduction/altered sexual ratio | Nakari and Erkomaa, 2003 |
| Other investigations | ||
| Saxitoxin | Morphological abnormalities and sensorimotor deficits | Lefebvre et al., 2004 |
| 1,2,3-trichlorobenzene | Reproductive impairment by non-polar narcosis | Roex et al., 2001 |
| Ammonium perchlorate | Reproductive performance and thyroid follicle histology | Patino et al., 2003 |
| Flavopiridol, Brefeldin A, Neomycin, and caspase inhibitors | Bioassays for assessing toxicity, angiogenesis, and apoptosis | Parng et al., 2002 |
| 7,12-dimethylbenz[a]anthracene (DMBA) | Neoplasia | Spitsbergen et al., 2000 |
| Triphenyltin acetate | Effects on survival, hatching success, and liver ultrastructure | Strmac and Braunbeck, 1999 |
The long-range goal was to develop the zebrafish as a model organism to identify AHR-regulated genes that mediate specific endpoints of TCDD toxicity. Yet, when the project was started, it was not known how or if zebrafish would respond to TCDD, what the endpoints of developmental toxicity would be, if the AHR signaling pathway in zebrafish would be similar to mammals, which PAS proteins (AHR1 vs. AHR2 and ARNT1 vs. ARNT2) were the key dimerization partners that mediated TCDD toxicity, and whether CYP1A induced by TCDD mediated toxicity.
How Do Zebrafish Respond to TCDD Exposure?
Fish are particularly sensitive to chemical exposure, especially during early development (Peterson et al., 1993, Walker and Peterson, 1994a). Zebrafish were no exception. After waterborne exposure to TCDD as newly fertilized eggs, they developed a severe phenotype consisting of pericardial and yolk sac edema, reduced heart size, impaired cardiovascular function resulting in reduced cardiac output and ischemia of peripheral tissues, impaired jaw development, hemorrhage, and uninflated swimbladder culminating in mortality. LC50 for zebrafish embryos exposed via water were reported as 2.5 ng TCDD/g egg (Henry et al., 1997) and 2.6 ng TCDD/g egg (Elonen et al., 1998) at 10 and 32 days postfertilization, respectively. Compared to embryos of over ten other species of fish (Elonen et al., 1998; Helder, 1980, 1991; Prince and Cooper, 1989; Walker and Peterson, 1991, 1994b; Walker et al., 1992; Wisk and Cooper, 1990a,b ), zebrafish embryos were the least sensitive to the adverse developmental effects of TCDD, whereas the most sensitive species were the lake and bull trout (Tanguay et al., 2003; Walker et al., 1991, 1992). As the least sensitive species, TCDD-exposed embryos of zebrafish, when compared to embryos of lake trout, may provide insight into the mechanistic basis for differences in sensitivity to TCDD developmental toxicity between fish species.
Endpoints of TCDD toxicity, manifested in the early life stages of zebrafish, are shown in Table 3. Likewise mortality, cardiovascular dysfunction, edema, and hemorrhages have also been reported in birds and mammals (Allen et al., 1977; Firestone, 1973; Ivnitski et al., 2001; Schwetz et al., 1973; Vos et al., 1974). Although there are great similarities in the endpoints of TCDD toxicity between embryos of different freshwater fish species (Tanguay et al., 2003), there are some differences. For example, physiological processes associated with hatching were unaffected in the zebrafish (Henry et al., 1997), whereas a low incidence of half-hatching was recorded for rainbow trout (Walker and Peterson, 1991). As rainbow trout are more mature than zebrafish at hatching, toxicants may have a greater impact on the ability of trout to hatch successfully. Manifestation of adverse effects of TCDD in the zebrafish are dependent on the developmental stage of exposure. Zebrafish exposed to TCDD immediately after fertilization develop edema as early as 72 hpf (Henry et al., 1997). Interestingly, if exposure to TCDD is delayed until after 96 hpf, edema is not observed (Belair et al., 2001). This suggests that developing zebrafish are especially vulnerable to TCDD shortly after hatching.
Endpoints of TCDD Developmental Toxicity in Zebrafish
Endpoints of toxicity . | References . |
|---|---|
| Mortality | a, b |
| Arrested growth | a, b |
| Craniofacial malformations | a, b, c, d |
| Yolk sac, pericardial and meningeal edema | a, b, d, e, f, g |
| Peripheral ischemia and disruption of erythropoiesis | a, f, h, i |
| Arrested gill development | a |
| Impaired swimbladder inflation | a |
| Altered apoptosis | h, j |
| Decreased number of neurons in the brain | j |
| Inhibition of fin regeneration | k |
| Inhibition of common cardinal vein regression | l |
| Reduced heart size and ventricular standstill | m |
Endpoints of toxicity . | References . |
|---|---|
| Mortality | a, b |
| Arrested growth | a, b |
| Craniofacial malformations | a, b, c, d |
| Yolk sac, pericardial and meningeal edema | a, b, d, e, f, g |
| Peripheral ischemia and disruption of erythropoiesis | a, f, h, i |
| Arrested gill development | a |
| Impaired swimbladder inflation | a |
| Altered apoptosis | h, j |
| Decreased number of neurons in the brain | j |
| Inhibition of fin regeneration | k |
| Inhibition of common cardinal vein regression | l |
| Reduced heart size and ventricular standstill | m |
Note. a: Henry et al., 1997; b: Elonen et al., 1998; c: Teraoka et al., 2002; d: Hill et al. 2004b; e: Prasch et al., 2003; f: Belair et al., 2001; g: Hill et al., 2004a; h: Dong et al., 2002; i: Dong et al., 2004; j: Hill et al., 2003; k: Zodrow et al., 2003; l: Bello et al., 2004; m: Antkiewicz et al., in press.
Endpoints of TCDD Developmental Toxicity in Zebrafish
Endpoints of toxicity . | References . |
|---|---|
| Mortality | a, b |
| Arrested growth | a, b |
| Craniofacial malformations | a, b, c, d |
| Yolk sac, pericardial and meningeal edema | a, b, d, e, f, g |
| Peripheral ischemia and disruption of erythropoiesis | a, f, h, i |
| Arrested gill development | a |
| Impaired swimbladder inflation | a |
| Altered apoptosis | h, j |
| Decreased number of neurons in the brain | j |
| Inhibition of fin regeneration | k |
| Inhibition of common cardinal vein regression | l |
| Reduced heart size and ventricular standstill | m |
Endpoints of toxicity . | References . |
|---|---|
| Mortality | a, b |
| Arrested growth | a, b |
| Craniofacial malformations | a, b, c, d |
| Yolk sac, pericardial and meningeal edema | a, b, d, e, f, g |
| Peripheral ischemia and disruption of erythropoiesis | a, f, h, i |
| Arrested gill development | a |
| Impaired swimbladder inflation | a |
| Altered apoptosis | h, j |
| Decreased number of neurons in the brain | j |
| Inhibition of fin regeneration | k |
| Inhibition of common cardinal vein regression | l |
| Reduced heart size and ventricular standstill | m |
Note. a: Henry et al., 1997; b: Elonen et al., 1998; c: Teraoka et al., 2002; d: Hill et al. 2004b; e: Prasch et al., 2003; f: Belair et al., 2001; g: Hill et al., 2004a; h: Dong et al., 2002; i: Dong et al., 2004; j: Hill et al., 2003; k: Zodrow et al., 2003; l: Bello et al., 2004; m: Antkiewicz et al., in press.
Mediation of TCDD Toxicity
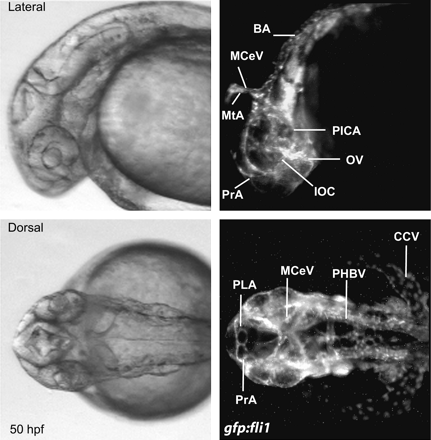
Certain polychlorinated dibenzo-p-dioxin (PCDD), dibenzofuran (PCDF), and biphenyl (PCB) congeners have been determined to exert toxicity through the AHR pathway. In comparison to mammals that have one AHR (reviews, Hahn, 2002; Hahn et al., 1997), zebrafish have two. AHR1 in zebrafish is an ortholog of the mammalian AHR, whereas AHR2 is a paralog to the type 1 receptors (Hahn et al., 1997). Multiple genes for ARNT, ARNT1, and ARNT2 have been identified in mammals (Drutel et al., 1996; Hirose et al., 1996; Hoffman et al., 1991; Li et al., 1994). Again due to the duplication event, an ARNT gene with multiple splice variants, zfARNT2b (Hsu et al., 2001; Tanguay et al., 2000) and recently zfARNT1 (Prasch et al., in press) has been identified in zebrafish. COS-7 cells have been used to investigate zebrafish genes via transactivation assays with a luciferase reporter. Such techniques have aided the discovery of AHR/ARNT heterodimers involved in TCDD toxicity (Andreasen et al., 2002a; Tanguay et al., 2000). zfAHR1 did dimerize with zfARNTs, but zfAHR2 exhibited a higher binding affinity to TCDD. ISH revealed zfahr2 mRNA is also more widely expressed than zfahr1 during early development, and is colocalized with zfarnt2a, b, and c in the same tissues as zfcyp1a (Andreasen et al., 2002b), a detoxification gene induced via the AHR pathway. Thus, zfAHR2 is considered to be the form of AHR involved in mediating TCDD developmental toxicity in zebrafish. Furthermore, although zfARNT2b may contribute to TCDD toxicity, zfARNT1 currently appears to be the far more essential transcription factor for this process. The conclusions were supported by the injection of a morpholino oligonucleotide (MO) against zfAHR2 preventing manifestation of TCDD toxicity in TCDD-exposed embryos (Fig. 2C) (Bello et al., 2004; Dong et al., 2004; Hill et al., 2004a; Prasch et al., 2003; Teraoka et al., 2003b). Likewise, zfARNT1 was shown to be the probable dimerization partner for zfAHR2 because a MO targeted against zfARNT1 rescued embryos from TCDD toxicity (Prasch et al., in press), whereas a MO targeted against zfARNT2 failed to protect against toxicity. Also, a zebrafish ARNT2 mutant still showed adverse effects when exposed to TCDD (Fig. 2D; Prasch et al., 2004).
Morpholino targeted against zfAHR2 rescued embryos from TCDD toxicity, whereas an ARNT2 mutant zebrafish shows the typical endpoints of TCDD toxicity. These results helped determine that AHR2 (not AHR1) and ARNT1 (not ARNT2) were the key dimerization partners required for AHR mediated TCDD developmental toxicity in zebrafish. Wild type embryos exposed to a vehicle control (A) and embryos injected with zfAHR2-MO exposed to 0.4 ng/ml TCDD (C) show no endpoints of TCDD toxicity, whereas wild type (B) and ARNT2 mutant embryos (D) exposed to 0.4 ng/ml TCDD show typical endpoints of toxicity such as P: pericardial edema, and C: craniofacial malformations. Embryos were exposed at 2 hpf for 1 h to TCDD and then allowed to develop in TCDD-free water prior to observation at 96 hpf.
The AHR/ARNT heterodimer causes alterations in gene expression by binding to xenobiotic response elements (XREs) upstream of target genes such as cyp1a (Schmidt and Bradfield, 1996). CYP1A antibodies have been used extensively to demonstrate induction in various tissues and especially in the zebrafish embryo vasculature (Andreasen et al., 2002b; Cantrell et al., 1998; Dong et al., 2002). Recently, knockdown of CYP1A with a MO failed to prevent several endpoints of TCDD developmental toxicity in zebrafish including reduced peripheral blood flow, pericardial edema, impaired jaw development, and disrupted erythropoiesis. These endpoints were assessed at three stages of development: 72, 96, and 144 hpf. So, although CYP1A can be a useful marker for AHR pathway induction, it was suggested that CYP1A was not involved in mediating these toxic responses to TCDD in zebrafish embryos (Carney et al., 2004). This is contrary to previous reports that suggested a role of CYP1A as a mediator of TCDD-induced toxicity in developing zebrafish. In the earlier studies, inhibitors for CYP1A and general CYPs rescued reduced blood flow in the dorsal midbrain of TCDD-treated zebrafish at 50 hpf (Dong et al., 2001, 2002) and knockdown of CYP1A with a MO rescued pericardial edema induced by TCDD at 72 hpf (Teraoka et al., 2003b).
Alternative Models to the Zebrafish
The closest model fish to zebrafish is medaka (Oryzias latipes). Medaka are of a similar size, mature after about two months and also lay large quantities of transparent eggs. The developmental stages of medaka were first published in English by Yamamoto (1975) and Kirchen and West (1976), although a more recent and more detailed version now also exists (Iwamatsu, 2004). Because medaka are eurythermal, they can survive a wide temperature range (0–40°C). The other main difference between them and the zebrafish is that they only have an expected lifespan of 1 year in the laboratory at 27°C. Although this would result in a large turnover of adults, they have been utilized for many years, particularly in Japan, and have proven to be a valuable resource. For example, they were adopted for carcinogenesis studies (Ishikawa et al., 1975; Matsushima and Sugimura, 1976) shortly after the first one performed with zebrafish (Stanton, 1965). They were the first organism used to demonstrate Y-linked inheritance (Aida, 1921) and provided much more information as a developmental and genetic model system (reviews: Ishikawa, 2000; Wittbrodt et al., 2002). Medaka was the first fish species in which a stable line of transgenics was established (Ozato et al., 1986). Transgenics produced today using the see-through mutation may have an advantage over those of the zebrafish because the transparent body of this line of fish allows effective visualization of the GFP expression pattern (Wakamatsu et al., 2001), and likewise, this mutant may also be useful for toxicity studies.
Complementary Model Fish Species
In contrast to the zebrafish, larger species such as the carp and goldfish, which are closely related to the zebrafish, may be more suited for experiments requiring larger amounts of tissue, such as gene expression profiling and proteomics, and are also more amenable to cell culture. Their size also permits greater access for anatomical investigations. Likewise, certain species have already been established as a good model for specific disciplines including toxicology. Fathead minnows, for example, have been used in a variety of toxicity studies, in particular those assessing endocrine disruption, and as such are noted for their regulatory acceptance both in the United States and the United Kingdom. Findings from these fish and other species that are genetically similar may be used to support those found in the zebrafish. The pufferfish (Fugu rubripes) has significant conservation of gene order, is closely related to the zebrafish, and although the genome is one-eighth the size of the human genome, it has a comparable number of genes due to the absence of “junk” DNA. Fugu has potential to provide important insights on genome organization and structure and gene function and regulation and, therefore, will continue to be an important complementary model organism for zebrafish as well as for numerous other species. As previously discussed, forward and reverse genetic screens in the medaka, together with further studies investigating organogenesis and embryonic patterning will provide valuable knowledge that can cross over into the zebrafish model. However, none of these other fish models alone provide the wide range of technical advantages afforded by the zebrafish.
CONCLUSION
The zebrafish, as a model vertebrate for toxicology, pharmacology, developmental biology, and genetics research, is only beginning. Research opportunities, such as the use of zebrafish in behavioral neuroscience, are in their infancy when compared to the use of laboratory rodents. As technology advances, mutant zebrafish, morpholinos, high-throughput screening and new bioassays for toxic and therapeutic endpoints in zebrafish will become more common. For toxicology, these advances, in addition to the accumulation of genetic and genomic infrastructure, will ultimately provide greater insight into the mechanisms of toxicity of chemicals, as well as aid in the discovery of new drugs for treating human disease.
We thank Drs. B. M. Weinstein and N. D. Lawson, Laboratory of Molecular Genetics, NICHD, NIH, Bethesda, MD, for providing the Fli1-eGFP transgenic zebrafish, and Sara Carney for providing the image of the zfAHR2 morphant. This work was supported by the University of Wisconsin Sea Grant Institute under grants from the National Sea Grant program, National Oceanic and Atmospheric Administration, U.S. Department of Commerce, Sea Grant Project numbers R/BT-16 and R/BT-17 (W.H. and R.E.P.), grants-in-aid for scientific research from the Japanese Ministry of Education, Culture, Sports, Science and Technology, and Gakujutsu-Frontier Cooperative Research from active research in Rakuno Gakuen University 2004–7 (H.T.)
References
Abnet, C. C., Tanguay, R. L., Heideman, W., and Peterson, R. E. (
Aida, T. (
Akimenko, M. A., and Ekker, M. (
Allen, J. R., Barsotti, D. A., Van Miller, J. P., Abrahamson, L. J., and Lalich, J. J. (
Amatruda, J. F., Shepard, J. L., Stern, H. M., and Zon, L. I. (
Amsterdam, A., Burgess, S., Golling, G., Chen, W., Sun, Z., Townsend, K., Farrington, S., Haldi, M., and Hopkins, N. (
Andersen, L., Holbech, H., Gessbo, A., Norrgren, L., and Petersen, G. I. (
Andersson, P. L., Berg, A. H., Bjerselius, R., Norrgren, L., Olsen, H., Olsson, P. E., Orn, S., and Tysklind, M. (
Ando, H., Furuta, T., Tsien, R.Y ., and Okamoto, H. (
Andreasen, E. A., Hahn, M. E., Heideman, W., Peterson, R. E., and Tanguay, R. L. (
Andreasen, E. A., Spitsbergen, J. M., Tanguay, R. L., Stegeman, J. J., Heideman, W., and Peterson, R. E. (
Antkiewicz, D. S., Burns, G. C., Carney, S. A., Peterson, R. E., and Heideman, W. (
Barut, B. A., and Zon, L. I. (
Belair, C. D., Peterson, R. E., and Heideman, W. (
Bello, S. M., Heideman, W., and Peterson, R. E. (
Blader, P., Fischer N., Gradwohl G., Guillemot G., and Strähle U. (
Blader, P., Plessy, C., and Strähle, U. (
Blechinger, S. R., Warren, J. T., Jr., Kuwada, J. Y., and Krone, P. H. (
Brion, F., Tyler, C. R., Palazzi, X., Laillet, B., Porcher, J. M., Garric, J., and Flammarion, P. (
Cantrell, S. M., Joy-Schlezinger, J., Stegeman, J. J., Tillitt, D. E., and Hannink, M. (
Carlsson, G., and Norrgren, L. (
Carlsson, G., Orn, S., Andersson, P. L., Soderstrom, H., and Norrgren, L. (
Carney, S. A., Peterson, R. E., and Heideman, W. (
Cazenave, C., Stein, C. A., Loreau, N., Thuong, N. T., Neckers, L. M., Subasinghe, C., Helene, C., Cohen, J. S., and Toulme, J. J. (
Chan, P. K., and Cheng, S. H. (
Chen, J. N., Haffter, P., Odenthal, J., Vogelsang, E., Brand, M., van Eeden, F. J., Furutani-Seiki, M., Granato, M., Hammerschmidt, M., Heisenberg, C. P., et al. (
Chen, W., Burgess, S., and Hopkins, N. (
Collodi, P., Kamei, Y., Ernst, T., Miranda, C., Buhler, D. R., and Barnes, D. W. (
Collodi, P., Kamei, Y., Sharps, A., Weber, D., and Barnes, D. (
Cooper, M. S., D'Amico, L. A., and Henry, C. A. (
Cooper, M. S., D'Amico, L. A., and Henry, C. A. (
Dave, G. (
Dave, G., and Xiu, R. Q. (
Dent, J. A., Polson, A. G., and Klymkowsky, M. W. (
Dodd, A., Curtis, P. M., Williams, L. C., and Love, D. R. (
Dooley, K., and Zon, L. I. (
Dong, W., Teraoka, H., Kondo, S., and Hiraga, T. (
Dong, W., Teraoka, H., Tsujimoto, Y., Stegeman, J. J., and Hiraga, T. (
Dong, W., Teraoka, H., Yamazaki, K., Tsukiyama, S., Imani, S., Imagawa, T., Stegeman, J. J., Peterson, R. E., and Hiraga, T. (
Driever, W., Stemple, D., Schier, A., and Solnica-Krezel, L. (
Drutel, G., Kathmann, M., Heron, A., Schwartz, J. C., and Arrang, J. M. (
Elonen, G. E., Spehar, R. L., Holcombe, G. W., Johnson, R. D., Fernandez, J. D., Erickson, R. J., Tietge, J. E., and Cook, P. M. (
Ensenbach, U., and Nagel, R. (
Fishman, M. C. (
Force, A., Lynch, M., Pickett, F. B., Amores, A., Yan, Y. L., and Postlethwait, J. (
Fox, K., Zauke, G. P., and Butte, W. (
Garrity, D. M., Childs, S., and Fishman, M. C. (
Gerull, B., Gramlich, M., Atherton, J., McNabb, M., Trombitas, K., Sasse-Klaassen, S., Seidman, J. G., Seidman, C., Granzier, H., Labeit, S., et al. (
Ghosh, C., and Collodi, P. (
Ghosh, C., Zhou, Y. L., and Collodi, P. (
Golling, G., Amsterdam, A., Sun, Z., Antonelli, M., Maldonado, E., Chen, W., Burgress, S., Haldi, M., Artzt, K., Farrington, S., et al. (
Gorge, G., and Nagel, R. (
Granato, M., van Eeden, F. J., Schach, U., Trowe, T., Brand, M., Furutani-Seiki, M., Haffter, P., Hammerschmidt, M., Heisenberg, C. P., Jiang, Y. J., et al. (
Guyon, J. R., Mosley, A. N., Zhou, Y., O'Brien, K. F., Sheng, X., Chiang, K., Davidson, A. J., Volinski, J. M., Zon, L. I., and Kunkel, L. M. (
Haffter, P., Granato, M., Brand, M., Mullins, M. C., Hammerschmidt, M., Kane, D. A., Odenthal, J., van Eeden, F. J., Jiang, Y. J., Heisenberg, C. P., et al. (
Hahn, M. E. (
Hahn, M. E., Karchner, S. I., Shapiro, M. A., and Perera, S. A. (
Halloran, M. C., Sato-Maeda, M., Warren, J. T., Su, F., Lele, Z., Krone, P. H., Kuwada, J. Y., and Shoji, W. (
Heasman, J., Holwill, S., and Wylie, C. C. (
Helder, T. (
Helder, T. (
Helmrich, A., and Barnes, D. (
Hen Chow, E. S., and Cheng, S. H. (
Henry, T. R., Nesbit, D. J., Heideman, W., and Peterson, R. E. (
Henry, T. R., Spitsbergen, J. M., Hornung, M. W., Abnet, C. C., and Peterson, R. E. (
Higashijima, S., Hotta, Y., and Okamoto, H. (
Hill, A. J., Bello, S. M., Prasch, A. L., Peterson, R. E., and Heideman, W. (
Hill, A. J., Howard, C. V., and Cossins, A. R. (
Hill, A., Howard C. V., and Cossins A. (
Hill, A., Howard, C. V., Strahle, U., and Cossins, A. (
Hill, J., Clarke, J. D., Vargesson, N., Jowett, T., and Holder, N. (
Hill, R. L., Jr., and Janz, D. M. (
Hirose, K., Morita, M., Ema, M., Mimura, J., Hamada, H., Fujii, H., Saijo, Y., Gotoh, O., Sogawa, K., and Fujii-Kuriyama, Y. (
Hoffman, E. C., Reyes, H., Chu, F. F., Sander, F., Conley, L. H., Brooks, B. A., and Hankinson, O. (
Holder, N., and Xu, Q. (
Hsu, H. J., Wang, W. D., and Hu, C. H. (
Incardona, J. P., Collier, T. K., and Scholz, N. L. (
Ishikawa, Y. (
Ishikawa, T., Shimamine, T., and Takayama, S. (
Ivnitski, I., Elmaoued, R., and Walker, M. K. (
Iwamatsu, T. (
Jacobson, J. L., Fein, G. G., Jacobson, S. W., Schwartz, P. M., and Dowler, J. K. (
Jiang, Y. J., Brand, M., Heisenberg, C. P., Beuchle, D., Furutani-Seiki, M., Kelsh, R. N., Warga, R. M., Granato, M., Haffter, P., Hammerschmidt, M., et al. (
Kazeto, Y., Place, A. R., and Trant, J. M. (
Kimmel, C. B., Ballard, W. W., Kimmel, S. R., Ullmann, B., and Schilling, T. T. (
Kimmel, C. B., and Warga, R. M. (
Kirchen, R. V., and West, W. R. (
Kishida, M., McLellan, M., Miranda, J. A., and Callard, G. V. (
Klymkowsky, M. W., and Hanken, J. (
Knapik, E. W. (
Krone, P. H., Lele, Z., and Sass, J. B. (
Kumar, K., and Ansari, B. A. (
Labrot, F., Narbonne, J. F., Ville, P., Saint Denis, M., and Ribera, D. (
Lanzky, P. F., and Halling-Sorensen, B. (
Lawson, N. D., and Weinstein, B. M. (
Lee, R. K. K., Stainier, D. Y. R., Weinstein, B. M., and Fishman, M. C. (
Lefebvre, K. A., Trainer, V. L., and Scholz, N. L. (
Levin, E. D., Chrysanthis, E., Yacisin, K., and Linney, E. (
Li, H., Dong, L., and Whitlock, J. P., Jr. (
Lin, S., Gaiano, N., Culp, P., Burns, J. C., Friedmann, T., Yee, J. K., and Hopkins, N. (
MacRae, C. A., and Peterson, R. T. (
Marra, M. A., Kucaba, T. A., Dietrich, N. L., Green, E. D., Brownstein, B., Wilson, R. K., McDonald, K. M., Hillier, L. W., McPherson, J. D., and Waterston, R. H. (
Matsushima, T., and Sugimura, T. (
Mattingly, C. J., McLachlan, J. A., and Toscano, W. A., Jr. (
McAllister, B. G., and Kime, D. E. (
Metscher, B. D., and Ahlberg, P. E. (
Milan, D. J., Peterson, T. A., Ruskin, J. N., Peterson, R. T., and MacRae, C. A. (
Nakari, T., and Erkomaa, K. (
Nasevicius, A., and Ekker, S. C. (
Nasevicius, A., and Ekker, S. C. (
Orn, S., Andersson, P. L., Forlin, L., Tysklind, M., and Norrgren, L. (
Oulmi, Y., and Braunbeck, T. (
Oxtoby, E., and Jowett, T. (
Ozato, K., Kondoh, H., Inohara, H., Iwamatsu, T., Wakamatsu, Y., and Okada, T. S. (
Parng, C., Seng, W. L., Semino, C., and McGrath, P. (
Patino, R., Wainscott, M. R., Cruz-Li, E. I., Balakrishnan, S., McMurry, C., Blazer, V. S., and Anderson, T. A. (
Perz-Edwards, A., Hardison, N. L., and Linney, E. (
Peterson, R. T., Link, B. A., Dowling, J. E., and Schreiber, S. L. (
Peterson, R. T., Mably, J. D., Chen, J. N., and Fishman, M. C. (
Peterson, R. E., Theobald, H. M., and Kimmel, G. L. (
Phillips, R. B., and Reed, K. M. (
Postlethwait, J., Amores, A., Force, A., and Yan, Y. L. (
Prasch, A. L., Heideman, W., and Peterson, R. E. (
Prasch, A. L., Tanguay, R. L., Heideman, W., and Peterson, R. E. (
Prasch, A. L., Teraoka, H., Carney, S. A., Dong, W., Hiraga, T., Stegeman, J. J., Heideman, W., and Peterson, R. E. (
Prince, R., and Cooper, K. R. (
Quint, E., Smith, A., Avaron, F., Laforest, L., Miles, J., Gaffield, W., and Akimenko, M. A. (
Ree, G. E., and Payne, J. F. (
Roche, H., Boge, G., and Peres, G. (
Roex, E. W., Giovannangelo, M., and van Gestel, C. A. (
Roex, E. W., Keijzers, R., and van Gestel, C. A. (
Rougier, F., Menudier, A., Bosgiraud, C., and Nicolas, J. A. (
Samson, J. C., Goodridge, R., Olobatuyi, F., and Weis, J. S. (
Sbrogna, J. L., Barresi, M. J., and Karlstrom, R. O. (
Schmidt, J. V., and Bradfield, C. A. (
Schreurs, R. H., Legler, J., Artola-Garicano, E., Sinnige, T. L., Lanser, P. H., Seinen, W., and Van der Burg, B. (
Schwetz, B. A., Norris, J. M., Sparsch, G. L., Rowe, U. K., Gehring, P. J., Emerson, J. L., and Gerbig, C. G. (
Sehnert, A. J., Huq, A., Weinstein, B. M., Walker, C., Fishman, M., and Stainier, D. Y. R. (
Skidmore, J. F. (
Spitsbergen, J. M., and Kent, M. L. (
Spitsbergen, J. M., Tsai, H. W., Reddy, A., Miller, T., Arbogast, D., Hendricks, J. D., and Bailey, G. S. (
Stainier, D. Y., Fouquet, B., Chen, J. N., Warren, K. S., Weinstein, B. M., Meiler, S. E., Mohideen, M. A., Neuhauss, S. C., Solnica-Krezel, L., Schier, A. F., et al. (
Stainier, D. Y. R., Lee, R. K., and Fishman, M. C. (
Stanton, M. F. (
Stenkamp, D. L., and Frey, R. A. (
Strmac, M., and Braunbeck, T. (
Stuart, G. W., McMurray, J. V., and Westerfield, M. (
Summerton, J., and Weller, D. (
Tanguay, R. L., Abnet, C. C., Heideman, W., and Peterson, R. E. (
Tanguay, R. L., Andreasen, E., Heideman, W., and Peterson, R. E. (
Tanguay, R. L., Andreasen, E., Walker, M. K., and Peterson, R. E. (
Teraoka, H., Dong, W., Hiraga, T. (
Teraoka, H., Dong, W., Ogawa, S., Tsukiyama, S., Okuhara, Y., Niiyama, M., Ueno, N., Peterson, R. E., and Hiraga, T. (
Teraoka, H., Dong, W., Tsujimoto, Y., Iwasa, H., Endoh, D., Ueno, N., Stegeman, J. J., Peterson, R. E., Hiraga, T. (
Thermes, V., Grabher, C., Ristoratore, F., Bourrat, F., Choulika, A., Wittbrodt, J., and Joly, J. S. (
Todd, N. E., and Van Leeuwen, M. (
Van den Belt, K., Verheyen, R., and Witters, H. (
van Eeden, F. J., Granato, M., Schach, U., Brand, M., Furutani-Seiki, M., Haffter, P., Hammerschmidt, M., Heisenberg, C. P., Jiang, Y. J., Kane, D. A., et al. (
van Heyningen, V. (
Vandersea, M. W., Fleming, P., McCarthy, R. A., and Smith, D. G. (
Vascotto, S. G., Beckham, Y., and Kelly, G. M. (
Vos, J. G., Moore, J. A., and Zinkl, J. G. (
Wakamatsu, Y., Pristyazhnyuk, S., Kinoshita, M., Tanaka, M., and Orzato, K. (
Walker, M. K., Hufnagle, L. C., Clayton, M. K., and Peterson, R. E. (
Walker, M. K., and Peterson, R. E. (
Walker, M. K., and Peterson, R. E. (
Walker, M. K., and Peterson, R. E. (
Ward, A. C., and Lieschke, G. J. (
Westerfield, M. (
Wiegand, C., Krause, E., Steinberg, C., and Pflugmacher, S. (
Willey, J. B., and Krone, P. H. (
Wienholds, E., Schulte-Merker, S., Walderich, B., and Plasterk, R. H. A. (
Wisk, J. D., and Cooper, K. R. (
Wisk, J. D., and Cooper, K. R. (
Wittbrodt, J., Shima, A., and Schartl, M. (
Xu, X., Meiler, S. E., Zhong, T. P., Mohideen, M., Crossley, D. A., and Bruggren, W. W. (
Yamazaki, K., Teraoka, H., Dong, W., Stegeman, J. J., Hiraga, T. (
Zodrow, J. M., and Tanguay, R. L. (
Zok, S., Gorge, G., Kalsch, W., and Nagel, R. (
Author notes
*School of Pharmacy, University of Wisconsin, Madison, Wisconsin, 53705; †Department of Toxicology, School of Veterinary Medicine, Rakuno Gakuen University, Ebetsu, Japan; and ‡Molecular and Environmental Toxicology Center, University of Wisconsin, Madison, Wisconsin, 53705

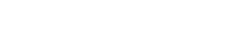


Comments